Product Info Summary
| SKU: | A00284-1 |
|---|---|
| Size: | 100 μg/vial |
| Reactive Species: | Human, Mouse, Rat |
| Host: | Rabbit |
| Application: | ELISA, Flow Cytometry, IF, ICC, WB |
Customers Who Bought This Also Bought
Product info
Product Name
Anti-NF-kB p65/RELA Antibody Picoband®
SKU/Catalog Number
A00284-1
Size
100 μg/vial
Form
Lyophilized
Description
Boster Bio Anti-NF-kB p65/RELA Antibody Picoband® catalog # A00284-1. Tested in ELISA, Flow Cytometry, IF, ICC, WB applications. This antibody reacts with Human, Mouse, Rat. The brand Picoband indicates this is a premium antibody that guarantees superior quality, high affinity, and strong signals with minimal background in Western blot applications. Only our best-performing antibodies are designated as Picoband, ensuring unmatched performance.
Storage & Handling
Store at -20˚C for one year from date of receipt. After reconstitution, at 4˚C for one month. It can also be aliquotted and stored frozen at -20˚C for six months. Avoid repeated freeze-thaw cycles.
Cite This Product
Anti-NF-kB p65/RELA Antibody Picoband® (Boster Biological Technology, Pleasanton CA, USA, Catalog # A00284-1)
Host
Rabbit
Contents
Each vial contains 4 mg Trehalose, 0.9 mg NaCl and 0.2 mg Na2HPO4.
Clonality
Polyclonal
Isotype
Rabbit IgG
Immunogen
E. coli-derived human NF-kB p65 recombinant protein (Position: F99-S551).
Cross-reactivity
No cross-reactivity with other proteins.
Reactive Species
A00284-1 is reactive to RELA in Human, Mouse, Rat
Observed Molecular Weight
65-70 kDa
Calculated molecular weight
60.2 kDa
Background of RELA
Transcription factor p65, also known as NFKB3 or NF-kB p65, is a protein that in humans is encoded by the RELA gene. It is mapped to 11q13.1. NFKB is an essential transcription factor complex involved in all types of cellular processes, including cellular metabolism, chemotaxis, etc, and it may play a role in inflammatory conditions of the peripheral nervous system. Phosphorylation and acetylation of NFKB3 are crucial post-translational modifications required for NFKB activation. It has also been shown to modulate immune responses, and activation of NFKB3 is positively associated with multiple types of cancer. In addition to that, NFKB3 antagonizes TNFR1-JNK proliferative signals in epidermis and plays a nonredundant role in restraining epidermal growth.
Antibody Validation
Boster validates all antibodies on WB, IHC, ICC, Immunofluorescence, and ELISA with known positive control and negative samples to ensure specificity and high affinity, including thorough antibody incubations.
Application & Images
Applications
A00284-1 is guaranteed for ELISA, Flow Cytometry, IF, ICC, WB Boster Guarantee
Recommend Dilution
| Application | Dilution | Species |
|---|---|---|
| Western blot | 0.1-0.5 μg/ml | |
| Immunocytochemistry/Immunofluorescence | 5 μg/ml | |
| Immunoprecipitation | 0.5-2 μg/ml | |
| Flow Cytometry (Fixed) | 1-3μg/1x106 cells | |
| ELISA | 0.1-0.5 μg/ml |
Tested application
Suggested blocking solution with 5% non-fat milk or BSA; (*)Recommended protein loading: 20-40 µg per lane
Validation Images & Assay Conditions
Click image to see more details
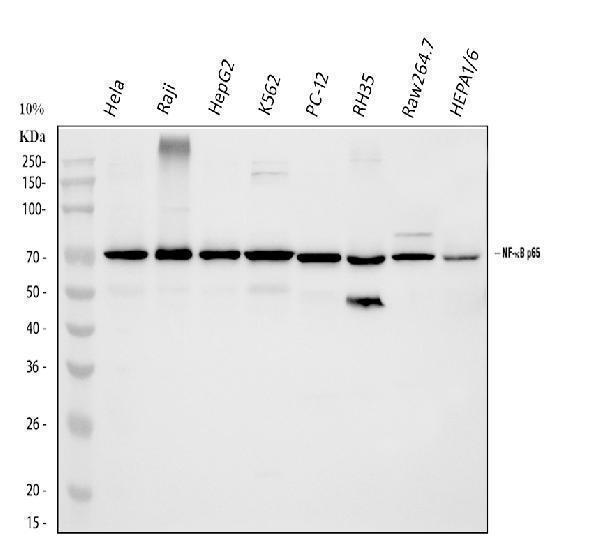
Western blot analysis of NF-kB p65 using anti-NF-kB p65 antibody (A00284-1).
