Product Info Summary
| SKU: | PA1806 |
|---|---|
| Size: | 100 μg/vial |
| Reactive Species: | Human, Mouse, Rat |
| Host: | Rabbit |
| Application: | WB |
Customers Who Bought This Also Bought
Product info
Product Name
Anti-Aurora B/AURKB Antibody Picoband®
SKU/Catalog Number
PA1806
BA3799-2 is an alternative SKU for this antibody, used in previous lots.
Size
100 μg/vial
Form
Lyophilized
Description
Boster Bio Anti-Aurora B/AURKB Antibody catalog # PA1806. Tested in WB applications. This antibody reacts with Human, Mouse, Rat. The brand Picoband indicates this is a premium antibody that guarantees superior quality, high affinity, and strong signals with minimal background in Western blot applications. Only our best-performing antibodies are designated as Picoband, ensuring unmatched performance.
Storage & Handling
Store at -20˚C for one year from date of receipt. After reconstitution, at 4˚C for one month. It can also be aliquotted and stored frozen at -20˚C for six months. Avoid repeated freeze-thaw cycles.
Cite This Product
Anti-Aurora B/AURKB Antibody Picoband® (Boster Biological Technology, Pleasanton CA, USA, Catalog # PA1806)
Host
Rabbit
Contents
Each vial contains 4 mg Trehalose, 0.9 mg NaCl and 0.2 mg Na2HPO4.
Clonality
Polyclonal
Isotype
Rabbit IgG
Immunogen
A synthetic peptide corresponding to a sequence at the N-terminus of mouse Aurora B, different from the related rat sequence by two anino acids.
Cross-reactivity
No cross-reactivity with other proteins
Reactive Species
PA1806 is reactive to Aurkb in Human, Mouse, Rat
Observed Molecular Weight
39 kDa
Calculated molecular weight
39.4 kDa
Background of Aurkb
AURKB (aurora kinase B) also called Aik2, AIM-1, ARK2, AurB, STK12, IPL1, PPP1R48 or STK5, localizes to microtubules near kinetochores, specifically to the specialized microtubules called K-fibers. Cell cycle and Northern blot analyses showed that STK12 is expressed in the S phase and persistently thereafter. Western blot analysis indicated that STK12 is localized in the midbodies during anaphase. Northern blot analysis detected strong expression of a 1.5-kb STK12 transcript in thymus, with weaker expression in small intestine, testis, colon, spleen, and brain. The AURKB gene is mapped on 17p13.1. Examination of the role of both kinases in the phosphorylation of CENPA revealed that the reaction is mediated sequentially by AURKA and AURKB in early mitosis. EB1 overexpression enhanced AURKB kinase activity, and knockdown of EB1 with small interfering RNA impaired AURKB activity. EB1 protected AURKB from dephosphorylation/inactivation by protein phosphatase-2A by blocking binding of PP2A to AURKB.
Antibody Validation
Boster validates all antibodies on WB, IHC, ICC, Immunofluorescence, and ELISA with known positive control and negative samples to ensure specificity and high affinity, including thorough antibody incubations.
Application & Images
Applications
PA1806 is guaranteed for WB Boster Guarantee
Recommend Dilution
| Application | Dilution | Species |
|---|---|---|
| Western blot | 0.1-0.5μg/ml | Human, Mouse, Rat |
Tested application
Suggested blocking solution with 5% non-fat milk or BSA; (*)Recommended protein loading: 20-40 µg per lane
Validation Images & Assay Conditions
Click image to see more details
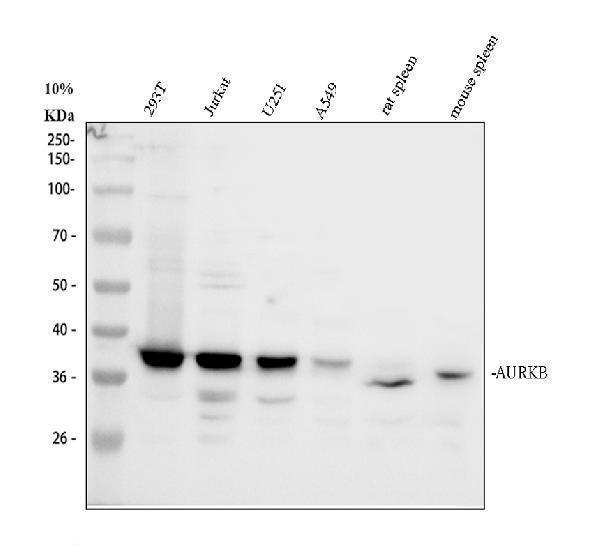
Western blot analysis of AURKB using anti-AURKB antibody (PA1806).
Electrophoresis was performed on a 10% SDS-PAGE gel at 80V (Stacking gel) / 120V (Resolving gel) for 2 hours. The sample well of each lane was loaded with 30 ug of sample under reducing conditions.
