Product Info Summary
| SKU: | PA1665 |
|---|---|
| Size: | 100 μg/vial |
| Reactive Species: | Human, Mouse, Rat |
| Host: | Rabbit |
| Application: | Flow Cytometry, IHC, WB |
Customers Who Bought This Also Bought
Product info
Product Name
Anti-CIAS1/NALP3/NLRP3 Antibody Picoband®
SKU/Catalog Number
PA1665
BA3677 is an alternative SKU for this antibody, used in previous lots.
Size
100 μg/vial
Form
Lyophilized
Description
Boster Bio Anti-CIAS1/NALP3/NLRP3 Antibody catalog # PA1665. Tested in Flow Cytometry, IHC, WB applications. This antibody reacts with Human, Mouse, Rat. The brand Picoband indicates this is a premium antibody that guarantees superior quality, high affinity, and strong signals with minimal background in Western blot applications. Only our best-performing antibodies are designated as Picoband, ensuring unmatched performance.
Storage & Handling
Store at -20˚C for one year from date of receipt. After reconstitution, at 4˚C for one month. It can also be aliquotted and stored frozen at -20˚C for six months. Avoid repeated freeze-thaw cycles.
Cite This Product
Anti-CIAS1/NALP3/NLRP3 Antibody Picoband® (Boster Biological Technology, Pleasanton CA, USA, Catalog # PA1665)
Host
Rabbit
Contents
Each vial contains 4 mg Trehalose, 0.9 mg NaCl and 0.2 mg Na2HPO4.
Clonality
Polyclonal
Isotype
Rabbit IgG
Immunogen
A synthetic peptide corresponding to a sequence at the N-terminus of human CIAS1, different from the related rat and mouse sequences by one amino acid.
Cross-reactivity
No cross-reactivity with other proteins
Reactive Species
PA1665 is reactive to NLRP3 in Human, Mouse, Rat
Observed Molecular Weight
110 kDa
Calculated molecular weight
118.2 kDa
Background of NLRP3
NLRP3 (NLR FAMILY, PYRIN DOMAIN-CONTAINING 3), also known as CIAS1, CRYOPYRIN, NALP3 or PYPAF1, is a protein that in humans is encoded by the NLRP3 (NOD-like receptor family, pryin domain containing 3) gene. The NLRP3 gene encodes a pyrin-like protein expressed predominantly in peripheral blood leukocytes. And the NLRP3 gene is mapped on 1q44. NLRP3 interacts with apoptosis-associated speck-like protein containing a CARD (ASC). The encoded protein may play a role in the regulation of inflammation and apoptosis. Mutation of the NALP3 nucleotide-binding domain reduced ATP binding, CASP1 activation, IL1B production, cell death, macromolecular complex formation, self-association, and association with ASC. Consistent with an essential role for Nlrp3 inflammasomes in antifungal immunity, Gross et al.showed that Nlrp3-deficient mice are hypersusceptible to C. albicans infection. Activation of the NLRP3 inflammasome in response to virus or to RNA was dependent upon lysosomal maturation and reactive oxygen species production in human cells. The NLRP3 inflammasome senses obesity-associated danger signals and contributes to obesity-induced inflammation and insulin resistance.
Antibody Validation
Boster validates all antibodies on WB, IHC, ICC, Immunofluorescence, and ELISA with known positive control and negative samples to ensure specificity and high affinity, including thorough antibody incubations.
Application & Images
Applications
PA1665 is guaranteed for Flow Cytometry, IHC, WB Boster Guarantee
Assay Dilutions Recommendation
The recommendations below provide a starting point for assay optimization. The actual working concentration varies and should be decided by the user.
Western blot, 0.1-0.5μg/ml, Human, Mouse, Rat
Immunohistochemistry (Paraffin-embedded Section), 2-5μg/ml, Human
Flow Cytometry(Fixed), 1-3μg/1x106 cells, Human
Positive Control
WB: rat thymus tissue, mouse thyus tissue, mouse RAW264.7 whole cell, mouse J774A.1 whole cell, human THP-1 whole cell
IHC: human colon cancer tissue
FCM: THP-1 cell
Validation Images & Assay Conditions

Click image to see more details
Western blot analysis of NLRP3 using anti-NLRP3 antibody (PA1665).
Electrophoresis was performed on a 8% SDS-PAGE gel at 80V (Stacking gel) / 120V (Resolving gel) for 2 hours. The sample well of each lane was loaded with 30 ug of sample under reducing conditions.
Lane 1: rat thymus tissue lysates,
Lane 2: mouse thyus tissue lysates,
Lane 3: mouse RAW264.7 whole cell lysates,
Lane 4: mouse J774A.1 whole cell lysates.
After electrophoresis, proteins were transferred to a nitrocellulose membrane at 150 mA for 50-90 minutes. Blocked the membrane with 5% non-fat milk/TBS for 1.5 hour at RT. The membrane was incubated with rabbit anti-NLRP3 antigen affinity purified polyclonal antibody (PA1665) at 0.5 μg/mL overnight at 4°C, then washed with TBS-0.1%Tween 3 times with 5 minutes each and probed with a goat anti-rabbit IgG-HRP secondary antibody (Catalog # BA1054) at a dilution of 1:5000 for 1.5 hour at RT. The signal is developed using an ECL Plus Western Blotting Substrate (Catalog # AR1196-200) with Tanon 5200 system. A specific band was detected for NLRP3 at approximately 110 kDa. The expected band size for NLRP3 is at 118kDa.

Click image to see more details
Western blot analysis of NLRP3 using anti-NLRP3 antibody (PA1665).
Electrophoresis was performed on a 8% SDS-PAGE gel at 80V (Stacking gel) / 120V (Resolving gel) for 2 hours. The sample well of each lane was loaded with 30 ug of sample under reducing conditions.
Lane 1:human THP-1 whole cell lysates.
After electrophoresis, proteins were transferred to a nitrocellulose membrane at 150 mA for 50-90 minutes. Blocked the membrane with 5% non-fat milk/TBS for 1.5 hour at RT. The membrane was incubated with rabbit anti-NLRP3 antigen affinity purified polyclonal antibody (PA1665) at 0.5 μg/mL overnight at 4°C, then washed with TBS-0.1%Tween 3 times with 5 minutes each and probed with a goat anti-rabbit IgG-HRP secondary antibody (Catalog # BA1054) at a dilution of 1:5000 for 1.5 hour at RT. The signal is developed using an ECL Plus Western Blotting Substrate (Catalog # AR1196-200) with Tanon 5200 system. A specific band was detected for NLRP3 at approximately 110 kDa. The expected band size for NLRP3 is at 118kDa.

Click image to see more details
IHC analysis of NLRP3 using anti-NLRP3 antibody (PA1665).
NLRP3 was detected in a paraffin-embedded section of human colon cancer tissue. Heat mediated antigen retrieval was performed in EDTA buffer (pH 8.0, epitope retrieval solution). The tissue section was blocked with 10% goat serum. The tissue section was then incubated with 2 μg/ml rabbit anti-NLRP3 Antibody (PA1665) overnight at 4°C. Peroxidase Conjugated Goat Anti-rabbit IgG was used as secondary antibody and incubated for 30 minutes at 37°C. The tissue section was developed using HRP Conjugated Rabbit IgG Super Vision Assay Kit (Catalog # SV0002) with DAB as the chromogen.

Click image to see more details
Flow Cytometry analysis of THP-1 cells using anti-NLRP3 antibody (PA1665).
Overlay histogram showing THP-1 cells stained with PA1665 (Blue line). To facilitate intracellular staining, cells were fixed with 4% paraformaldehyde and permeabilized with permeabilization buffer. The cells were blocked with 10% normal goat serum. And then incubated with rabbit anti-NLRP3 Antibody (PA1665, 1 μg/1x106 cells) for 30 min at 20°C. DyLight®488 conjugated goat anti-rabbit IgG (BA1127, 5-10 μg/1x106 cells) was used as secondary antibody for 30 minutes at 20°C. Isotype control antibody (Green line) was rabbit IgG (1 μg/1x106) used under the same conditions. Unlabelled sample without incubation with primary antibody and secondary antibody (Red line) was used as a blank control.
Click image to see more details
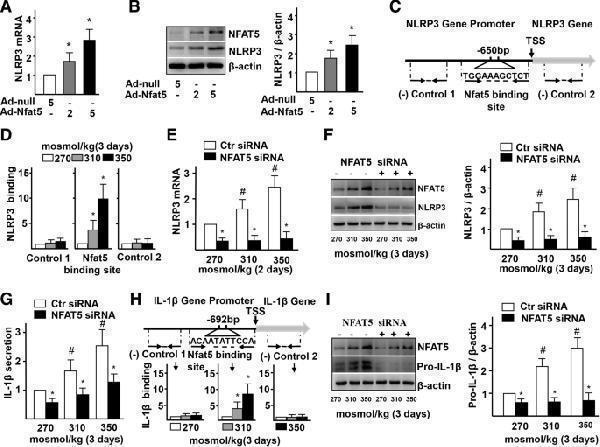
High-salt-elevated NFAT5 mediates transcription of NLRP3 and IL-1β in ECs. a-b mRNA and protein levels of NLRP3 in ECs treated with Adenovirus-null (Ad-null, 5 MOI) and Adenovirus-NFAT5 (Ad-NFAT5, 2 MOI or 5 MOI). c-d High-salt increases binding of NFAT5 to the promoter region of IL-1β. Diagram showing the region of the NFAT5 binding site upstream of the transcription start site (TSS) of NLRP3, and the regions that were used to analyze NFAT5 binding by ChIP. ChIP results are relative to 270 mosmol/kg. e-f mRNA and protein levels of NLRP3 in ECs treated with high-salt and transfected with Ctr siRNA or NFAT5 siRNA. g Protein secretion of IL-1β in ECs treated with high-salt and transfected with Ctr siRNA or NFAT5 siRNA. h High-salt increases binding of NFAT5 to the promoter region of IL-1β. i Protein levels of pro-IL-1β in ECs treated with high-salt and transfected with Ctr siRNA or NFAT5 siRNA. All data were presented as mean ± SEM, N ≥ 3. * p < 0.05
Index in PubMed under a CC BY license. PMID: 31429763
Click image to see more details
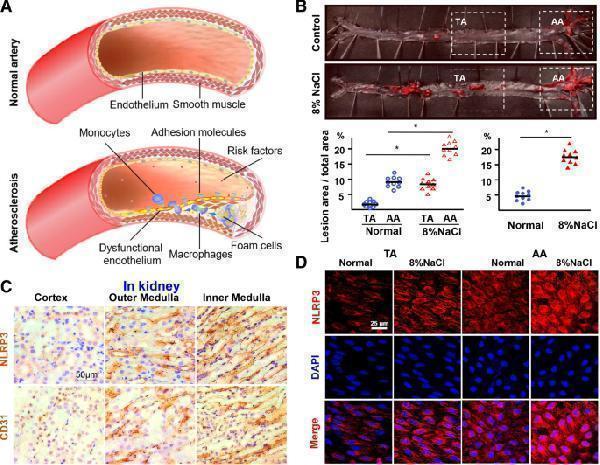
High-salt intake predisposes atherosclerosis and upregulates NLRP3 expression. a Schematic shows the process of atherosclerosis formation. b Oil Red O staining of aortas and quantification of percentage lesion areas in the thoracic aorta (TA) and aortic arch (AA) of ApoE −/− mice ( n = 10) fed with a normal or high-salt diet for 12 weeks. c Immunohistochemistry staining for NLRP3 and CD31 in the kidney at 4 weeks. Nuclei, hematoxylin staining. d En face immunofluorescent staining of NLRP3 of ECs in TA and AA of ApoE−/− mice fed with a normal or high-salt diet for 4 weeks. See Additional file : Figure S1D for NLRP3 mRNA qPCR. All data were presented as mean ± SEM, N ≥ 9. * p < 0.05
Index in PubMed under a CC BY license. PMID: 31429763
Click image to see more details
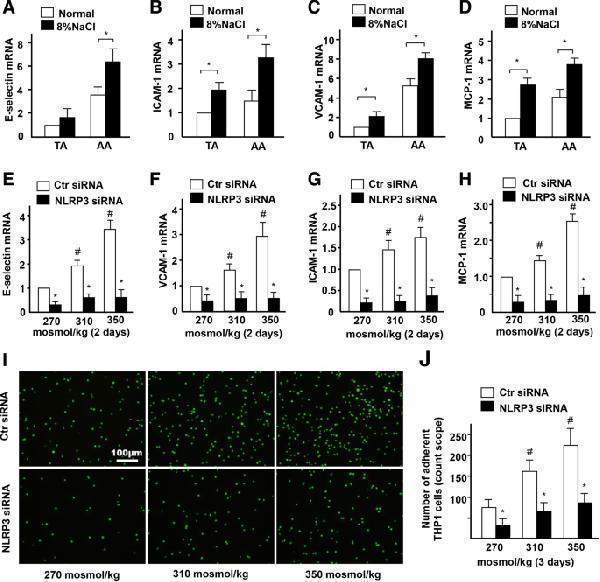
High-salt elevates endothelial inflammation via NLRP3. a-d Quantification of mRNA levels of E-selectin, ICAM-1, VCAM-1 and MCP-1 in TA and AA of ApoE−/− mice fed with a normal or high-salt diet. e-h Quantification of mRNA levels of E-selectin, VCAM-1, ICAM-1, and MCP-1 in ECs treated with high-salt and transfected with Control siRNA or NLRP3 siRNA. i, j The adhesion of Calcein-labeled THP-1 monocytes in ECs treated with high-salt and transfected with Control siRNA or NLRP3 siRNA. All data were presented as mean ± SEM, N ≥ 3. * p < 0.05
Index in PubMed under a CC BY license. PMID: 31429763
Click image to see more details
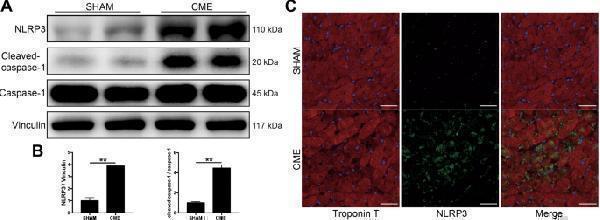
NLRP3 and cleaved-caspase-1 are upregulated in mice heart 3 days after CME intervention. A Representative immunoblots of NLRP3 and cleaved-caspase-1 expression in sham and CME mice hearts. B Densitometric analysis of relative protein expressions, Vinculin is used as loading control. n = 4 per group. Data represent the mean ± SEM of three replicates. C Representative images of immunofluorescent double staining of Troponin T and NLRP3 in cardiac tissues. Scale bars = 50 µm. n = 4 per group. ∗ P < 0.05, ∗ ∗ P < 0.01.