Electrophoresis was performed on a 5-20% SDS-PAGE gel at 70V (Stacking gel) / 90V (Resolving gel) for 2-3 hours. The sample well of each lane was loaded with 30 ug of sample under reducing conditions.
Lane 1: human Hela whole cell lysates,
Lane 2: human Raji whole cell lysates,
Lane 3: human HepG2 whole cell lysates,
Lane 4: human K562 whole cell lysates,
Lane 5: rat PC-12 whole cell lysates,
Lane 6: rat RH35 whole cell lysates,
Lane 7: mouse RAW264.7 whole cell lysates,
Lane 8: mouse HEPA1/6 whole cell lysates.
After electrophoresis, proteins were transferred to a nitrocellulose membrane at 150 mA for 50-90 minutes. Blocked the membrane with 5% non-fat milk/TBS for 1.5 hour at RT. The membrane was incubated with rabbit anti-NF-kB p65 antigen affinity purified polyclonal antibody (Catalog # A00284-1) at 0.5 μg/mL overnight at 4°C, then washed with TBS-0.1%Tween 3 times with 5 minutes each and probed with a goat anti-rabbit IgG-HRP secondary antibody at a dilution of 1:5000 for 1.5 hour at RT. The signal is developed using an Enhanced Chemiluminescent detection (ECL) kit (Catalog # EK1002) with Tanon 5200 system. A specific band was detected for NF-kB p65 at approximately 70 kDa. The expected band size for NF-kB p65 is at 60 kDa.
Click image to see more details
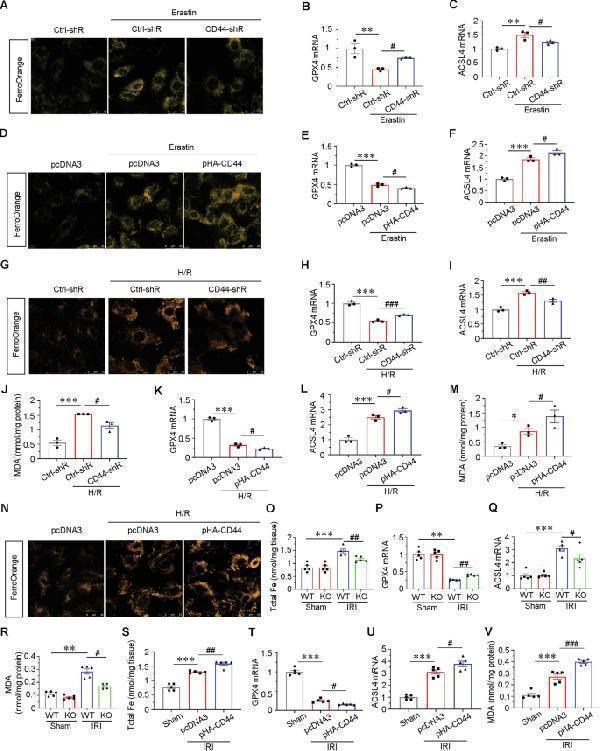
CD44 promotes tubular cell injury and AKI partially through ferroptosis. A HKC-8 was transfected with Ctrl-shR or CD44-shR plasmid and then treated with 5 μM erastin for 24 h. Representative micrographs show the Fe 2+ content via FerroOrange staining in different groups, as indicated. Scale bar, 25 μm. B and C Quantitative results of QPCR showing relative ( B ) GPX4 and ( C ) ACSL4 mRNA levels among different groups. ** P < 0.01 versus Ctrl-shR group; # P < 0.05 versus erastin with ctrl-shR ( n = 3). D HKC-8 was transfected with pcDNA3 or pHA-CD44 overexpression plasmid and then treated with 5 μM erastin for 24 h. Representative micrographs show the Fe 2+ content via FerroOrange staining in different groups, as indicated. Scale bar, 25 μm. E and F Quantitative result of QPCR showing relative ( E ) GPX4 and ( F ) ACSL4 mRNA levels among different groups. *** P < 0.001 versus pcDNA3 group; # P < 0.05 versus erastin with pcDNA3 ( n = 3). G Representative micrographs show the Fe 2+ content via FerroOrange staining in different groups, as indicated. Scale bar, 25 μm. H and I Quantitative result of QPCR showing relative H GPX4 and I ACSL4 mRNA level among different groups. *** P < 0.001 versus Ctrl-shR group; ## P < 0.01, ### P < 0.001 versus H/R with ctrl-shR ( n = 3). J Quantitative result showing MDA content among different groups. *** P < 0.001 versus Ctrl-shR group; # P < 0.05 versus H/R with ctrl-shR ( n = 3). K and L Quantitative result of QPCR showing relative ( K ) GPX4 and ( L ) ACSL4 mRNA levels among different groups. *** P < 0.001 versus pcDNA3 group; # P < 0.05 versus H/R with pcDNA3 ( n = 3). M Quantitative result showing MDA content among different groups. *** P < 0.001 versus pcDNA3 group; # P < 0.05 versus H/R with pcDNA3 ( n = 3). N Representative micrographs show the Fe 2+ content via FerroOrange staining in different groups, as indicated. Scale bar, 25 μm. O Quantitative result showing total iron content in kidney tissue among different groups. *** P < 0.001 versus wild-type mice upon sham group; ## P < 0.01 versus wild-type mice upon IRI group ( n = 5). P and Q Quantitative result of QPCR showing relative P GPX4 and Q ACSL4 mRNA levels among different groups. ** P < 0.01, *** P < 0.001 versus wild-type mice upon sham group; # P < 0.05, ## P < 0.01 versus wild-type mice upon IRI group ( n = 5). R Quantitative result showing MDA content in kidney tissue among different groups. ** P < 0.01 versus wild-type mice upon sham group; # P < 0.05 versus wild-type mice upon IRI group ( n = 5). S Quantitative result showing total iron content in kidney tissue among different groups. *** P < 0.001 versus sham group; ## P < 0.01 versus IRI group injected with pcDNA3 ( n = 5). T and U Quantitative result of QPCR showing relative S GPX4 and T ACSL4 mRNA level among different groups. *** P < 0.001 versus sham group; # P < 0.05 versus IRI group injected with pcDNA3 ( n = 5). V Quantitative result showing MDA content in kidney tissue among different groups. *** P < 0.001 versus sham group; ### P < 0.001 versus IRI group injected with pcDNA3 ( n = 5).
Index in PubMed under a CC BY license. PMID: 39979265
Click image to see more details
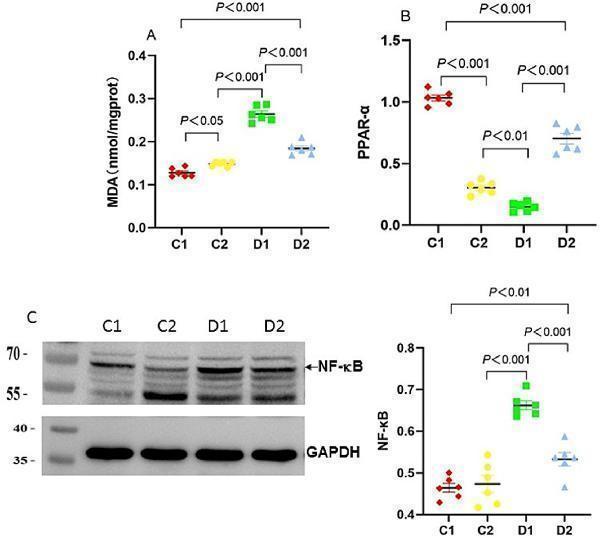
Effects of n-6 PUFA on liver lipid peroxidation and the inflammatory marker NF-κB in rats with NASH induced by a choline-deficient diet. (A) Liver MDA levels, (B) PPAR-α mRNA expression in the liver. Data are expressed as mean ±SEM; n = 6/group. (C) NF-κB protein expression (~65 kDa) in the liver as analyzed by Western blotting, normalized to GAPDH, with a representative blot (left) and quantification (right). Protein molecular weight standards (kDa) are labeled on the left of each blot. Data are expressed as mean ± SEM; n = 6/group.
