Product Info Summary
| SKU: | M03855 |
|---|---|
| Size: | 100 μl/vial |
| Reactive Species: | Human |
| Host: | Rabbit |
| Application: | Flow Cytometry, IF, ICC, WB |
Customers Who Bought This Also Bought
Product info
Product Name
Anti-ATAD2 Rabbit Monoclonal Antibody
SKU/Catalog Number
M03855
Size
100 μl/vial
Form
Liquid
Description
Boster Bio Anti-ATAD2 Rabbit Monoclonal Antibody catalog # M03855. Tested in WB, ICC/IF, Flow Cytometry applications. This antibody reacts with Human.
Storage & Handling
Store at -20°C for one year. For short term storage and frequent use, store at 4°C for up to one month. Avoid repeated freeze-thaw cycles.
Cite This Product
Anti-ATAD2 Rabbit Monoclonal Antibody (Boster Biological Technology, Pleasanton CA, USA, Catalog # M03855)
Host
Rabbit
Contents
Rabbit IgG in stabilizing components, phosphate buffered saline, pH 7.4, 150mM NaCl, 0.02% sodium azide and 50% glycerol.
*This antibody is supplied in a stabilized formulation.
Compatibility with conjugation reactions depends on the chemistry of the conjugation method used.
For conjugation methods that are not compatible with the stabilizing components present in this formulation, a carrier-free antibody format is required.
Clonality
Monoclonal
Clone Number
21A07
Isotype
IgG
Immunogen
A synthesized peptide derived from human ATAD2
Reactive Species
M03855 is reactive to ATAD2 in Human
Observed Molecular Weight
180 kDa
Calculated molecular weight
158.6 kDa
Antibody Validation
Boster validates all antibodies on WB, IHC, ICC, Immunofluorescence, and ELISA with known positive control and negative samples to ensure specificity and high affinity, including thorough antibody incubations.
Application & Images
Applications
M03855 is guaranteed for Flow Cytometry, IF, ICC, WB Boster Guarantee
Recommend Dilution
WB 1:500-2000
ICC/IF 1:50-200
FC 1:50
Validation Images & Assay Conditions

Click image to see more details
All lanes use the Antibody at 1:2K dilution for 1 hour at room temperature.

Click image to see more details
Western blot analysis of ATAD2 expression in MCF-7 cell lysate.
Click image to see more details
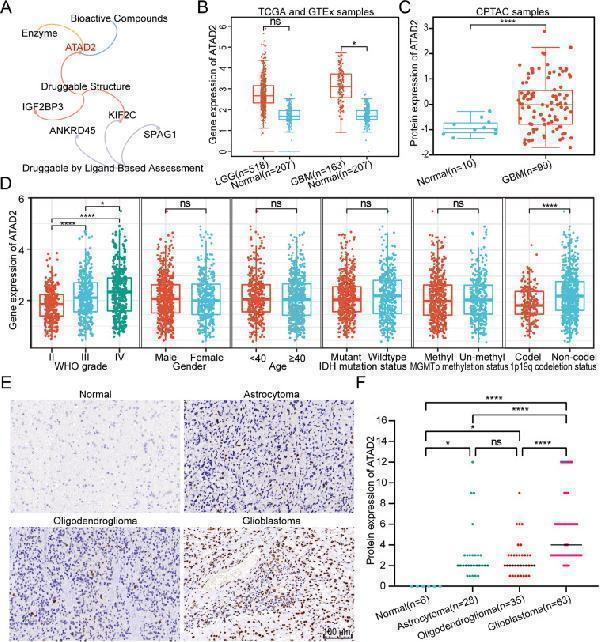
The expression of the druggable target ATAD2 in glioma. (A) Visualization of DepMap database-based druggability analysis of CTARS-related proteins using a network Venn diagram. (B) Analysis using the GEPIA database analysis revealed significant upregulation of ATAD2 mRNA expression in glioblastoma (GBM). (C) The UALCAN database analysis showed significant up-regulation of ATAD2 protein in GBM. (D) The expression of ATAD2 mRNA across different clinicopathological characteristics was analyzed in the CGGA cohort. Statistical analyses utilized the Kruskal–Wallis H test followed by post hoc Dunn's test for WHO grade comparisons and the Wilcoxon rank-sum test for other parameters. (E) Representative cases showing immunohistochemical staining across various clinicopathological tissue types of gliomas. Scale bar: 100 μm. (F) Statistical analysis of the IHC scores based on the staining intensity and the percentage of positive cells. Statistical analysis was performed with the Kruskal–Wallis H test and by post hoc Dunn's test. Significance levels are indicated as follows: ns, not significant; ∗, P < 0.05; ∗∗∗∗, P < 0.0001.