Index in PubMed under a CC BY license. PMID: 33436548
Click image to see more details
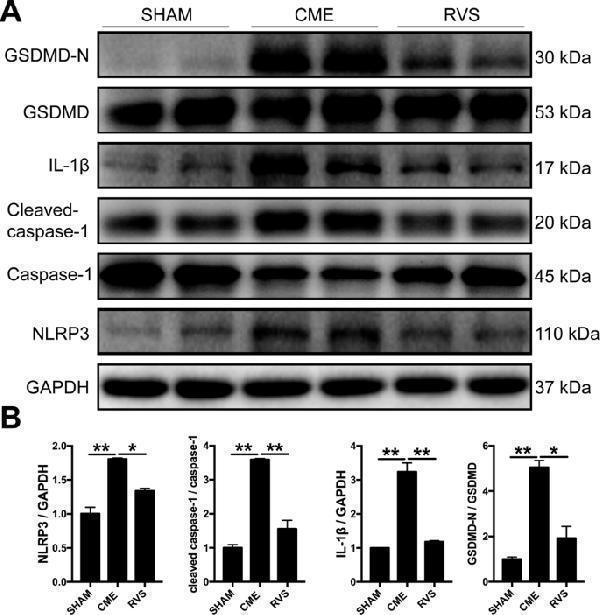
RVS restrains the activation of the NLRP3 inflammasome at 3 days after CME intervention. A Representative immunoblots for myocardial NLRP3, caspase-1, cleaved-caspase-1, IL-1β, GSDMD, GSDMD-N in the hearts of sham, CME and RVS-treated mice. B The densitometric analysis of relative protein expressions, GAPDH is used as loading control. n = 4 per group. Data represent the mean ± SEM of three replicates. ∗ P < 0.05, ∗ ∗ P < 0.01.
Index in PubMed under a CC BY license. PMID: 33436548
Click image to see more details
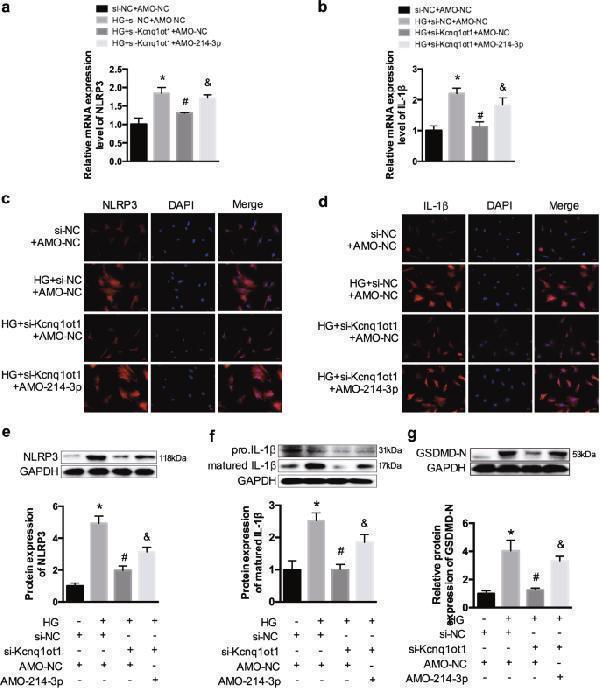
The texts overlap in figure 6g. So I want to replace it with a new in the attachment. Kcnq1ot1/miR-214-3p pathway regulates inflammation in fibroblasts a , b qRT-PCR was conducted to detect expression of NLRP3 and IL-1β. c , d The expression levels of NLRP3 and IL-1β were determined by immunofluorescence. e – g Western blot was performed to determine the expression of NLRP3, IL-1β, and GSDMD-N. * P < 0.05 compared with the si-NC + AMO-NC group, # P < 0.05 compared with the HG + si-Kcnq1ot1 + AMO-NC group, & P < 0.05 compared with the HG + si-Kcnq1ot1 + AMO-214-3p group. n = 3 in each group
Index in PubMed under a CC BY license. PMID: 30250027
Click image to see more details
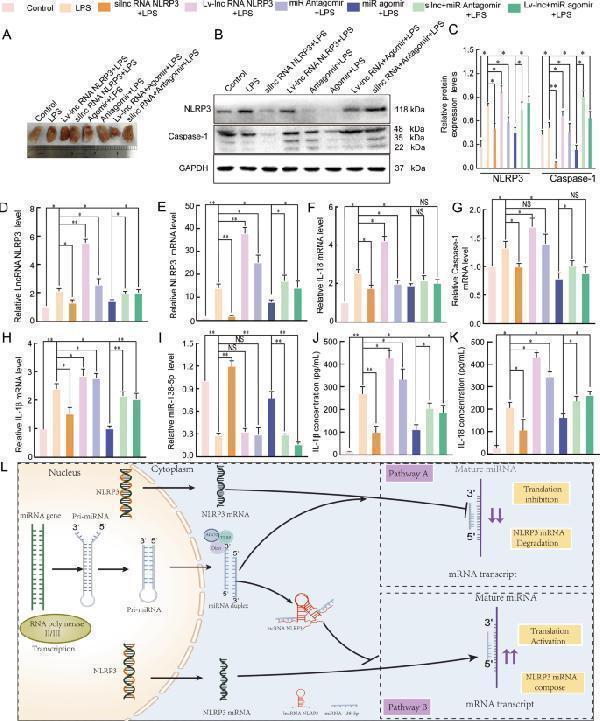
The mechanism by which the lncRNA NLRP3/miR-138-5p/NLRP3 ceRNET functions in the inflammatory response. The lungs of rats were injected with PBS in the control group and LPS-treated rats were further treated with si-r-lncRNA NLRP3, Lv-lncRNA NLRP3, agomiR-138-5p, antagomiR-138-5p, Lv-lncRNA NLRP3 + agomiR-138-5p, and si-r-lnc NLRP3 + antagomiR-138-5p. A Morphometric changes in the appearance of the lungs that had been fixed in 4% paraformaldehyde for 24 h at 25 °C in each group. B , C The protein expression levels of NLRP3 and caspase-1 in rat lung tissues. qRT-PCR assays were used to analyse mRNA expression of D lncRNA NLRP3, E NLRP3, F IL-18, G Caspase-1, H IL-1β, and I miR-138-5p in the lung tissues of rats. ELISA analysis of the IL-1β ( J ) and IL-18 ( K ) levels in the culture supernatant. L Graphical summary of the role of the lncRNA NLRP3/miR-138-5p/NLRP3 ceRNET in acute lung injury. β-Actin was used as the reference. The data are presented as mean ± SE ( n = 6). * P < 0.05; ** P < 0.01; *** P < 0.001; NS, no statistically significant difference.
Index in PubMed under a CC BY license. PMID: 34599154
Click image to see more details
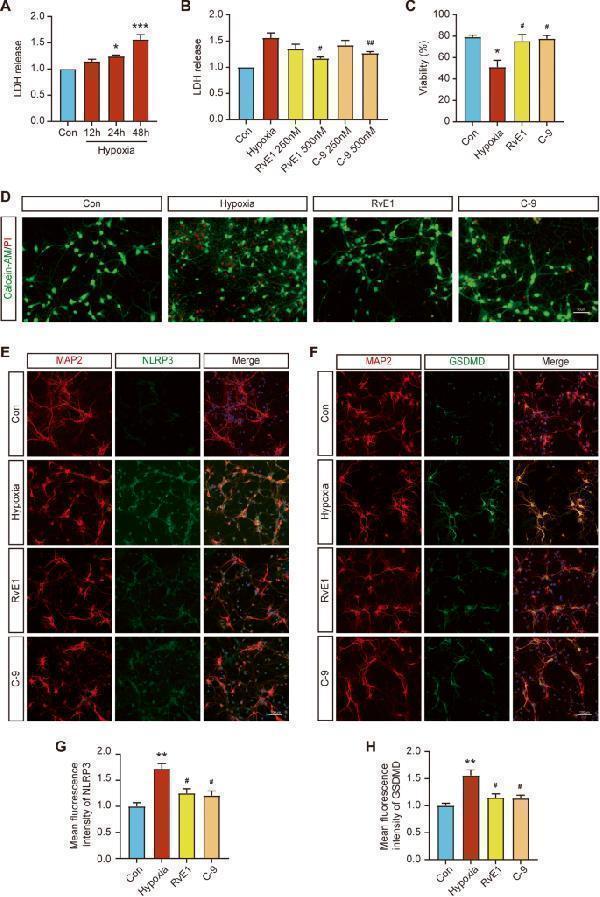
ChemR23 activation ameliorated pyroptosis after chronic hypoxic stimulation in primary rat hippocampal neurons. A Changes in LDH release in the cell culture supernatant with time under hypoxic conditions. n = 4 per group. B Effects of different concentrations of RvE1 and C-9 on the release of LDH in primary neurons under hypoxic stimulation. n = 4 per group. C Cell viability of each group under different treatment conditions. n = 3 per group. D Fluorescence images of primary neurons stained with calcein AM (live cells, green fluorescence) and PI (dead cells, red fluorescence) after different treatments. Scale bar: 50 μm. n = 3 per group. E Double immunostaining of NLRP3 with MAP2 in primary neurons of different groups. Scale bar: 100 μm. n = 3 for each group. F Double immunostaining of GSDMD with MAP2 in primary neurons of different groups. Scale bar: 100 μm. n = 3 for each group. G The statistics of the mean fluorescence intensity of NLRP3 in each group. n = 3 for each group. H The statistics of the mean fluorescence intensity of GSDMD in each group. n = 3 for each group. Data are presented as the mean ± SEM. *P < 0.05, **P < 0.01, ***P < 0.001 vs. Con; #P < 0.05, ##P < 0.01 vs. Hypoxia.
