Product Info Summary
| SKU: | PA1030-1 |
|---|---|
| Size: | 100 μg/vial |
| Reactive Species: | Human |
| Host: | Rabbit |
| Application: | IF, IHC, ICC, WB |
Customers Who Bought This Also Bought
Product info
Product Name
Anti-DUT Antibody Picoband®
SKU/Catalog Number
PA1030-1
BA2182 is an alternative SKU for this antibody, used in previous lots.
Size
100 μg/vial
Form
Lyophilized
Description
Boster Bio Anti-DUT Antibody catalog # PA1030-1. Tested in ICC/IF, IHC, WB applications. This antibody reacts with Human. The brand Picoband indicates this is a premium antibody that guarantees superior quality, high affinity, and strong signals with minimal background in Western blot applications. Only our best-performing antibodies are designated as Picoband, ensuring unmatched performance.
Storage & Handling
Store at -20˚C for one year from date of receipt. After reconstitution, at 4˚C for one month. It can also be aliquotted and stored frozen at -20˚C for six months. Avoid repeated freeze-thaw cycles.
Cite This Product
Anti-DUT Antibody Picoband® (Boster Biological Technology, Pleasanton CA, USA, Catalog # PA1030-1)
Host
Rabbit
Contents
Each vial contains 4 mg Trehalose, 0.9 mg NaCl and 0.2 mg Na2HPO4.
Clonality
Polyclonal
Isotype
Rabbit IgG
Immunogen
A synthetic peptide corresponding to a sequence at the C-terminus of human DUT, different from the related rat and mouse sequences by two amino acids.
Cross-reactivity
No cross-reactivity with other proteins
Reactive Species
PA1030-1 is reactive to DUT in Human
Observed Molecular Weight
18 kDa
Calculated molecular weight
26.6 kDa
Background of DUT
Deoxyuridine triphosphate nucleotidohydrolase (dUTPase) is responsible for maintaining low intracellular levels of dUTP, thus preventing the incorporation of dUTP into DNA. dUTPase activity/expression can be down-regulated using siRNA specifically targeted to dUTPase mRNA and dUTPase plays a role in DNA nucleotide metabolism. This protein, present predominantly in the cytoplasm, contains 252 amino acids with a Mr of 26,704. It exhibits 35% identity with the E.coli dUTPase and 53% identity with the Saccharomyces cerevisiae enzyme. The nuclear and mitochondrial forms of dUTPase are encoded by the same gene with isoform-specific transcripts arising through the use of alternative 5-prime exons. Human dUTPase exhibits 92% identity with rat. Moreover, this enzyme has profound effects on the efficacy of agents that target thymidylate biosynthesis.
Antibody Validation
Boster validates all antibodies on WB, IHC, ICC, Immunofluorescence, and ELISA with known positive control and negative samples to ensure specificity and high affinity, including thorough antibody incubations.
Application & Images
Applications
PA1030-1 is guaranteed for IF, IHC, ICC, WB Boster Guarantee
Assay Dilutions Recommendation
The recommendations below provide a starting point for assay optimization. The actual working concentration varies and should be decided by the user.
Western blot, 0.1-0.5μg/ml, Human
Immunohistochemistry (Paraffin-embedded Section), 2-5μg/ml, Human
Immunocytochemistry/Immunofluorescence, 5 μg/ml, Human
Positive Control
WB: human HEL whole cell, human Jurkat whole cell, human SH-SY5Y whole cell, human Hela whole cell
IHC: human tonsil tissue
ICC/IF: Hela cell
Validation Images & Assay Conditions
Click image to see more details
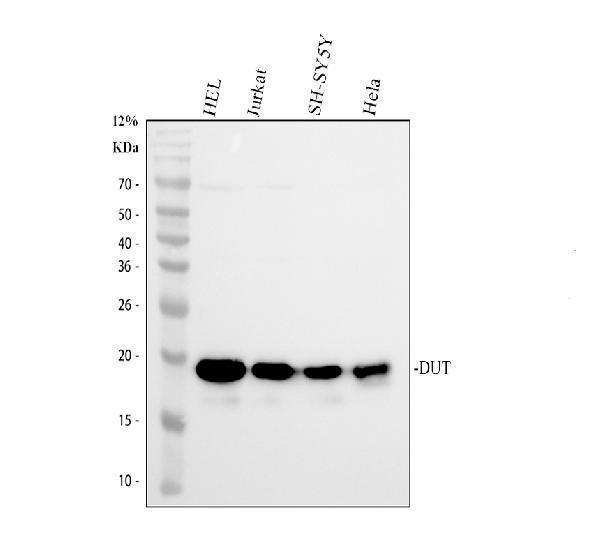
Western blot analysis of DUT using anti-DUT antibody (PA1030-1).