Index in Genes & Diseases under a CC BY license. DOI: 10.1016/j.gendis.2025.101810
Click image to see more details
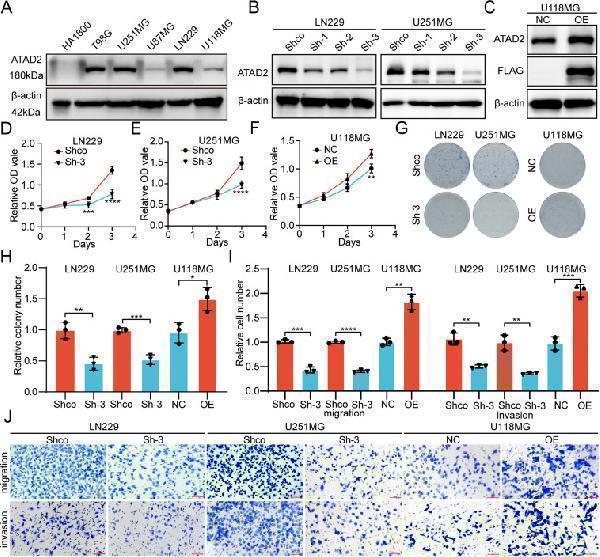
The impact of ATAD2 on the malignant phenotype of glioma cells. (A) The Western blot analysis shows the differential expression of ATAD2 in a normal astrocyte cell line (HA1800) and various glioma cell lines. (B) The Western blot analysis demonstrates the efficacy of three ATAD2 shRNAs in reducing ATAD2 expression in the LN229 and U251MG cell lines. (C) The Western blot analysis revealed the expression levels of ATAD2 and the FLAG-tagged protein following ATAD2 overexpression in U118MG cells. (D–F) The CCK-8 assay shows changes in cell proliferation after knockdown of ATAD2 in LN229 and U251MG cells, and its overexpression in U118MG cells. (G, H) Colony formation assay and statistical analysis. (I, J) Migration and invasion assays and their statistical analysis. The data are presented as means ± SD. For panels (D–F), two-way ANOVA was performed, while unpaired two-tailed Student's t-tests were used for panels (H, I). Statistical significance is denoted as follows: ∗, P < 0.05; ∗∗, P < 0.01; ∗∗∗, P < 0.001; ∗∗∗∗, P < 0.0001.