Index in PubMed under a CC BY license. PMID: 37932279
Click image to see more details
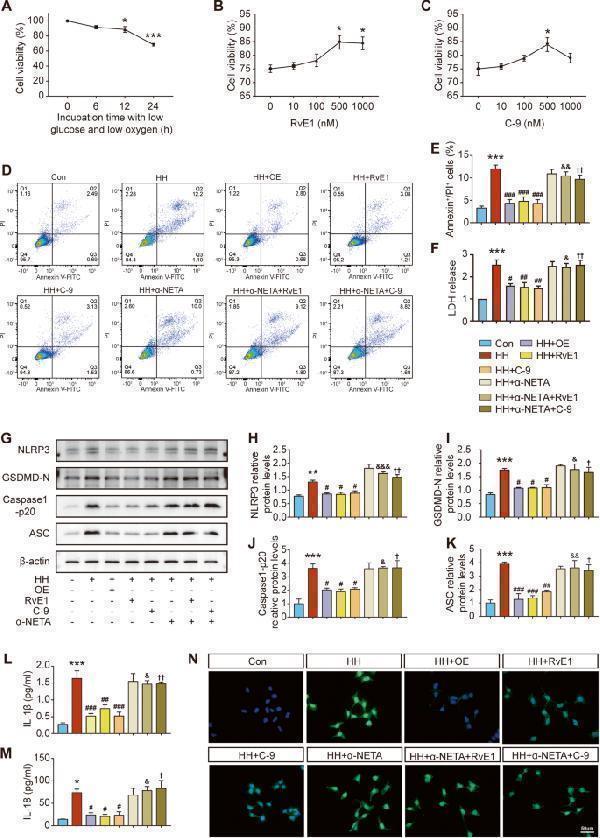
Effects of ChemR23 activation on pyroptosis in SH-SY5Y cells after hypoglycaemic and hypoxic stimulation. A Changes in cell viability with time under hypoglycaemic and hypoxic conditions. n = 4 per group. B , C Effects of different concentrations of RvE1 and C-9 on cell viability. n = 4 per group. D , E The proportion of Annexin V/PI double positive cells analysed by flow cytometry. n = 3 per group. F LDH levels in the cell culture supernatant. n = 3 per group. G – K Western blot analysis of NLRP3, GSDMD-N, Caspase-1 p20 and ASC. n = 3 per group. L , M Concentrations of IL-1β and IL-18 in the cell culture supernatant were measured by ELISA. n = 4 per group. (N) Representative immunofluorescence staining of NLRP3 (green) with the nuclei counterstained by DAPI. Scale bar: 50 μm. n = 3 per group. Data are presented as the mean ± SEM. **P < 0.01, ***P < 0.001 vs. Con; #P < 0.05, ##P < 0.01, ###P < 0.001 vs. HH; &P < 0.05, &&P < 0.01, &&&P < 0.001 vs. HH+RvE1; †P < 0.05, ††P < 0.01, †††P < 0.001 vs. HH + C-9.
Index in PubMed under a CC BY license. PMID: 37932279
Click image to see more details
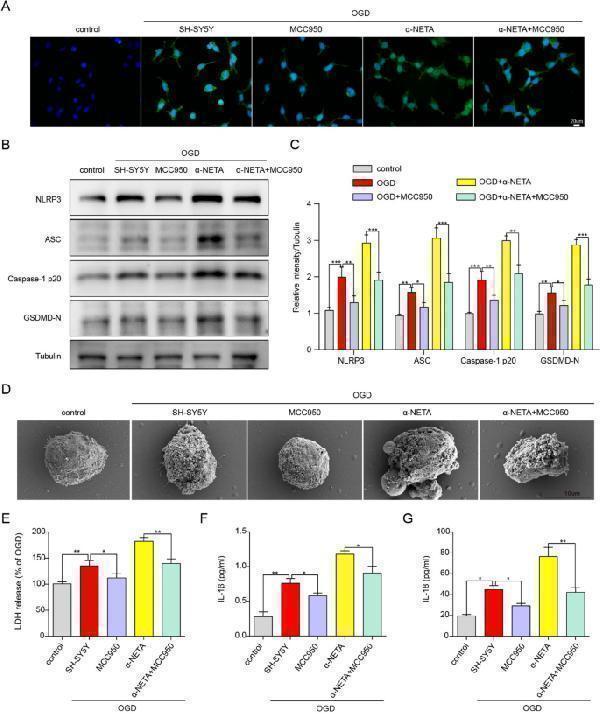
The neuroprotective effect of ChemR23 against OGD-induced neuronal pyroptosis was NLRP3 dependent. A Representative immunofluorescent images of NLRP3 in cultured neurons. Scale bar = 20 μm. B , C Immunoblots and quantitative analysis of NLRP3, ASC, caspase-1 p20, GSDMD-N in treated neurons after OGD. D Representative scanning electron microscopy pictures of neurons. Scale bar = 10 μm. E LDH release was assessed and quantified in neurons subjected to OGD for 4 h. F , G ELISA analysis for IL-1β and IL-18 levels in supernatants of neuron subjected to OGD for 4 h. At least three independent experiments were repeated. Data are represented as mean ± SD. *P < 0.05, **P < 0.01, ***P < 0.001
Index in PubMed under a CC BY license. PMID: 38178174
Click image to see more details
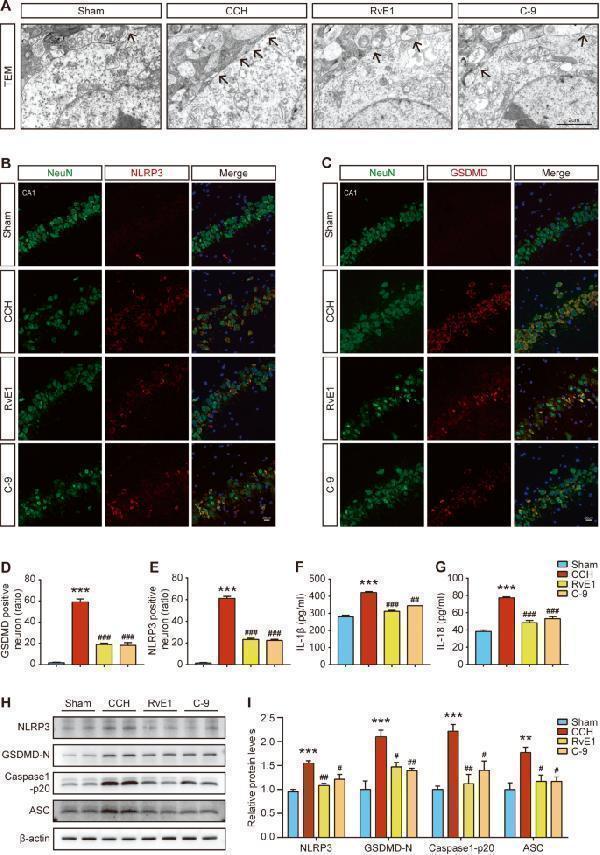
Effects of ChemR23 activation with RvE1 or C-9 on neuronal pyroptosis in CCH rats. A Representative transmission electron micrographs of hippocampal neurons in different treatment groups. Black arrowhead: membrane pores. Scale bar: 2 μm. n = 3 per group. B , C Representative double immunofluorescence staining of GSDMD/NeuN and NLRP3/NeuN in the rat hippocampus. Scale bar: 20 μm. n = 3 per group. D , E Quantitative statistics of GSDMD/NeuN or NLRP3/NeuN double-positive cells in rat hippocampus. n = 3 per group. F , G ELISA results of IL-1 and IL-18 analysis in the rat hippocampus. n = 4 per group. H , I Representative immunoblotting bands and the expression levels of NLRP3, GSDMD-N, Caspase-1 p20 and ASC in the rat hippocampus. n = 4 per group. Data are presented as the mean ± SEM. **P < 0.01, ***P < 0.001 vs. sham; #P < 0.05, ##P < 0.01, ###P < 0.001 vs. CCH.
Index in PubMed under a CC BY license. PMID: 37932279
Click image to see more details
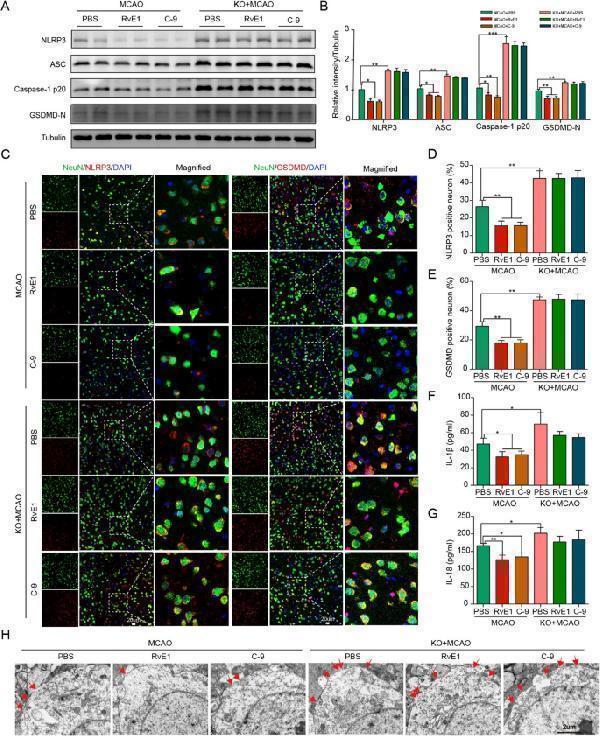
RvE1 and C-9 ameliorated NLRP3 inflammasome-mediated pyroptosis in MCAO mice. A , B Immunoblots and quantitative analysis of NLRP3, ASC, caspase-1 p20, GSDMD-N (n = 3/per group). C – E Double immunofluorescent staining of NLRP3 or GSDMD with NeuN in ischemic penumbra region and quantitative analysis at Day 1 after MCAO (n = 3/per group). Scale bar = 20 μm. F , G The ELISA assays for IL-1β and IL-18 levels in ipsilateral brain tissues subjected to ischemia at Day 1 (n = 4/per group). H Representative microphotographs and quantification of neuronal death based on NeuN and TUNEL assay in the ischemic ipsilateral brain regions at Day 1 after MCAO (n = 3/per group). Scale bar = 2 μm. Data are represented as mean ± SD. *P < 0.05, **P < 0.01, ***P < 0.001
Index in PubMed under a CC BY license. PMID: 38178174
Click image to see more details
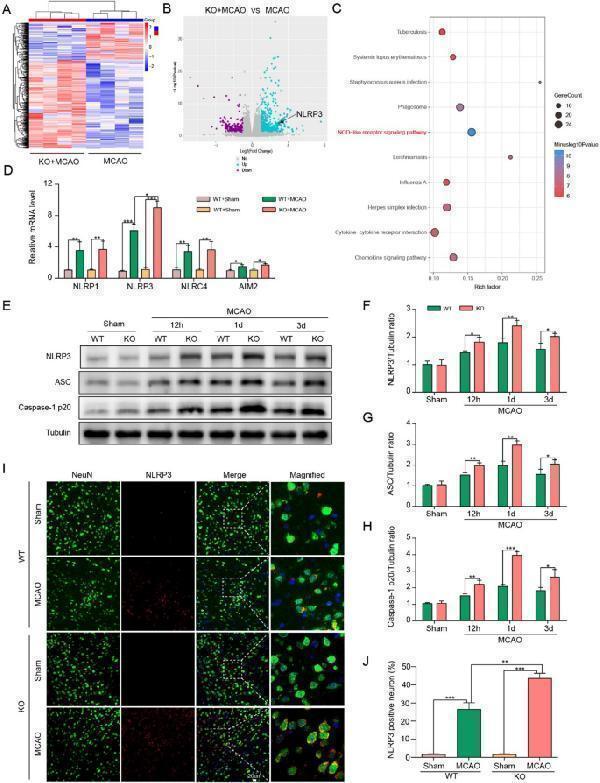
ChemR23 deletion amplified NLRP3 inflammasome activation in cerebral ischemia injury. A , B Thermograms and volcano plots showing the comparison between ChemR23 KO group and WT group at Day 1 after MCAO (n = 4/per group). C The KEGG pathway enrichment analysis (n = 4/per group). D qPCR analysis of the infammasome sensor genes NLRP1, NLRP3, NLRC4, and AIM2 ipsilateral hemispheres of WT mice and KO mice subjected to MCAO at Day 1 (n = 3/per group). E – H Western blotting and quantitative analysis of NLRP3, ASC, caspase-1 p20 expression in ischemic penumbra tissue at 12 h, Day 1 and Day 3 after MCAO (n = 3/per group). I , J Representative immunofluorescence staining images of NLRP3 were co-stained with NeuN in peri-infarct areas and their quantification at Day 1 after MCAO (n = 3/per group). Scale bar = 20 μm. Data are represented as mean ± SD. *P < 0.05, **P < 0.01, ***P < 0.001
Index in PubMed under a CC BY license. PMID: 38178174
Click image to see more details
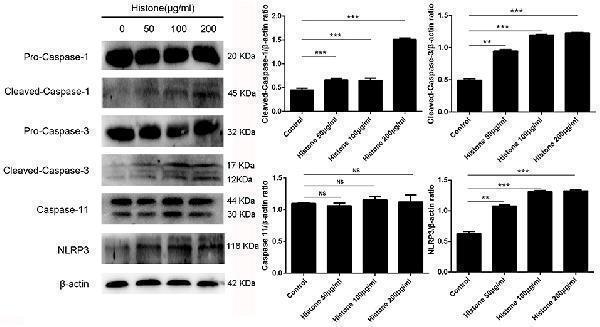
Histone induce BMECs pyroptosis is dependent on activations of caspase 1, caspase 3 and NLRP3. BMECs were incubated with stimulated histone for 16 h, and the activities of these proteins were determined by western blotting. Date are presented as mean ± SEM ( n = 5). P -values of < 0.05 were considered significant (** P < 0.01, *** P < 0.001, and “ns” means not significant).