Electrophoresis was performed on a 12% SDS-PAGE gel at 80V (Stacking gel) / 120V (Resolving gel) for 2 hours. The sample well of each lane was loaded with 30 ug of sample under reducing conditions.
Lane 1: human HEL whole cell lysates,
Lane 2: human Jurkat whole cell lysates,
Lane 3: human SH-SY5Y whole cell lysates,
Lane 4: human Hela whole cell lysates.
After electrophoresis, proteins were transferred to a nitrocellulose membrane at 150 mA for 50-90 minutes. Blocked the membrane with 5% non-fat milk/TBS for 1.5 hour at RT. The membrane was incubated with rabbit anti-DUT antigen affinity purified polyclonal antibody (PA1030-1) at 0.5 μg/mL overnight at 4°C, then washed with TBS-0.1%Tween 3 times with 5 minutes each and probed with a goat anti-rabbit IgG-HRP secondary antibody at a dilution of 1:5000 for 1.5 hour at RT. The signal is developed using an ECL Plus Western Blotting Substrate (Catalog # AR1196-200) with Tanon 5200 system. A specific band was detected for DUT at approximately 18 kDa. The expected band size for DUT is at 18, 27 kDa.
Click image to see more details
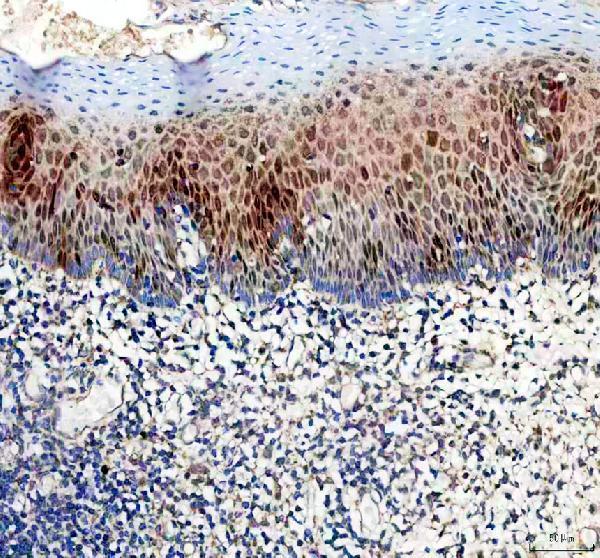
IHC analysis of DUT using anti-DUT antibody (PA1030-1).
DUT was detected in a paraffin-embedded section of human tonsil tissue. Heat mediated antigen retrieval was performed in EDTA buffer (pH 8.0, epitope retrieval solution). The tissue section was blocked with 10% goat serum. The tissue section was then incubated with 2 μg/ml rabbit anti-DUT Antibody (PA1030-1) overnight at 4°C. Peroxidase Conjugated Goat Anti-rabbit IgG was used as secondary antibody and incubated for 30 minutes at 37°C. The tissue section was developed using HRP Conjugated Rabbit IgG Super Vision Assay Kit (Catalog # SV0002) with DAB as the chromogen.
Click image to see more details
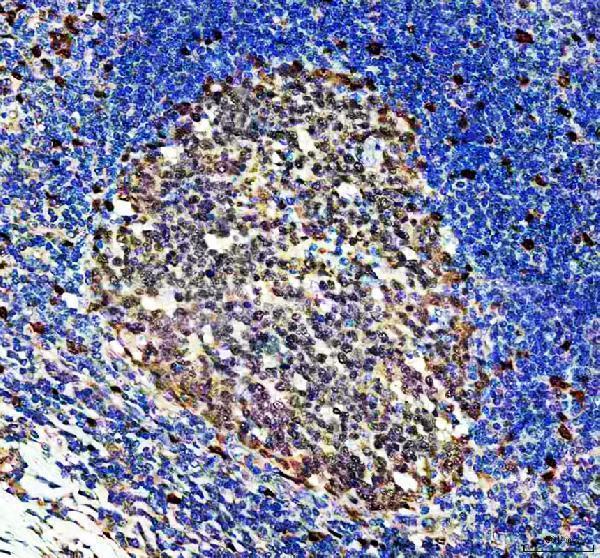
IHC analysis of DUT using anti-DUT antibody (PA1030-1).