Index in Genes & Diseases under a CC BY license. DOI: 10.1016/j.gendis.2025.101810
Click image to see more details
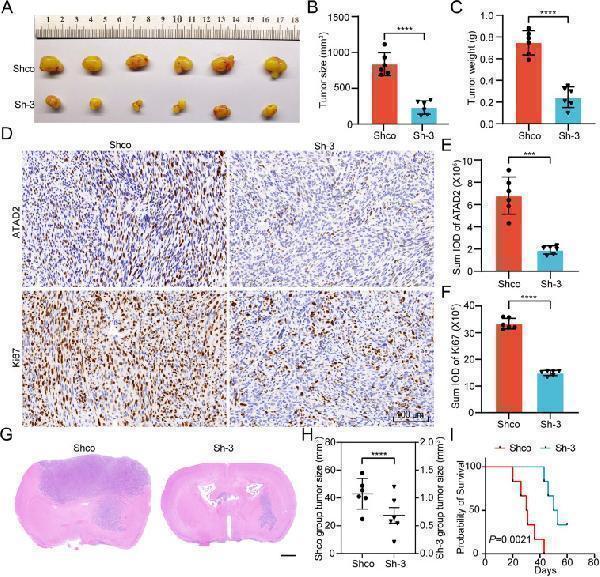
The impact of ATAD2 knockdown on Glioma tumorigenesis in vivo. (A) Gross morphological observations were conducted on subcutaneous tumors induced by LN229 cells in the Shco and Sh-3 groups of nude mice. (B) Statistical analysis of the subcutaneous tumor size measurements. (C) Statistical analysis of the subcutaneous xenograft tumor weights. (D) Representative images of IHC staining for ATAD2 and Ki67. Scale bar:100 μm. (E, F) The Image-Pro Plus software (version 6) was used to compute the sum integrated optical density of ATAD2 and Ki-67 IHC staining, followed by statistical analysis. (G) Representative HE staining images of intracranial orthotopic xenografts formed by LN229 cells in the Shco and Sh-3 groups. Scale bar: 1 mm. (H) Statistical analysis of intracranial orthotopic tumor size measurements. (I) Kaplan–Meier survival curve of nude mice with orthotopic intracranial tumors (Log-rank test). The data are presented as means ± SD, two-tailed paired Student's t-test was used to compare the results in (B), (C), (E), and (F); and two-tailed unpaired Student's t-test was used to compare the results in (H). Each group included n = 6 mice. Statistical significance is denoted as follows: ∗∗∗, P < 0.001; ∗∗∗∗, P < 0.0001.
Index in Genes & Diseases under a CC BY license. DOI: 10.1016/j.gendis.2025.101810
Click image to see more details
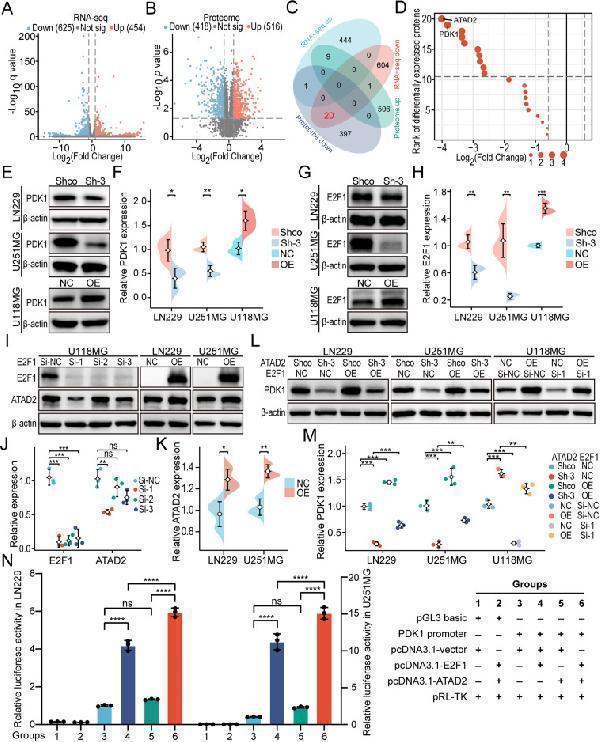
ATAD2-E2F1 positive feedback loop regulates the expression of PDK1. (A) Volcano plot of differentially expressed genes from RNA-seq analysis of ATAD2 knockdown and control conditions in LN229 cells. (B) Volcano plot of differentially expressed proteins from proteome analysis of ATAD2 knockdown and control conditions in LN229 cells. (C) The Venn diagram illustrates the overlap between the up-regulated and down-regulated mRNAs and proteins. (D) The fold change ranking plot illustrates 20 commonly down-regulated proteins. (E, F) Western blot analysis confirmed that ATAD2 enhances the expression of PDK1. (G, H) Western blot analysis confirmed that E2F1 up-regulates the expression of ATAD2. (I–K) Western blot analysis validated that E2F1 enhances the expression of ATAD2. (L, M) Western blot analysis showed that ATAD2 cooperates with E2F1 to up-regulate the expression of PDK1. (N) The dual luciferase reporter gene assays revealed that ATAD2 and E2F1 synergistically enhance the promoter activity of PDK1. The data are presented as means ± SD. Statistical comparisons were performed using unpaired two-tailed Student's t-test for figures (F), (H), and (K), and one-way ANOVA followed by Tukey's post hoc test for figures (J), (M), and (N). Statistical significance is denoted as follows: ns, not significant; ∗, P < 0.05; ∗∗, P < 0.01; ∗∗∗, P < 0.001; ∗∗∗∗, P < 0.0001.