Index in PubMed under a CC BY license. PMID: 31156617
Click image to see more details
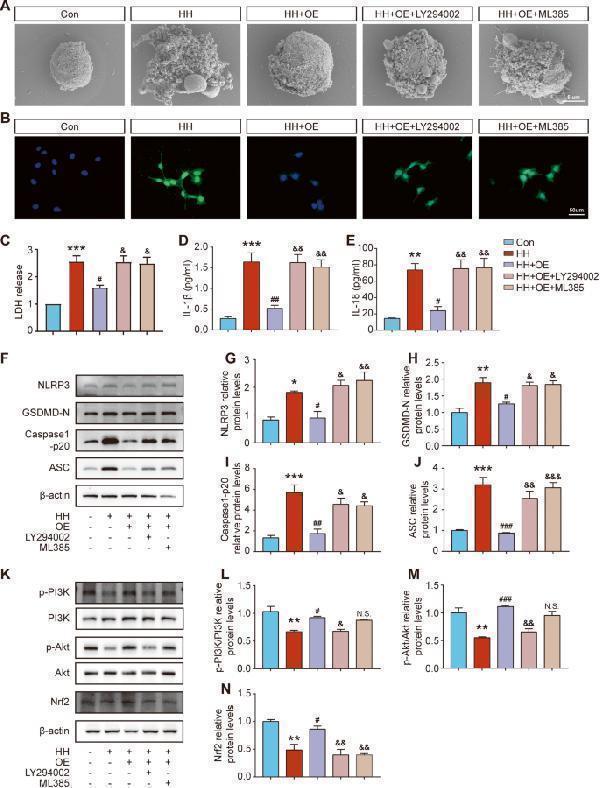
ChemR23 activation inhibited neuronal pyroptosis via the PI3K/AKT/Nrf2 signalling pathway in SH-SY5Y cells. A Scanning electron microscopy analysis showing the cell morphology in different treatment groups. n = 3 per group. B Representative immunofluorescence staining of NLRP3 (green) with the nuclei counterstained by DAPI. Scale bar: 50 μm. n = 3 per group. C LDH levels in the cell culture supernatant. n = 3 per group. D , E ELISA of IL-1β and IL-18 levels. n = 3 per group. F – J The levels of NLRP3, GSDMD-N, Caspase-1 p20 and ASC were evaluated by western blot. n = 3 per group. K – N The expression levels of p-PI3K/PI3K, p-AKT/AKT and Nrf2 were evaluated by western blot. n = 3 per group. Data are presented as the mean ± SEM. *P < 0.05, **P < 0.01, ***P < 0.001 vs. Con; #P < 0.05, ##P < 0.01, ###P < 0.001 vs. HH; &P < 0.05, &&P < 0.01, &&&P < 0.001 vs. HH + OE; N.S. means not significant vs. HH + OE.
Index in PubMed under a CC BY license. PMID: 37932279
Click image to see more details
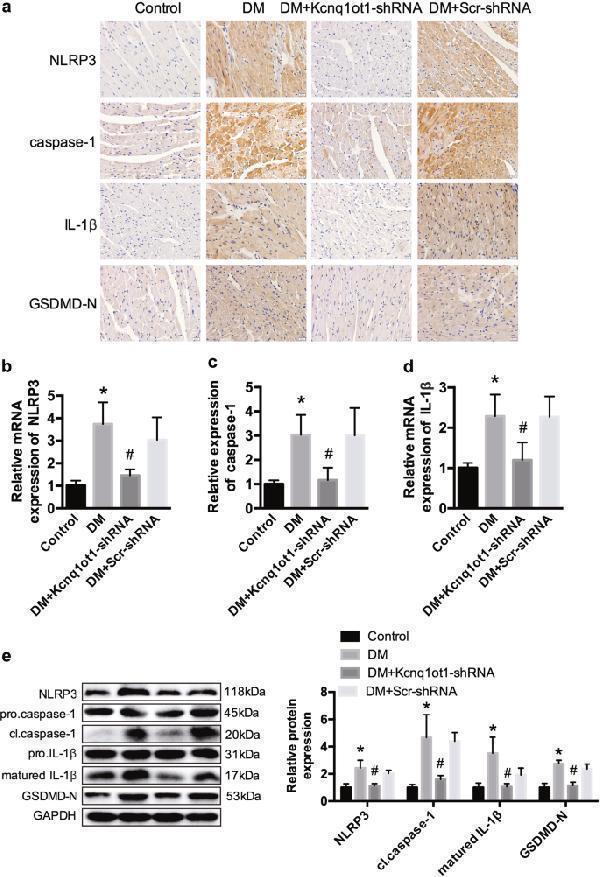
Kcnq1ot1 is involved in the regulation of pyroptosis in vivo a Immunohistochemistry analysis was performed to detect the expression NLRP3, caspase-1, IL-1β, and GSDMD-N. Scale bar, 20 μm. b – d qRT-PCR was conducted to analyze the mRNA expression of NLRP3, caspase-1 and IL-1β. e Western blot was conducted to determine the protein expression of NLRP3, caspase-1 and IL-1β. * P < 0.05 compared with the control group, # P < 0.05 compared with the DM + Scr-shRNA group. n = 5 in each group
Index in PubMed under a CC BY license. PMID: 30250027
Click image to see more details
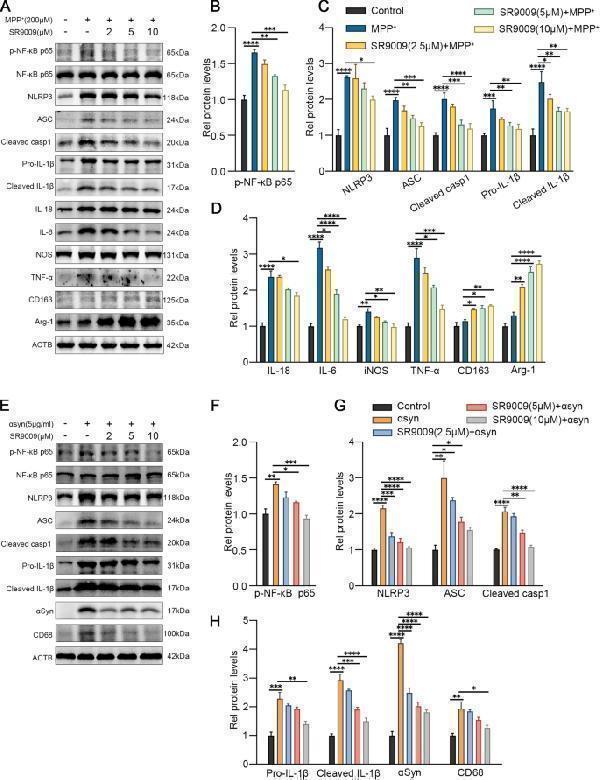
Activation of Rev-erbα inhibited microglial activation induced by MPP + and αsyn PFF. The representative western blot bands ( A ) and the statistical graph ( B – D ) of p-NF-κB p65, NLRP3, ASC, cleaved caspase-1, IL-1β, IL-18, IL-6, TNF-a, iNOS, Arg-1 and CD163 protein expressions. BV2 cells were pretreated with SR9009 (2 μM, 5 μM, 10 μM) for 1 h, then incubated with MPP + for 24 h. The representative western blot bands ( E ) and the statistical graph ( F–H ) of p-NF-κB p65, NLRP3, ASC, cleaved caspase-1, IL-1β, CD68 and αsyn protein expressions. BV2 cells were pretreated with SR9009 (2 μM, 5 μM, 10 μM) for 1 h, then incubated with αsyn pre-formed-fibril for 6 h. The p-NF-κB p65 level was normalized to the total of NF-κB p65, and the rest protein levels were normalized to β-actin. Data were presented as mean ± SEM ( n = 3). (* p < 0.05, ** p < 0.01, *** p < 0.001, **** or p < 0.0001 by One-way ANOVA test)
Index in PubMed under a CC BY license. PMID: 35668454
Click image to see more details
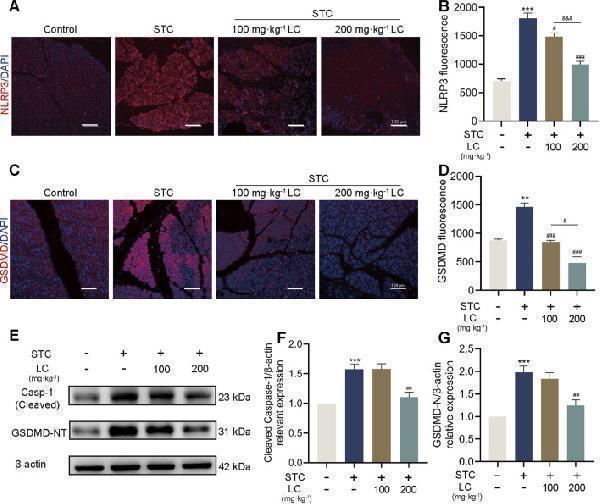
CPT1A protects against SAP in mice by inhibiting the NLRP3/GSDMD-mediated pyroptosis signalling pathway. (A–D) Immunofluorescence staining of NLRP3 and GSDMD in the pancreas of mice received different treatments. Scale bar: 100 μm. (E–G) Protein levels of cleaved Caspase-1 and GSDMD-NT in the pancreas of mice received different treatments. β-actin served as the loading control. All data are presented as means ± SEM, n = 3–5. * p < 0.05, ** p < 0.01, *** p < 0.001 vs. Control group. # p < 0.05, ## p < 0.01, ### p < 0.001 vs. STC group. & p < 0.05, &&& p < 0.001 vs. LC (100 mg/kg) treatment group.