DUT was detected in a paraffin-embedded section of human tonsil tissue. Heat mediated antigen retrieval was performed in EDTA buffer (pH 8.0, epitope retrieval solution). The tissue section was blocked with 10% goat serum. The tissue section was then incubated with 2 μg/ml rabbit anti-DUT Antibody (PA1030-1) overnight at 4°C. Peroxidase Conjugated Goat Anti-rabbit IgG was used as secondary antibody and incubated for 30 minutes at 37°C. The tissue section was developed using HRP Conjugated Rabbit IgG Super Vision Assay Kit (Catalog # SV0002) with DAB as the chromogen.
Click image to see more details
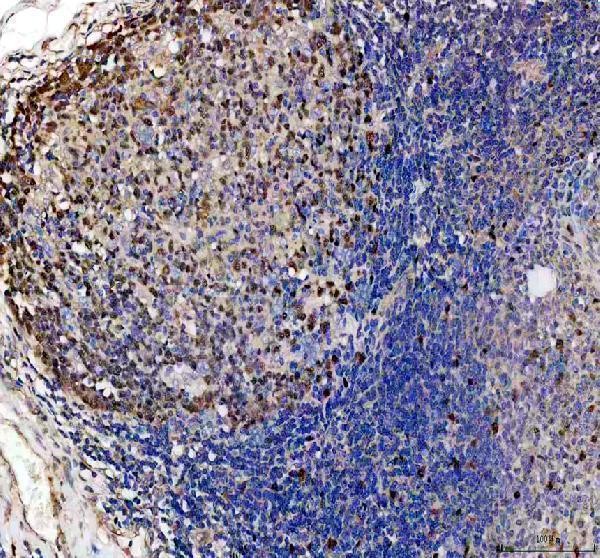
IHC analysis of DUT using anti-DUT antibody (PA1030-1).
DUT was detected in a paraffin-embedded section of human tonsil tissue. Heat mediated antigen retrieval was performed in EDTA buffer (pH 8.0, epitope retrieval solution). The tissue section was blocked with 10% goat serum. The tissue section was then incubated with 2 μg/ml rabbit anti-DUT Antibody (PA1030-1) overnight at 4°C. Peroxidase Conjugated Goat Anti-rabbit IgG was used as secondary antibody and incubated for 30 minutes at 37°C. The tissue section was developed using HRP Conjugated Rabbit IgG Super Vision Assay Kit (Catalog # SV0002) with DAB as the chromogen.
Click image to see more details
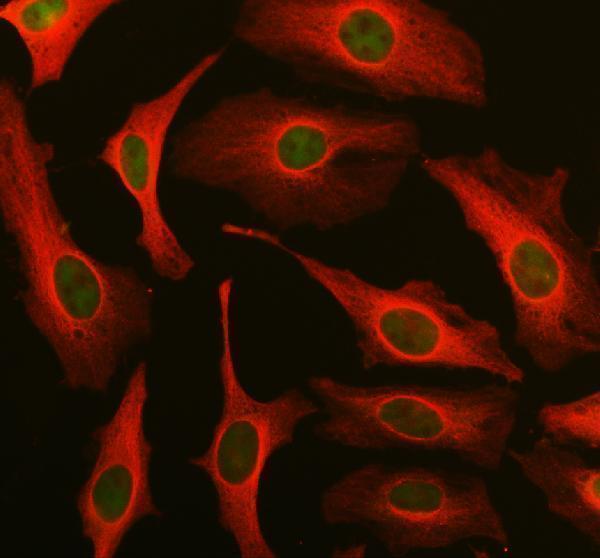
IF analysis of DUT using anti-DUT antibody (PA1030-1) and anti-Alpha Tubulin antibody (M03989-3).
DUT was detected in an immunocytochemical section of Hela cells. Enzyme antigen retrieval was performed using IHC enzyme antigen retrieval reagent (AR0022) for 15 mins. The cells were blocked with 10% goat serum. And then incubated with 5 μg/mL rabbit anti-DUT Antibody (PA1030-1) and mouse anti-Alpha Tubulin antibody (M03989-3) overnight at 4°C. Fluoro488 Conjugated Goat Anti-Rabbit IgG (BA1127) and Cy3 Conjugated Goat Anti-Mouse IgG (BA1031) were used as secondary antibody at 1:500 dilution and incubated for 30 minutes at 37°C. Visualize using a fluorescence microscope and filter sets appropriate for the label used.
Specific Publications For Anti-DUT Antibody Picoband® (PA1030-1)
Loading publications
Recommended Resources
Here are featured tools and databases that you might find useful.