Index in Genes & Diseases under a CC BY license. DOI: 10.1016/j.gendis.2025.101810
Click image to see more details
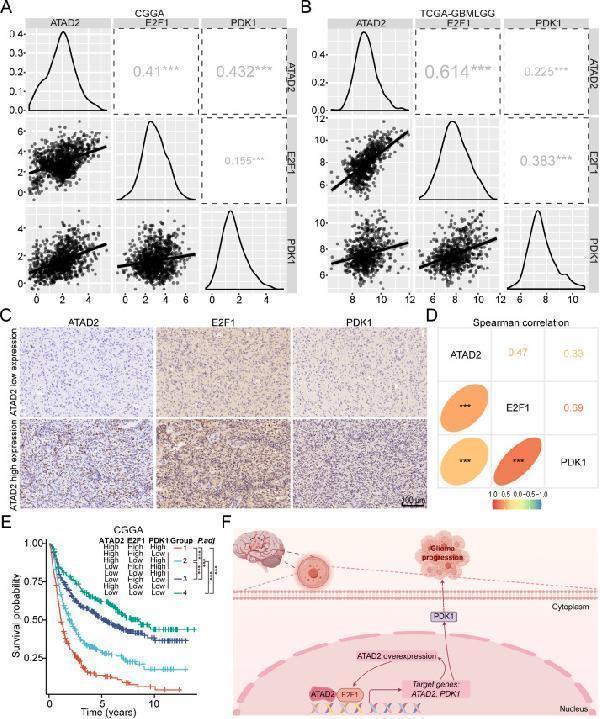
The clinical significance of the ATAD2-E2F1-PDK1 axis in glioma. (A, B) Analysis using the GlioVis database showed a positive correlation between the expressions of ATAD2, E2F1, and PDK1, as determined by the Spearman correlation test. (C) Representative cases showed the expression of E2F1 and PDK1 in glioma clinical specimens with low and high expression of ATAD2. Scale bar: 100 μm. (D) Spearman's correlation analysis of ATAD2, E2F1, and PDK1 expression levels in glioma clinical specimens. (E) Kaplan–Meier analysis and log-rank test of survival rates across distinct co-expression groups of ATAD2, E2F1, and PDK1 in the CGGA cohort. P values were adjusted for multiple group comparisons using the Bonferroni method. (F) Graphical diagram illustrating that ATAD2 promotes glioma progression and synergizes with E2F1 to increase PDK1 expression. The figure was created using Figdraw (www.figdraw.com). Statistical significance is denoted as follows: ∗∗∗, P < 0.001.
Index in Genes & Diseases under a CC BY license. DOI: 10.1016/j.gendis.2025.101810

Click image to see more details
Immunofluorescent analysis using the Antibody at 1:50 dilution.

Click image to see more details
Immunofluorescent analysis using the Antibody at 1:150 dilution.
Specific Publications For Anti-ATAD2 Rabbit Monoclonal Antibody (M03855)
Loading publications
Recommended Resources
Here are featured tools and databases that you might find useful.
- Boster's Pathways Library
- Protein Databases
- Bioscience Research Protocol Resources
- Data Processing & Analysis Software
- Photo Editing Software
- Scientific Literature Resources
- Research Paper Management Tools
- Molecular Biology Software
- Primer Design Tools
- Bioinformatics Tools
- Phylogenetic Tree Analysis
Customer Reviews
Have you used Anti-ATAD2 Rabbit Monoclonal Antibody?
Share your experimental results or join a short interview to earn up to $1,000 in product credits or other rewards.
0 Reviews For Anti-ATAD2 Rabbit Monoclonal Antibody
Customer Q&As
Have a question?
Find answers in Q&As, reviews.
Can't find your answer?
Submit your question