Index in PubMed under a CC BY license. PMID: 40718711
Click image to see more details
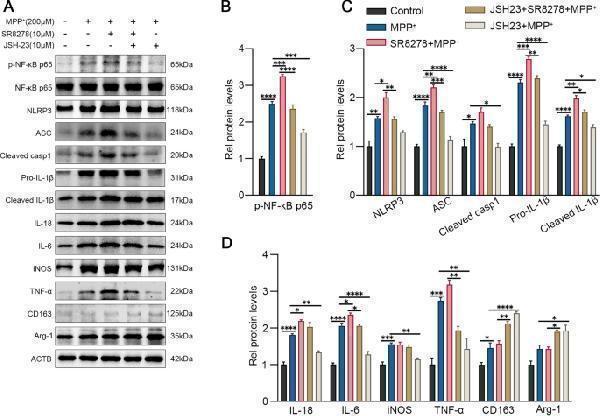
Rev-erbα regulates microglial activation by NF-κB inflammasome pathway. The representative western blot bands ( A ) and the statistical graph ( B – D ) of p-NF-κB p65, NLRP3, ASC, cleaved caspase-1, IL-1β, IL-18, IL-6, TNF-a, iNOS, Arg-1 and CD163 protein expressions. BV2 cells were pretreated with JSH-23 (10 μM) or SR8278 (10 μM) for 1 h, then incubated with MPP + for 24 h. The p-NF-κB p65 level was normalized to the total of NF-κB p65, and the rest protein levels were normalized to β-actin. Data were presented as mean ± SEM (n = 3). (* p < 0.05, ** p < 0.01, *** p < 0.001, **** or p < 0.0001 by One-way ANOVA test)
Index in PubMed under a CC BY license. PMID: 35668454
Click image to see more details
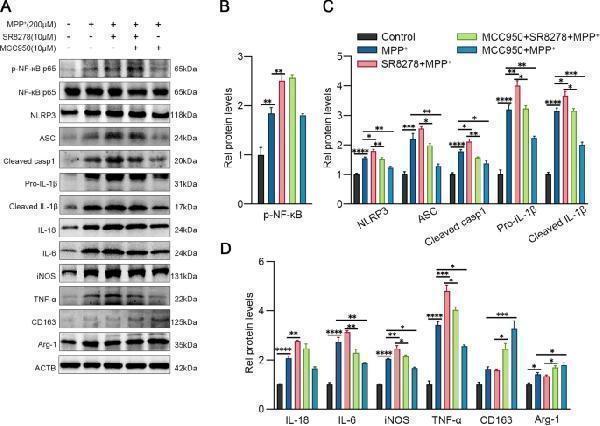
Rev-erbα regulates microglial activation by NLRP3 inflammasome pathway. The representative western blot bands ( A ) and the statistical graph ( B – D ) of p-NF-κB p65, NLRP3, ASC, cleaved caspase-1, IL-1β, IL-18, IL-6, TNF-a, iNOS, Arg-1 and CD163 protein expressions. BV2 cells were pretreated with MCC950 (10 μM) or SR8278 (10 μM) for 1 h, then incubated with MPP + for 24 h. The p-NF-κB p65 level was normalized to the total of NF-κB p65, and the rest protein levels were normalized to β-actin. Data were presented as mean ± SEM ( n = 3). (* p < 0.05, ** p < 0.01, *** p < 0.001, **** or p < 0.0001 by One-way ANOVA test)
Index in PubMed under a CC BY license. PMID: 35668454
Click image to see more details
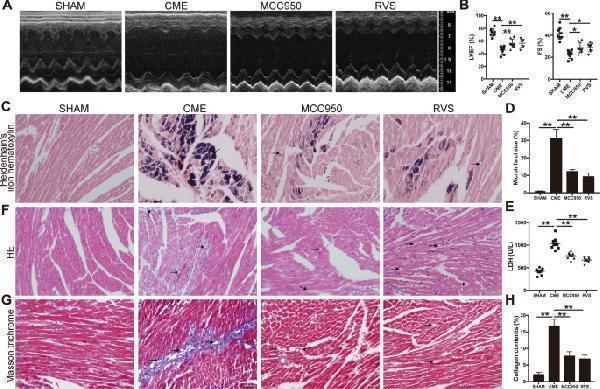
MCC950 and RVS protect against CME-induced cardiac dysfunction and injury. A Representative M-mode echocardiograms for each group at 3 days after CME intervention. Mice were treated with RVS and the selective NLRP3 inflammasome inhibitor MCC950 three days prior and after CME intervention at 40 and 20 mg/kg/d respectively. B Left ventricular ejection fraction (LVEF) and fractional shortening (FS) are measured using Doppler echocardiography. n = 8 to 10 per group. Data represent the mean ± SEM. C Representative images of Heidenhain’s iron hematoxylin staining for each group to visualize microinfarct areas, the microinfarct areas are stained into dark gray. D The quantitative analysis of microinfarct areas. n = 8 to 10 per group. The quantification is representative of the percentage of microinfarct area in 5 fields ± SEM by random. E Serum lactate dehydrogenase (LDH) levels are measured in each group. n = 8 to 10 per group. Data represent the mean ± SEM. F Representative images of HE staining to visualize the local micro-infracted lesions for each group. G Representative images of Masson trichrome staining of the ventricular sections of each group. H The quantitative analysis of collagen contents. n = 8 to 10 per group. The quantification is representative of the percentage of collagen contents in 5 fields ± SEM by random. Scale bars = 50 µm. Black arrows indicate the microspheres. ∗ P < 0.05, ∗ ∗ P < 0.01.
Index in PubMed under a CC BY license. PMID: 33436548
Click image to see more details
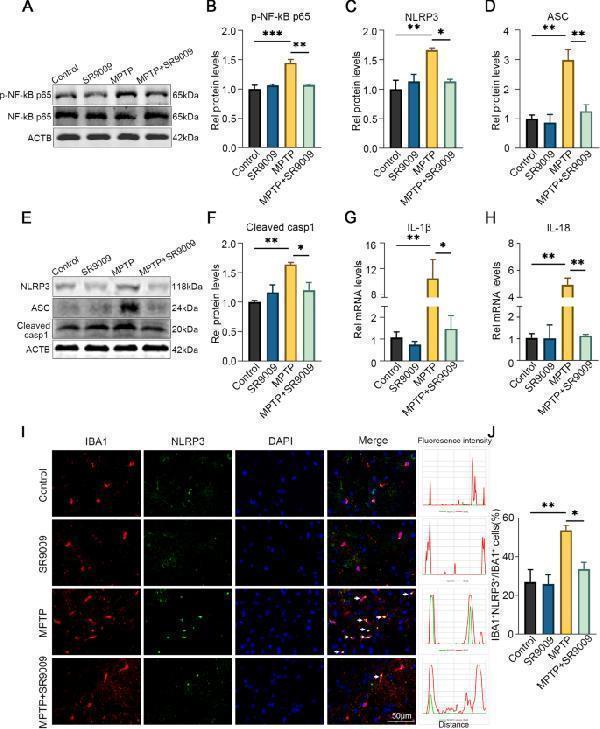
SR9009 suppresses NLRP3 inflammasome activation in the SN of MPTP-induced mice. The representative western blot bands ( A ) and the statistical graph ( B ) of p-NF-κB p65 and NF-κB p65 in the SN. The representative western blot bands ( E ) and the statistical graph ( C – D , F ) of NLRP3, ASC and cleaved-caspase-1 in the SN. The real-time PCR results of IL-1β ( G ) and IL-18 ( H ) in the SN. I Representative double-immunofluorescent staining of IBA1 (red) and NLRP3 (green) in the SN and the statistical graph ( J ) of IBA1 + NLRP3 + /IBA1 + cells. Scale bar, 50 μm. n = 3–4 for each group. The p-NF-κB p65 level was normalized to the total of NF-κB p65, and the rest protein levels were normalized to β-actin. Data were presented as mean ± SEM. (* p < 0.05, ** p < 0.01, or *** p < 0.001 by One-way ANOVA test)
Index in PubMed under a CC BY license. PMID: 35668454
Click image to see more details
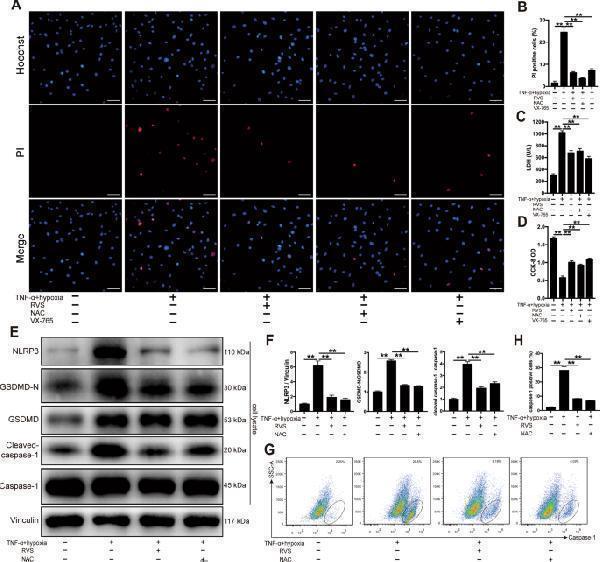
RVS and ROS scavenger inhibit the activation of the NLRP3 inflammasome and pyroptosis in H9c2 cells. A The H9c2 cells are stimulated with 40 ng/ml TNF-α combined with hypoxia for 12 h to active the NLRP3 inflammasome. The caspase-1 selective inhibitor VX-765 and ROS scavenger N-acetylcysteine (NAC) are added 2 h before the stimulation at 20 μM and 5 mM respectively. The pyroptotic cells are determined by Hoechst 33342/PI staining, wherein the nuclei are stained to blue by Hoechst 33342, and the pyroptotic cells are stained to red by PI. Scale bars = 50 μm. B The quantitative analysis of PI positive cells. n = 6 per group. The quantification is representative of the percentage of pyroptotic cells in 5 fields ± SEM by random. C LDH release is measured by a cytotoxicity detection LDH kit. n = 6 per group. D Cell viability is measured by a CCK-8 kit, and the optical densities (ODs) are compared among the groups. n = 6 per group. E Representative immunoblots for NLRP3, caspase-1, cleaved-caspase-1, GSDMD, GSDMD-N in H9c2 cells. F The densitometric analysis of relative protein expressions. Vinculin is used as a loading control. n = 4 per group. G Cells with active caspase-1 in each group are stained with FLICA (FAM-YVAD-FMK) probe and detected using flow cytometer. H The percentages of caspase-1 positive cells are compared among the groups. n = 6 per group. Data represent the mean ± SEM. All experiments were repeated three times. ∗ P < 0.05, ∗ ∗ P < 0.01.
Index in PubMed under a CC BY license. PMID: 33436548
Click image to see more details
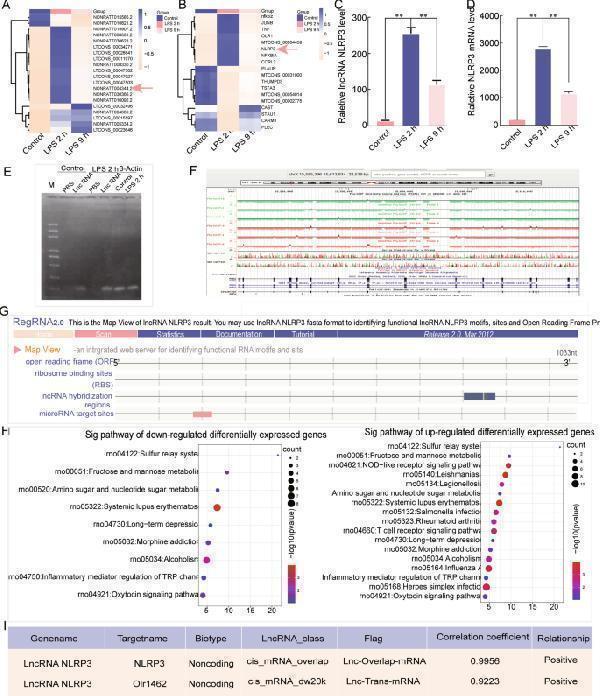
Dysregulated transcription of lncRNA NLRP3 and NLRP3 in LPS-treated NR8383 AM cells as determined by RNA-seq and bioinformatics analysis. A The heat map lists the top 20 differentially expressed lncRNAs and mRNAs in NR8383 AM after treatment with PBS, LPS for 2 h, and LPS for 9 h. A , B RNA-seq analysis shows the quantified gene expression of lncRNA NLRP3 and NLRP3 in AM cells in the negative control, LPS 2 h, and LPS 9 h groups. C , D Agarose gel electrophoresis analysis shows the quantified expression of lncRNA NLRP3 and NLRP3 in NR8383 cells. β-Actin served as the control. E The conservation of lncRNA NLRP3 was predicted and analysed by the UCSC Genome Browser. F The lncRNA NLRP3 potential protein-coding and binding sites were analysed with RNA 2.0 tools. G The results show that lncRNA NLRP3 has no protein-coding capability. H Gene Ontology and Kyoto Encyclopedia of Genes and Genomes pathway analysis were used to analyze differentially expressed genes. I The relationship between lncRNA NLRP3 and NLRP3, and the correlation coefficient is listed. * P < 0.05; ** P < 0.01; *** P < 0.001; NS, no statistically significant difference.