- Boster's Pathways Library
- Protein Databases
- Bioscience Research Protocol Resources
- Data Processing & Analysis Software
- Photo Editing Software
- Scientific Literature Resources
- Research Paper Management Tools
- Molecular Biology Software
- Primer Design Tools
- Bioinformatics Tools
- Phylogenetic Tree Analysis
Customer Reviews
Have you used Anti-DUT Antibody Picoband®?
Share your experimental results or join a short interview to earn up to $1,000 in product credits or other rewards.
0 Reviews For Anti-DUT Antibody Picoband®
Customer Q&As
Have a question?
Find answers in Q&As, reviews.
Can't find your answer?
Submit your question
7 Customer Q&As for Anti-DUT Antibody Picoband®
Question
We are currently using anti-DUT antibody PA1030-1 for human tissue, and we are satisfied with the WB results. The species of reactivity given in the datasheet says human, mouse, rat. Is it true that the antibody can work on canine tissues as well?
Verified Customer
Verified customer
Asked: 2020-04-07
Answer
The anti-DUT antibody (PA1030-1) has not been tested for cross reactivity specifically with canine tissues, but there is a good chance of cross reactivity. We have an innovator award program that if you test this antibody and show it works in canine you can get your next antibody for free. Please contact me if I can help you with anything.
Boster Scientific Support
Answered: 2020-04-07
Question
I see that the anti-DUT antibody PA1030-1 works with IHC, what is the protocol used to produce the result images on the product page?
Verified Customer
Verified customer
Asked: 2019-12-17
Answer
You can find protocols for IHC on the "support/technical resources" section of our navigation menu. If you have any further questions, please send an email to support@bosterbio.com
Boster Scientific Support
Answered: 2019-12-17
Question
I was wanting to use your anti-DUT antibody for IHC for human brain on frozen tissues, but I want to know if it has been tested for this particular application. Has this antibody been tested and is this antibody a good choice for human brain identification?
N. Collins
Verified customer
Asked: 2019-07-23
Answer
You can see on the product datasheet, PA1030-1 anti-DUT antibody has been tested for IHC, WB on human, mouse, rat tissues. We have an innovator award program that if you test this antibody and show it works in human brain in IHC-frozen, you can get your next antibody for free.
Boster Scientific Support
Answered: 2019-07-23
Question
We want to test anti-DUT antibody PA1030-1 on human brain for research purposes, then I may be interested in using anti-DUT antibody PA1030-1 for diagnostic purposes as well. Is the antibody suitable for diagnostic purposes?
Verified Customer
Verified customer
Asked: 2019-03-27
Answer
The products we sell, including anti-DUT antibody PA1030-1, are only intended for research use. They would not be suitable for use in diagnostic work. If you have the means to develop a product into diagnostic use, and are interested in collaborating with us and develop our product into an IVD product, please contact us for more discussions.
Boster Scientific Support
Answered: 2019-03-27
Question
Is there a BSA free version of anti-DUT antibody PA1030-1 available?
K. Li
Verified customer
Asked: 2017-05-29
Answer
Thanks for your recent telephone inquiry. I can confirm that some lots of this anti-DUT antibody PA1030-1 are BSA free. For now, these lots are available and we can make a BSA free formula for you free of charge. It will take 3 extra days to prepare. If you require this antibody BSA free again in future, please do not hesitate to contact me and I will be pleased to check which lots we have in stock that are BSA free.
Boster Scientific Support
Answered: 2017-05-29
Question
Will PA1030-1 anti-DUT antibody work on parafin embedded sections? If so, which fixation method do you recommend we use (PFA, paraformaldehyde, other)?
Z. Parker
Verified customer
Asked: 2016-04-29
Answer
It shows on the product datasheet, PA1030-1 anti-DUT antibody as been validated on IHC. It is best to use PFA for fixation because it has better tissue penetration ability. PFA needs to be prepared fresh before use. Long term stored PFA turns into formalin, as the PFA molecules congregate and become formalin.
Boster Scientific Support
Answered: 2016-04-29
Question
I appreciate helping with my inquiry over the phone. Here are the WB image, lot number and protocol we used for brain using anti-DUT antibody PA1030-1. Let me know if you need anything else.
M. Zhao
Verified customer
Asked: 2015-09-22
Answer
Thanks for the data. You have provided everything we needed. Our lab team are working to resolve your inquiry as quickly as possible, and we appreciate your patience and understanding! Please let me know if there is anything you need in the meantime.
Boster Scientific Support
Answered: 2015-09-22