Lane 1: human 293T whole cell lysates,
Lane 2: human Jurkat whole cell lysates,
Lane 3: human U251 whole cell lysates,
Lane 4: human A549 whole cell lysates,
Lane 5: rat spleen tissue lysates,
Lane 6: mouse spleen tissue lysates.
After electrophoresis, proteins were transferred to a nitrocellulose membrane at 150 mA for 50-90 minutes. Blocked the membrane with 5% non-fat milk/TBS for 1.5 hour at RT. The membrane was incubated with rabbit anti-AURKB antigen affinity purified polyclonal antibody (PA1806) at 0.5 μg/mL overnight at 4°C, then washed with TBS-0.1%Tween 3 times with 5 minutes each and probed with a goat anti-rabbit IgG-HRP secondary antibody (Catalog # BA1054) at a dilution of 1:5000 for 1.5 hour at RT. The signal is developed using an ECL Plus Western Blotting Substrate (Catalog # AR1196-200) with Tanon 5200 system. A specific band was detected for AURKB at approximately 39 kDa. The expected band size for AURKB is at 39 kDa.
Specific Publications For Anti-Aurora B/AURKB Antibody Picoband® (PA1806)
Loading publications
Recommended Resources
Here are featured tools and databases that you might find useful.
- Boster's Pathways Library
- Protein Databases
- Bioscience Research Protocol Resources
- Data Processing & Analysis Software
- Photo Editing Software
- Scientific Literature Resources
- Research Paper Management Tools
- Molecular Biology Software
- Primer Design Tools
- Bioinformatics Tools
- Phylogenetic Tree Analysis
Customer Reviews
Have you used Anti-Aurora B/AURKB Antibody Picoband®?
Share your experimental results or join a short interview to earn up to $1,000 in product credits or other rewards.
0 Reviews For Anti-Aurora B/AURKB Antibody Picoband®
Customer Q&As
Have a question?
Find answers in Q&As, reviews.
Can't find your answer?
Submit your question
4 Customer Q&As for Anti-Aurora B/AURKB Antibody Picoband®
Question
We were well pleased with the WB result of your anti-Aurora B/AURKB antibody. However we have been able to see positive staining in lung nucleus. chromosome. chromosome using this antibody. Is that expected? Could you tell me where is AURKB supposed to be expressed?
Verified Customer
Verified customer
Asked: 2020-03-04
Answer
According to literature, lung does express AURKB. Generally AURKB expresses in nucleus. chromosome. chromosome, centromere. Regarding which tissues have AURKB expression, here are a few articles citing expression in various tissues:
Cervix carcinoma, Pubmed ID: 18669648, 18691976, 20068231
Cervix carcinoma, and Erythroleukemia, Pubmed ID: 23186163
Liver, and Spleen, Pubmed ID: 9858806
Lung, Lymph, and Muscle, Pubmed ID: 15489334
Boster Scientific Support
Answered: 2020-03-04
Question
We want using your anti-Aurora B/AURKB antibody for histone h3-s28 phosphorylation studies. Has this antibody been tested with western blotting on tissue lysate? We would like to see some validation images before ordering.
Verified Customer
Verified customer
Asked: 2019-10-25
Answer
Thanks for your inquiry. This PA1806 anti-Aurora B/AURKB antibody is tested on rat liver tissue, tissue lysate, 22rv cell lysate, hela cell lysate, sw620 cell lysate. It is guaranteed to work for WB in mouse, rat. Our Boster guarantee will cover your intended experiment even if the sample type has not been be directly tested.
Boster Scientific Support
Answered: 2019-10-25
Question
We have observed staining in mouse cervix carcinoma erythroleukemia. Are there any suggestions? Is anti-Aurora B/AURKB antibody supposed to stain cervix carcinoma erythroleukemia positively?
Verified Customer
Verified customer
Asked: 2019-06-03
Answer
Based on literature cervix carcinoma erythroleukemia does express AURKB. Based on Uniprot.org, AURKB is expressed in oocyte, liver spleen, lung, lymph muscle, cervix carcinoma, cervix carcinoma erythroleukemia, among other tissues. Regarding which tissues have AURKB expression, here are a few articles citing expression in various tissues:
Cervix carcinoma, Pubmed ID: 18669648, 18691976, 20068231
Cervix carcinoma, and Erythroleukemia, Pubmed ID: 23186163
Liver, and Spleen, Pubmed ID: 9858806
Lung, Lymph, and Muscle, Pubmed ID: 15489334
Boster Scientific Support
Answered: 2019-06-03
Question
We are currently using anti-Aurora B/AURKB antibody PA1806 for rat tissue, and we are happy with the WB results. The species of reactivity given in the datasheet says mouse, rat. Is it true that the antibody can work on goat tissues as well?
V. Brown
Verified customer
Asked: 2013-01-10
Answer
The anti-Aurora B/AURKB antibody (PA1806) has not been validated for cross reactivity specifically with goat tissues, though there is a good chance of cross reactivity. We have an innovator award program that if you test this antibody and show it works in goat you can get your next antibody for free. Please contact me if I can help you with anything.
Boster Scientific Support
Answered: 2013-01-10