Index in PubMed under a CC BY license. PMID: 34599154
Click image to see more details
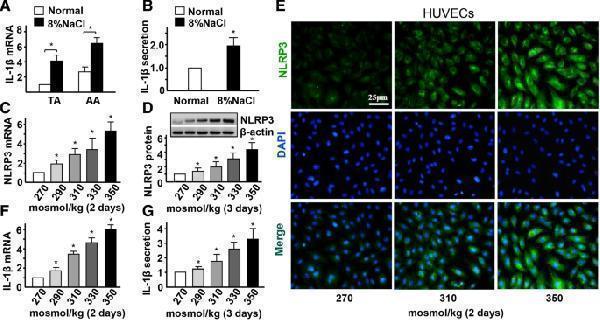
High-salt enhances NLRP3 inflammasome activation in ECs. a Quantification of mRNA levels of IL-1β in TA and AA of ApoE −/− mice fed with a normal or high-salt diet for 4 weeks. b Quantification of protein levels of IL-1β in the serum of ApoE −/− mice fed with a normal or high-salt diet for 4 weeks. c, d Quantification of mRNA levels and protein levels of NLRP3 by HUVECs in hypertonic medium (290, 310, 330 and 350 mosmol/kg) with isomolar 270 mosmol/kg as the control, evaluated by RT-qPCR and Western blotting with β-actin as the internal control. e Immunofluorescent staining of NLRP3 in HUVECs with iso- and hyper-osmotic media. f, g mRNA and protein expression of IL-1β in HUVECs exposed to iso- and hyper-osmotic media, evaluated by RT-qPCR and ELISA. All data were presented as mean ± SEM, N ≥ 3. * p < 0.05
Index in PubMed under a CC BY license. PMID: 31429763
Click image to see more details
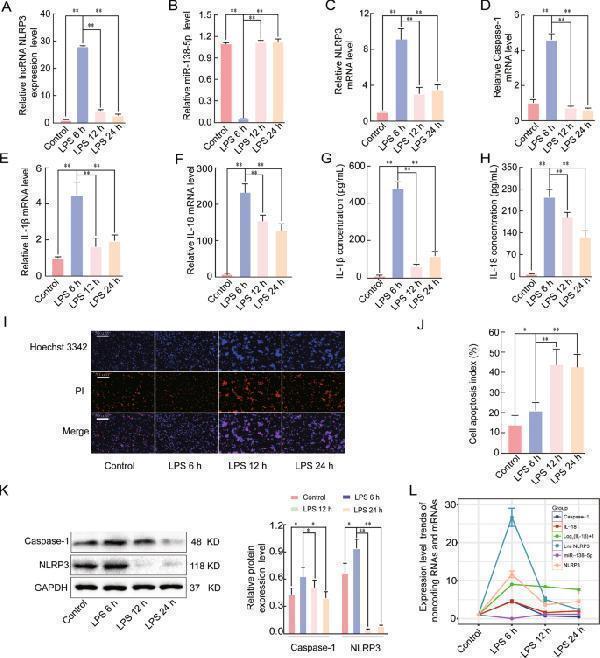
Effects of LPS on the lncRNA NLRP3, miR-138-5p, NLRP3, Caspase-1, IL-1β, and IL-18 expression levels in early ALI. A qRT-PCR assay was used to analyse the mRNA expression of A LncRNA NLRP3, B miR-138-5p, C NLRP3, D Caspase-1, E IL-1β, and F IL-18 in LPS-induced ALI. β-Actin was used as the reference gene. G , H ELISA analysis of the IL-1β and IL-18 levels in the culture supernatant. Cell apoptosis was determined by Hoechst 33342 and PI dual staining assays ( I ) and counted ( J ). The expression of NLRP3 and caspase-1 in the NR8383 AM cells from the four groups was analysed by western blotting ( K ). L Expression trends of lncRNA NLRP3, NLRP3, caspase-1, IL-1β, IL-18, and miR-138-5p in the negative control group and groups treated with LPS for 6, 12, and 24 h. The data are presented as mean ± SE ( n = 6). * P < 0.05; ** P < 0.01; *** P < 0.001; NS, no statistically significant difference.
Index in PubMed under a CC BY license. PMID: 34599154
Click image to see more details
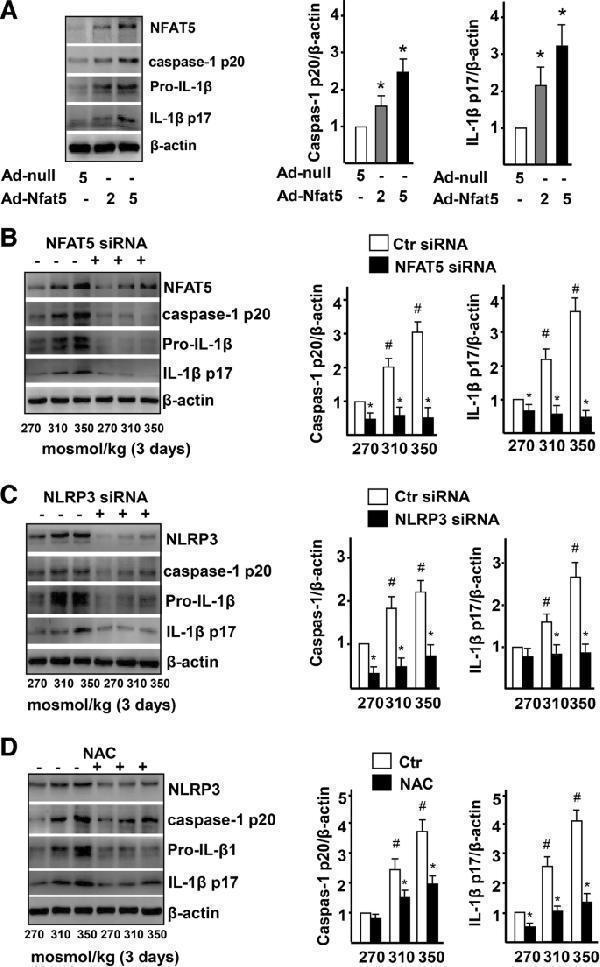
High-salt activates NRLP3 inflammasome in ECs via NFAT5. a Immunoblot of NFAT5, caspase-1 p20, pro-IL-1β, and IL-1β p17, and quantification of caspase-1 activity and mature IL-1β in ECs treated with Adenovirus-null (Ad-null, 5 MOI) and Adenovirus-NFAT5 (Ad-NFAT5, 2 MOI or 5 MOI). See Additional file : Figure S4 for caspase-1 activity. b Immunoblot images of NFAT5, caspase-1 p20, pro-IL-1β, and IL-1β p17, and quantification of active caspase-1 and mature IL-1β in ECs treated with high-salt and transfected with Ctr siRNA or NFAT5 siRNA. c Immunoblot images of NLRP3, caspase-1 p20, pro-IL-1β, and IL-1β p17, and quantification of active caspase-1 and mature IL-1β in ECs treated with high-salt and transfected with Ctr siRNA or NLRP3 siRNA. d Immunoblot images of NLRP3, caspase-1 p20, pro-IL-1β, and IL-1β p17, and quantification of active caspase-1 and mature IL-1β in ECs treated with high-salt, treated with NAC. All data were presented as mean ± SEM, N ≥ 3. * p < 0.05
Index in PubMed under a CC BY license. PMID: 31429763
Click image to see more details
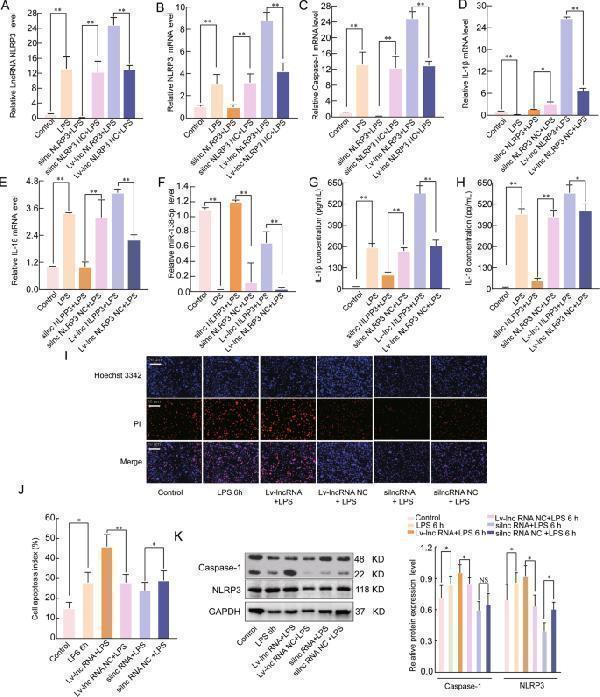
LncRNA NLRP3 regulates the inflammatory response during ALI through NLRP3 inflammasomes. A qRT-PCR assay was used to analyse the mRNA expression of A lncRNA NLRP3, B NLRP3, C Caspase-1, D IL-18, E IL-1β, and F miR-138-5p in LPS -induced ALI. β-Actin was used as the reference gene. G , H ELISA was used to analyse the IL-1β and IL-18 levels in the culture supernatants. I , J Cell apoptosis was determined by Hoechst 33342 and PI dual staining assays ( I ) and counted ( J ). K Western blotting was used to analyse the protein expression of NLRP3 and caspase-1 after lncRNA NLRP3 overexpression in the cytoplasm. The data are presented as mean ± SE ( n = 6). * P < 0.05; ** P < 0.01; *** P < 0.001; NS, no statistically significant difference.
Index in PubMed under a CC BY license. PMID: 34599154
Click image to see more details
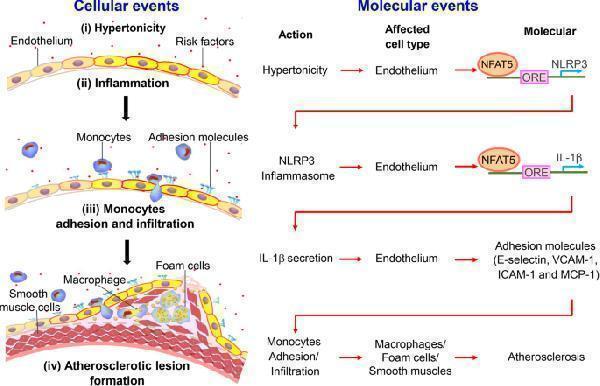
Schematic summarizes the mechanism that NLRP3 inflammasome activation in endothelium mediates hypertonic stress-induced atherosclerosis via NFAT5. Schematic illustration of the process. Stage i: Hypertonicity → NFAT5-dependent NLRP3 gene transcription → NLRP3 inflammasome activation. Stage ii: NLRP3 inflammasome activation → NFAT5-transcription-mediated pro-IL-1β → IL-1β secretion. Stage iii: IL-1β secretion → adhesive molecules → monocytes adhesion and infiltration. Stage iv: The activation of endothelial innate immunity promotes macrophage-driven foam cells and phenotype conversion of smooth muscle cells, contributing to the formation of atherosclerosis
Index in PubMed under a CC BY license. PMID: 31429763
Click image to see more details
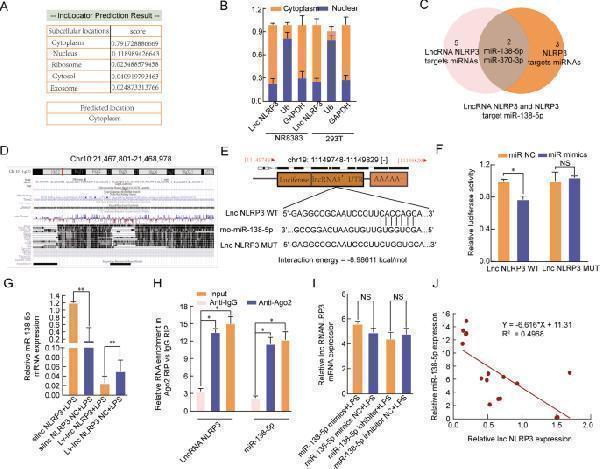
LncRNA NLRP3 functions as a sponge for miR-138-5p. A LncLocator was used to investigate the distribution of lncRNA NLRP3. B LncRNA NLRP3 was mainly located in the cytoplasm. C Venn diagram of miRDB predicting miR-138-5p and miR-370-3p sponged by lncRNA NLRP3 and NLRP3. D , E The predicted miR-138-5p-binding sites in the lncRNA NLRP3 3′-UTR. F miR-138-5p mimics notably reduced the luciferase activity of the lncRNA NLRP3-Wt group. G silncRNA NLRP3 significantly increased miR-138-5p expression; however, overexpression of lncRNA NLRP3 reduced miR-138-5p expression in LPS-treated NR8383 AM cells. H RIP assays revealed that Ago2-containing beads enriched the expression of miR-138-5p and NLRP3. I The miR-138-5p inhibitor and miR-138-5p mimics had no effects on lncRNA NLRP3 expression in LPS-treated NR8383 AM cells. The data are presented as mean ± SE ( n = 6). J LncRNA NLRP3 expression was negatively correlated with miR-138-5p expression in LPS-treated NR8383 AM cells. * P < 0.05; ** P < 0.01; *** p < 0.001; NS, no statistically significant difference.
Index in PubMed under a CC BY license. PMID: 34599154
Click image to see more details
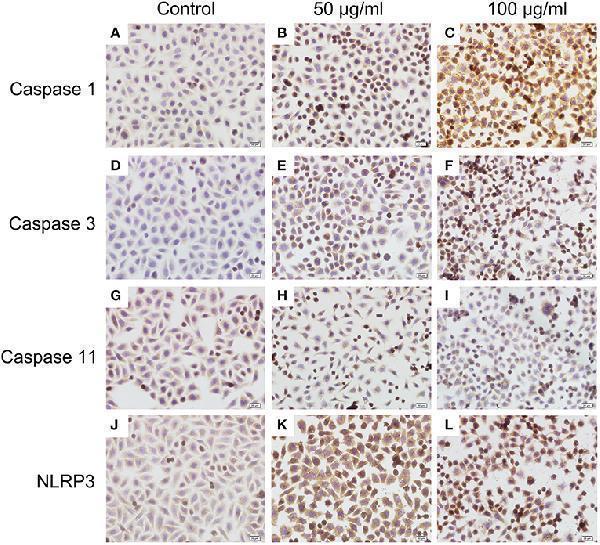
Immunocytochemistry analysis of histone induced activations of caspase 1, caspase 3, and NLRP3 (400×). BMECs were cultured briefly on cover glasses (pre-treated with poly-L –lysine, 0.1 mg/mL, Sigma-Aldrich) and incubated with histone (50 and 100 μg/mL) for 16 h. The samples were visualized with DAB, counterstained with hematoxylin and observed by inverted microscope. Three independent experiments were carried out by light microscope analyses.
