Recombinant Technology Concept Overview
A recombinant protein is a protein produced artificially by an expression host organism,
aka expression system, that is transfected with a recombinant gene (target gene) isolated from
another organism. The purpose is to produce the target gene encoded protein in large quantities for
medical, research and academic uses.
History of recombinant protein technology
Early 1950s discovery of DNA plasmids directly led to the field of recombinant DNA
(rDNA). Peter Lobban from Stanford University Medical School first proposed the recombinant DNA
idea in the early 1970s. The first successful production of recombinant proteins is documented
in 1972 and 1973 from Stanford and UCSF. Several people were awarded the Nobel Prize for
contribution to the recombinant technology.
The pharmaceutical industry has been utilizing recombinant technology for producing
protein drugs since the early 1980s. One of the most impactful achievements of recombinant
protein technology is the production of recombinant protein insulin. Before recombinant protein
insulin is made, diabetes treatment largely depended on insulin from animal sources. The
recombinant insulin significantly lowered treatment cost and enabled a consistent and sufficient
worldwide supply for diabetes treatment. The discovery of restriction enzymes and ligases
catapulted the field to the ability to insert foreign genetic material and proliferate in a
bacterial system.
What does a restriction enzyme do?
Restriction enzymes, more precisely restriction endonucleases, cleave DNA with high
specificity in pre-defined cleaving patterns, making them the backbone of recombinant technology.
Under the guide of their enzymatic properties, restriction endonucleases are able to cleave plasmids
and insert the foreign gene of interest. Further characterization analysis is realized with the
enhanced expression.
Individual restriction enzymes recognize specific sequences, referred to as restriction
sequences. Restriction sequences are palindromic sequences, usually 4 to 8 base pairs long.
Palindromic sequences read the same in both the upstream and downstream direction. When restriction
enzymes find these restriction sequences, they cleave the covalent bond of either the pyrimidine (T
and C) or purine (A and G), generating a 5’ phosphate and a 3’ hydroxyl group at the cleavage
point.
This cleavage leaves behind blunt ends with no overlap,
therefore they can no longer base pair with other nucleotides. Other types of restriction
endonucleases
leave sticky ends. These sticky ends are single strand overhangs that extend
beyond the cleavage site that may further bond with other nucleotides. Sticky
ends come in handy when the same restriction enzyme is used to cleave both the
fragment of interest and the vector. This cleavage will leave identical
overhangs that a ligase will bind (ligation), resulting in your recombinant
product (rDNA, protein, etc.).
Expression systems
There are several common expression systems for producing recombinant proteins, such as
E. Coli, Yeast, Insect Cell lines such as Sf21, mammalian cells such as CHO, HEK, and human cell
lines. These expression systems act as the factory for recombinant protein production, where the
cellular mechanisms involved in protein production use the recombinant gene in the vector as a
blueprint to mass produce the target protein.
E. Coli is the first organism, and still often the first choice, to be used as an
expression system. It is easy to use, however cannot produce protein with post-translational
modification (PTM). E. Coli lacks the cellular mechanisms of eukaryotic cells which are necessary
for PTM of proteins. Thus, for producing simple proteins and for applications that do not require
proteins to have PTM, E. Coli is a sufficient and convenient expression system. For more complex
applications, more advanced expression systems are required.
Expression Vectors
A vector is a tool for manipulating DNA. It is the transport vehicle for the gene of the
protein of interest. There are many types of vectors; some common vectors are plasmids and reverse
transcription viruses. The DNA that encodes the protein is first synthesized in vitro, or cloned
from native host DNA templates, then inserted into the vector DNA. The host organism is processed to
take up vectors. Expression vectors, once inside the corresponding host cells, will be both
transcribed and translated into proteins. They are engineered specifically to enhance the expression
of target proteins.
Vectors to Recombinant Proteins
Vectors are known for producing high numbers of copies, as it is exceedingly efficient
in self-replication. It is utilized as a carrier of foreign passenger DNA, to be introduced at the
restriction enzyme’s cleavage site. Engineered vectors may have many recognition sites; >20
recognition sites are referred to collectively as Multiple Cloning Sites (MCS). There are many
commercially available vector types for cloning and expression studies. Vectors are chosen for their
ability to overexpress and produce high numbers of the proteins of interest from the carrier
plasmid.
Just like the decision of which vector to use, the importance of the choice of
appropriate biological system, often bacterial, cannot be understated. These systems are referred to
as “protein factories”, as the necessity of a robust system is mandatory for overexpression and
protein production.
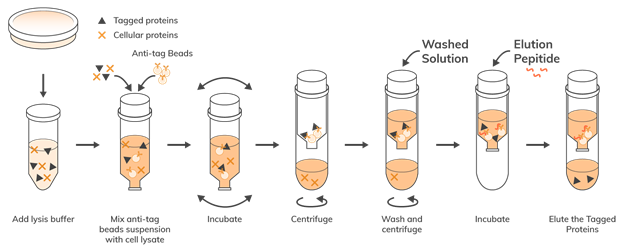
Once cloning of the recombinant protein of interest is completed, the protein must be
purified. Structural and functional studies, along with high-throughput and large-scale production,
require proteins to be >95% pure.