Index in PubMed under a CC BY license. PMID: 31156617
Click image to see more details
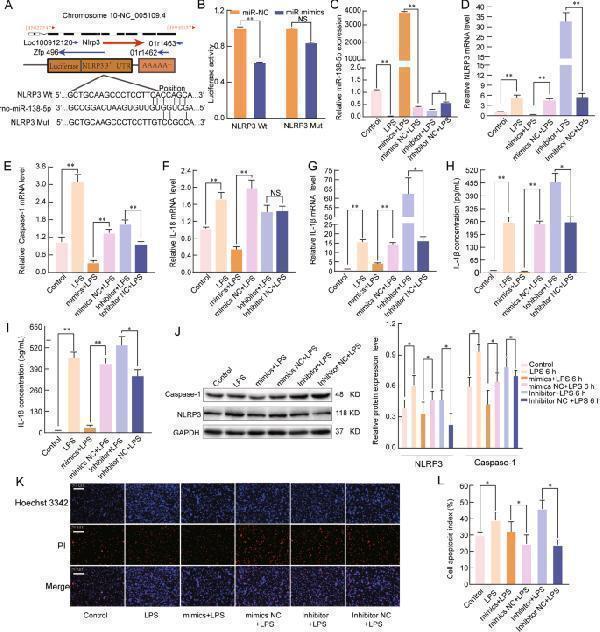
miR-138-5p regulates the inflammatory response by targeting NLRP3. A The predicted miR-138-5p-binding sites in the NLRP3 mRNA 3′-UTR. B A firefly luciferase reporter containing either wild-type or mutant NLRP3 was cotransfected into NR8383 AM cells with miR-138-5p mimics NC or miR-138-5p mimics. qRT-PCR assays were used to analyse the mRNA expression of C miR-138-5p, D NLRP3, E Caspase-1, F IL-18, and G IL-1β in the NR8383 AM cells ( n = 6). β-Actin was used as a reference gene. H , I ELISA was used to analyse the IL-1β and IL-18 levels in the culture supernatants. J Western blotting assay of the protein expression levels of NLRP3 and Caspase-1. K , L Cell apoptosis was determined by Hoechst 33342 and PI dual staining assays ( K ) and counted ( L ). The data are presented as mean ± SE ( n = 6). * P < 0.05; ** P < 0.01; *** P < 0.001; NS, no statistically significant difference.
Index in PubMed under a CC BY license. PMID: 34599154
Click image to see more details
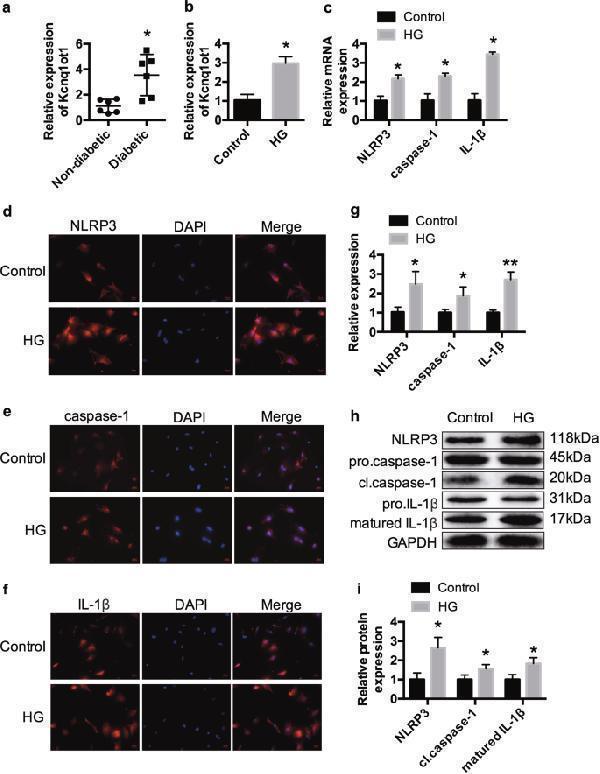
Kcnq1ot1 and pytoptosis are activated in HG-treated cardiac fibroblasts qRT-PCR was performed to measure the Kcnq1ot1 expression level in the serums of non-diabetic and diabetic patients a . * P < 0.05 compared with the non-diabetic group. n = 6 in each group. Cardiac fibroblasts of neonatal C57BL/6 mice were incubated with 5.5 mmol/L glucose (Control) or 30 mmol/L (high glucose, HG) for 24 h. The expression levels of Kcnq1ot1 were detected by qRT-PCR b . The expression levels of NLRP3, caspase-1 and IL-1β were determined by qRT-PCR c , immunofluorescence d – g and western blot h , i . * P < 0.05 compared with the control group. n = 3 in each group
Index in PubMed under a CC BY license. PMID: 30250027
Click image to see more details
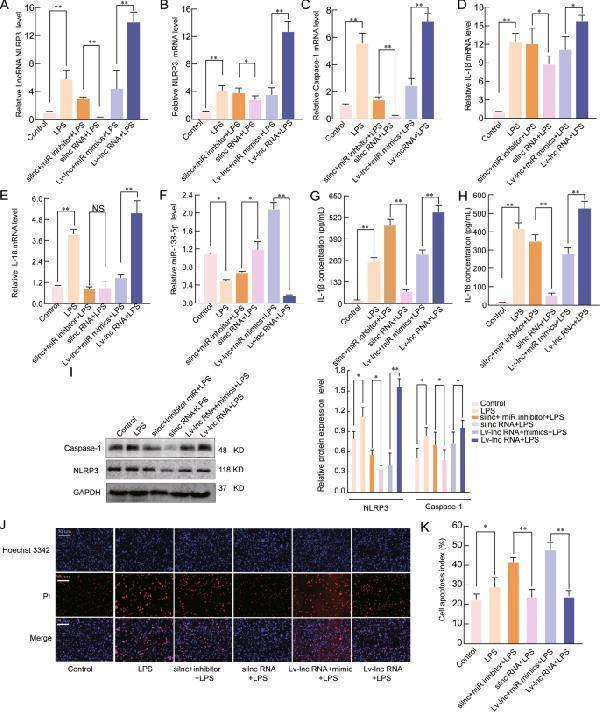
LncRNA NLRP3 regulates the inflammatory response through the lncRNA NLRP3/miR-138-5p/NLRP3 ceRNET in vitro. miR-138-5p suppression reversed the effects of silncRNA NLRP3 on the mRNA expression of A lncRNA NLRP3, B NLRP3, C Caspase-1, D IL-1β, E IL-18, and F miR-138-5p in NR8383 alveolar macrophage (AMs) cells. β-Actin was used as the reference gene. G , H ELISA analysis of the IL-1β and IL-18 levels in the culture supernatant. I Western blotting assay of the protein expression levels of NLRP3 and caspase-1. J , K Cell apoptosis was determined by Hoechst 33342 and PI dual staining assays ( J ) and counted ( K ). The data are presented as mean ± SE ( n = 6). * P < 0.05; ** P < 0.01; *** P < 0.001; NS, no statistically significant difference.
Index in PubMed under a CC BY license. PMID: 34599154
Click image to see more details
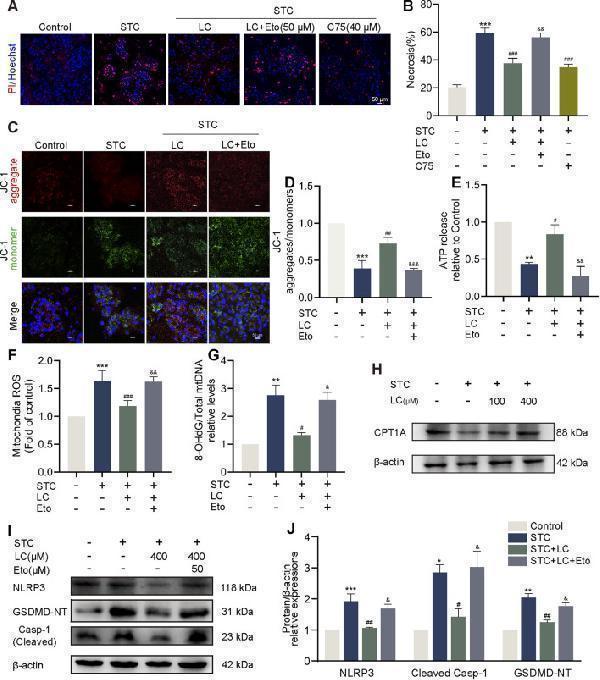
LC protects against STC-induced acinar cell pyroptosis via CPT1A. (A,B) Representative images and quantification of Hoechst 33342 (blue) and PI (red) staining of primary acinar cells received different treatments. Scale bar: 50 μm. (C,D) JC-1 fluorescent staining of primary acinar cells in different treatment groups. Scale bar: 50 μm. (E) The release of ATP levels in the supernatant of cultured primary acinar cells received different treatments. (F) Quantification of mtROS in primary acinar cells in different treatment groups. (G) The ratio of 8-OHdG to total mtDNA content in primary acinar cells received different treatments. (H) Protein levels of CPT1A and β-actin in the primary acinar cells received different treatments. (I,J) Protein levels of NLRP3, cleaved Caspase-1 and GSDMD-NT in the primary acinar cell received different treatments. β-actin served as the loading control (n = 3). Data are presented as means ± SEM, n = 3–5. * p < 0.05, ** p < 0.01, *** p < 0.001 vs. Control group. # p < 0.05, ## p < 0.01, ### p < 0.001, vs. STC group. & p < 0.05, && p < 0.01, &&& p < 0.001 vs. STC+LC treatment group. STC: sodium taurocholate; LC: L-carnitine; Eto: Etomoxir.
Index in PubMed under a CC BY license. PMID: 40718711
Click image to see more details
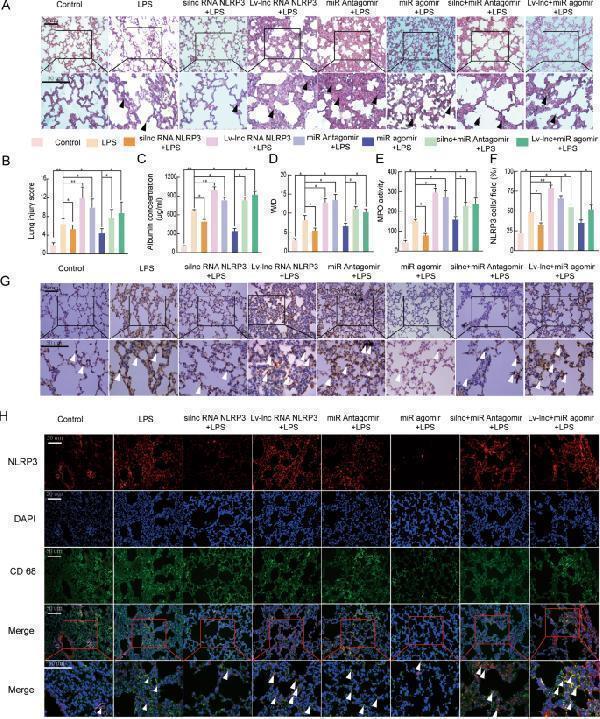
LncRNA NLRP3/miR-138-5p/NLRP3 functions via the ceRNET during the NLRP3-triggered inflammatory response in vivo. Rat lungs were injected with PBS in the control group and LPS-treated rats were further treated with si-r-lncRNA NLRP3, Lv-lncRNA NLRP3, agomiR-138-5p, antagomiR-138-5p, Lv-lncRNA NLRP3 + agomiR-138-5p, and si-r-lncRNA NLRP3 + antagomiR-138-5p. A Lung tissue samples were collected 6 h after establishing LPS-induced ALI to analyse the histopathological changes (×200, ×400). The black arrow indicates neutrophil infiltration, pulmonary oedema, alveolar wall thickening, and alveolar haemorrhage. B The lung injury score was determined via H&E staining, a representative histological analysis ( n = 6 animals per group). C ELISA was used to measure the BALF albumin content. D Detection of the lung W/D ratio in rats. E MPO activity in the lung tissues of rats. F , G Immunohistochemical detection of the NLRP3 contents in rat lung tissues (×200, ×400). H The inflammatory response in NR8383 AM cells was suppressed by si-r-lncRNA NLRP3 and miR-138-5p mimics alone or in combination, as shown by the decreased number of cells colabeled with CD68 (green) and NLRP3 (red). LncRNA NLRP3 overexpression, miR-138-5p inhibition, and NLRP3 augmented the inflammatory response in LPS-induced ALI with more NLRP3 and CD68 anchored in the plasma membrane of the AM cells. The data are presented as mean ± SE ( n = 6). * P < 0.05; ** P < 0.01; *** P < 0.001; NS, no statistically significant difference.
Index in PubMed under a CC BY license. PMID: 34599154
Specific Publications For Anti-CIAS1/NALP3/NLRP3 Antibody Picoband® (PA1665)
Loading publications
Recommended Resources
Here are featured tools and databases that you might find useful.
- Boster's Pathways Library
- Protein Databases
- Bioscience Research Protocol Resources
- Data Processing & Analysis Software
- Photo Editing Software
- Scientific Literature Resources
- Research Paper Management Tools
- Molecular Biology Software
- Primer Design Tools
- Bioinformatics Tools
- Phylogenetic Tree Analysis
Customer Reviews
Have you used Anti-CIAS1/NALP3/NLRP3 Antibody Picoband®?
Share your experimental results or join a short interview to earn up to $1,000 in product credits or other rewards.
0 Reviews For Anti-CIAS1/NALP3/NLRP3 Antibody Picoband®
Customer Q&As
Have a question?
Find answers in Q&As, reviews.
Can't find your answer?
Submit your question
18 Customer Q&As for Anti-CIAS1/NALP3/NLRP3 Antibody Picoband®
Question
Do you have a BSA free version of anti-CIAS1/NALP3/NLRP3 antibody PA1665 available?
Verified Customer
Verified customer
Asked: 2020-04-09
Answer
We appreciate your recent telephone inquiry. I can confirm that some lots of this anti-CIAS1/NALP3/NLRP3 antibody PA1665 are BSA free. For now, these lots are available and we can make a BSA free formula for you free of charge. It will take 3 extra days to prepare. If you require this antibody BSA free again in future, please do not hesitate to contact me and I will be pleased to check which lots we have in stock that are BSA free.
Boster Scientific Support
Answered: 2020-04-09
Question
Is this PA1665 anti-CIAS1/NALP3/NLRP3 antibody reactive to the isotypes of NLRP3?
Verified Customer
Verified customer
Asked: 2020-04-02
Answer
The immunogen of PA1665 anti-CIAS1/NALP3/NLRP3 antibody is A synthetic peptide corresponding to a sequence at the N-terminus of human CIAS1(12-31aa RYLEDLEDVDLKKFKMHLED), different from the related rat and mouse sequences by one amino acid. Could you tell me which isotype you are interested in so I can help see if the immunogen is part of this isotype?
Boster Scientific Support
Answered: 2020-04-02
Question
I appreciate helping with my inquiry over the phone. Here are the WB image, lot number and protocol we used for umbilical cord blood using anti-CIAS1/NALP3/NLRP3 antibody PA1665. Let me know if you need anything else.
Verified Customer
Verified customer
Asked: 2020-03-16
Answer
We appreciate the data. You have provided everything we needed. Our lab team are working to resolve your inquiry as quickly as possible, and we appreciate your patience and understanding! Please let me know if there is anything you need in the meantime.
Boster Scientific Support
Answered: 2020-03-16
Question
We have seen staining in rat blood. Are there any suggestions? Is anti-CIAS1/NALP3/NLRP3 antibody supposed to stain blood positively?
Verified Customer
Verified customer
Asked: 2020-02-06
Answer
From what I have seen in literature blood does express NLRP3. From what I have seen in Uniprot.org, NLRP3 is expressed in blood, brain, colon, umbilical cord blood, among other tissues. Regarding which tissues have NLRP3 expression, here are a few articles citing expression in various tissues:
Brain, Pubmed ID: 14702039, 16710414
Colon, Pubmed ID: 15489334
Umbilical cord blood, Pubmed ID: 11042152
Boster Scientific Support
Answered: 2020-02-06
Question
We purchased anti-CIAS1/NALP3/NLRP3 antibody for Flow Cytometry on brain a few years ago. I am using mouse, and We are going to use the antibody for ICC next. I would like examining brain as well as blood in our next experiment. Could you please give me some suggestion on which antibody would work the best for ICC?
Verified Customer
Verified customer
Asked: 2020-01-16
Answer
I have checked the website and datasheets of our anti-CIAS1/NALP3/NLRP3 antibody and I see that PA1665 has been validated on mouse in both Flow Cytometry and ICC. Thus PA1665 should work for your application. Our Boster satisfaction guarantee will cover this product for ICC in mouse even if the specific tissue type has not been validated. We do have a comprehensive range of products for ICC detection and you can check out our website bosterbio.com to find out more information about them.
Boster Scientific Support
Answered: 2020-01-16
Question
Does PA1665 anti-CIAS1/NALP3/NLRP3 antibody work on parafin embedded sections? If so, which fixation method do you recommend we use (PFA, paraformaldehyde, other)?
Verified Customer
Verified customer
Asked: 2019-11-28
Answer
As indicated on the product datasheet, PA1665 anti-CIAS1/NALP3/NLRP3 antibody as been validated on WB. It is best to use PFA for fixation because it has better tissue penetration ability. PFA needs to be prepared fresh before use. Long term stored PFA turns into formalin, as the PFA molecules congregate and become formalin.
Boster Scientific Support
Answered: 2019-11-28
Question
Does anti-CIAS1/NALP3/NLRP3 antibody PA1665 work for WB with umbilical cord blood?
Verified Customer
Verified customer
Asked: 2019-11-27
Answer
According to the expression profile of umbilical cord blood, NLRP3 is highly expressed in umbilical cord blood. So, it is likely that anti-CIAS1/NALP3/NLRP3 antibody PA1665 will work for WB with umbilical cord blood.
Boster Scientific Support
Answered: 2019-11-27
Question
I see that the anti-CIAS1/NALP3/NLRP3 antibody PA1665 works with WB, what is the protocol used to produce the result images on the product page?
L. Taylor
Verified customer
Asked: 2019-10-29
Answer
You can find protocols for WB on the "support/technical resources" section of our navigation menu. If you have any further questions, please send an email to support@bosterbio.com
Boster Scientific Support
Answered: 2019-10-29
Question
I am looking for to test anti-CIAS1/NALP3/NLRP3 antibody PA1665 on human umbilical cord blood for research purposes, then I may be interested in using anti-CIAS1/NALP3/NLRP3 antibody PA1665 for diagnostic purposes as well. Is the antibody suitable for diagnostic purposes?
Verified Customer
Verified customer
Asked: 2019-10-18
Answer
The products we sell, including anti-CIAS1/NALP3/NLRP3 antibody PA1665, are only intended for research use. They would not be suitable for use in diagnostic work. If you have the means to develop a product into diagnostic use, and are interested in collaborating with us and develop our product into an IVD product, please contact us for more discussions.
Boster Scientific Support
Answered: 2019-10-18
Question
I am interested in using your anti-CIAS1/NALP3/NLRP3 antibody for positive regulation of interleukin-13 production studies. Has this antibody been tested with western blotting on u87 cell lysate? We would like to see some validation images before ordering.
Verified Customer
Verified customer
Asked: 2019-10-08
Answer
I appreciate your inquiry. This PA1665 anti-CIAS1/NALP3/NLRP3 antibody is tested on a549 cell lysate, u87 cell lysate, cem cell lysate, u937 cells, rat kidney. It is guaranteed to work for Flow Cytometry, IF, IHC-P, ICC, WB in human, mouse, rat. Our Boster guarantee will cover your intended experiment even if the sample type has not been be directly tested.
Boster Scientific Support
Answered: 2019-10-08
Question
My question regarding product PA1665, anti-CIAS1/NALP3/NLRP3 antibody. I was wondering if it would be possible to conjugate this antibody with biotin. I would need it to be without BSA or sodium azide. I am planning on using a buffer exchange of sodium azide with PBS only. Would there be problems for me to conjugate the antibody and store it in -20 degrees in small aliquots?
Verified Customer
Verified customer
Asked: 2019-09-23
Answer
We do not advise storing this antibody with PBS buffer only in -20 degrees. If you want to store it in -20 degrees it is best to add some cryoprotectant like glycerol. If you want carrier free PA1665 anti-CIAS1/NALP3/NLRP3 antibody, we can provide it to you in a special formula with trehalose and/or glycerol. These molecules will not interfere with conjugation chemistry and provide a good level of protection for the antibody from degradation. Please be sure to specify this in your purchase order.
Boster Scientific Support
Answered: 2019-09-23
Question
I was wanting to use your anti-CIAS1/NALP3/NLRP3 antibody for WB for human umbilical cord blood on frozen tissues, but I want to know if it has been tested for this particular application. Has this antibody been tested and is this antibody a good choice for human umbilical cord blood identification?
Verified Customer
Verified customer
Asked: 2019-07-16
Answer
It shows on the product datasheet, PA1665 anti-CIAS1/NALP3/NLRP3 antibody has been tested for Flow Cytometry, IF, IHC-P, ICC, WB on human, mouse, rat tissues. We have an innovator award program that if you test this antibody and show it works in human umbilical cord blood in IHC-frozen, you can get your next antibody for free.
Boster Scientific Support
Answered: 2019-07-16
Question
Does anti-CIAS1/NALP3/NLRP3 antibody PA1665 work on primate IHC-P with colon?
Verified Customer
Verified customer
Asked: 2019-05-16
Answer
Our lab technicians have not tested anti-CIAS1/NALP3/NLRP3 antibody PA1665 on primate. You can run a BLAST between primate and the immunogen sequence of anti-CIAS1/NALP3/NLRP3 antibody PA1665 to see if they may cross-react. If the sequence homology is close, then you can perform a pilot test. Keep in mind that since we have not validated primate samples, this use of the antibody is not covered by our guarantee. However we have an innovator award program that if you test this antibody and show it works in primate colon in IHC-P, you can get your next antibody for free.
Boster Scientific Support
Answered: 2019-05-16
Question
We were happy with the WB result of your anti-CIAS1/NALP3/NLRP3 antibody. However we have been able to see positive staining in blood golgi apparatus membrane. note=(microbial using this antibody. Is that expected? Could you tell me where is NLRP3 supposed to be expressed?
R. Banerjee
Verified customer
Asked: 2019-03-19
Answer
From literature, blood does express NLRP3. Generally NLRP3 expresses in cytoplasm, cytosol, golgi apparatus membrane. note=(microbial. Regarding which tissues have NLRP3 expression, here are a few articles citing expression in various tissues:
Brain, Pubmed ID: 14702039, 16710414
Colon, Pubmed ID: 15489334
Umbilical cord blood, Pubmed ID: 11042152
Boster Scientific Support
Answered: 2019-03-19
Question
Hello, I currently trying to establish NLRP3 staining (IF) in cryo sections from colon using at the moment an polyclonal NLRP3 antibody from abcam (ab4207, goat). My current antibody shows unspecific staining of colon mucus and epithelia. Did you find a way to counteract this? Is it possible with your aNLRP3-Antibody, to stain NLRP3 specific in cryo colon sections?
Verified Customer
Verified customer
Asked: 2018-10-08
Answer
We are uncertain if PA1665 is specific in cryo colon sections. Please suggest the customer to purchase a sample size to test.
Boster Scientific Support
Answered: 2018-10-08
Question
Is a blocking peptide available for product anti-CIAS1/NALP3/NLRP3 antibody (PA1665)?
M. Edwards
Verified customer
Asked: 2016-08-09
Answer
We do provide the blocking peptide for product anti-CIAS1/NALP3/NLRP3 antibody (PA1665). If you would like to place an order for it please contact support@bosterbio.com and make a special request.
Boster Scientific Support
Answered: 2016-08-09
Question
We are currently using anti-CIAS1/NALP3/NLRP3 antibody PA1665 for human tissue, and we are content with the Flow Cytometry results. The species of reactivity given in the datasheet says human, mouse, rat. Is it likely that the antibody can work on feline tissues as well?
J. Roberts
Verified customer
Asked: 2015-09-18
Answer
The anti-CIAS1/NALP3/NLRP3 antibody (PA1665) has not been validated for cross reactivity specifically with feline tissues, but there is a good chance of cross reactivity. We have an innovator award program that if you test this antibody and show it works in feline you can get your next antibody for free. Please contact me if I can help you with anything.
Boster Scientific Support
Answered: 2015-09-18
Question
See below the WB image, lot number and protocol we used for umbilical cord blood using anti-CIAS1/NALP3/NLRP3 antibody PA1665. Please let me know if you require anything else.
E. Evans
Verified customer
Asked: 2014-01-29
Answer
Thank you very much for the data. Our lab team are working to resolve this as quickly as possible, and we appreciate your patience and understanding! You have provided everything we needed. Please let me know if there is anything you need in the meantime.
Boster Scientific Support
Answered: 2014-01-29