Index in PubMed under a CC BY license. PMID: 40626231
Click image to see more details
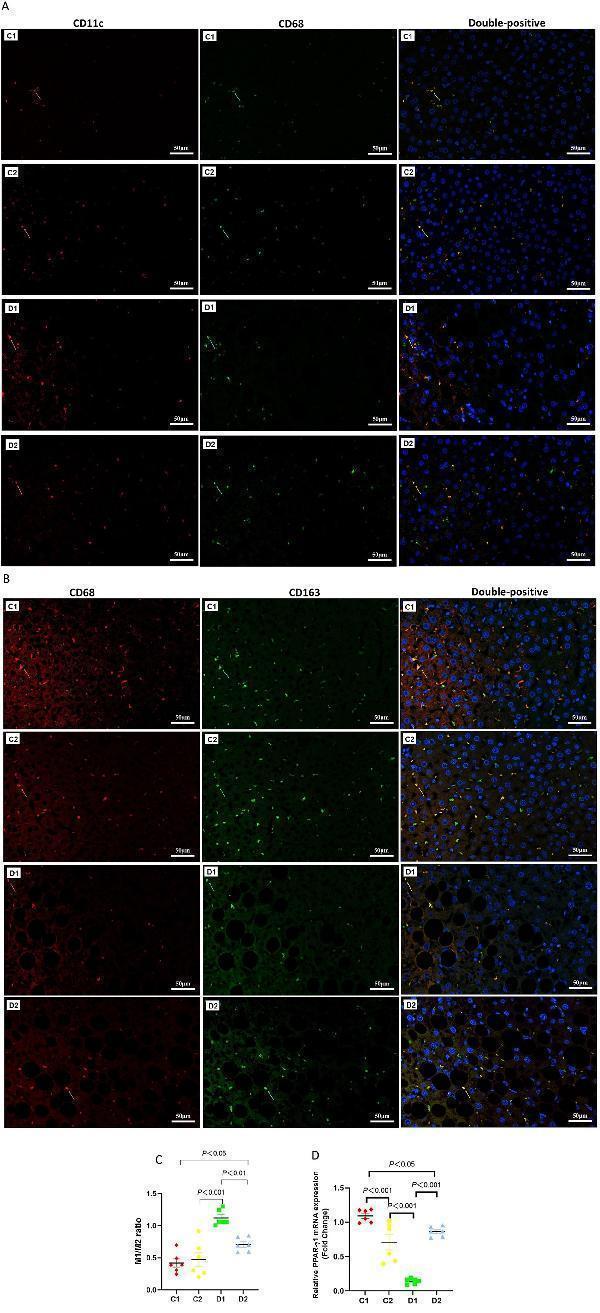
Effects of n-6 PUFA on liver macrophage phenotype in rats with NASH induced by a choline-deficient diet. (A) M1-type Kupffer cells (KCs) identified by double staining: red arrows show CD11c-positive cells, green arrows show CD68-positive cells, and yellow arrows highlight CD11c and CD68 double-positive M1-type KCs (Scale bar – 50 μM). (B) M2-type KCs identified similarly, with red arrows indicating CD163-positive cells, green arrows showing CD68-positive cells, and yellow arrows marking CD163 and CD68 double-positive M2-type KCs (Scale bar – 50 μM). For (A,B) (see ) for full-size photomicrographs. (C) M1/M2 phenotype ratio (unitless), calculated as the proportion of CD68 + CD11c + to CD68 + CD163 + cells. (D) Relative PPAR-γ2 mRNA expression (fold change normalized to GAPDH) in the liver, which is linked to macrophage polarization and inflammation. Data are expressed as mean ±SEM; n = 6/group.
Index in PubMed under a CC BY license. PMID: 40626231
Click image to see more details
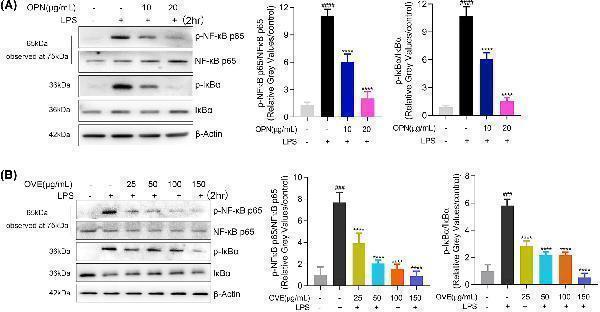
Effects of OPN and OVE on activation of NF-ĸB pathways in LPS-stimulated RAW264.7 cells. Cells were pretreated with OVE or OPN at different concentrations for 1 h. Expression levels of p-NF-ĸB p65, NF-ĸB p65, p-IĸBα, and IĸBα were detected after 24 h of LPS treatment. (A) OPN treatment. (B) OVE treatment. All experiments were carried out in triplicates and data are presented as means ± SDs; one-way ANOVA analysis was adopted for multiple comparisons; ###P < 0.001, ####P < 0.0001, compared to the untreated control group; ****P < 0.0001, compared to the LPS control group.
Index in PubMed under a CC BY license. PMID: 39455284
Click image to see more details
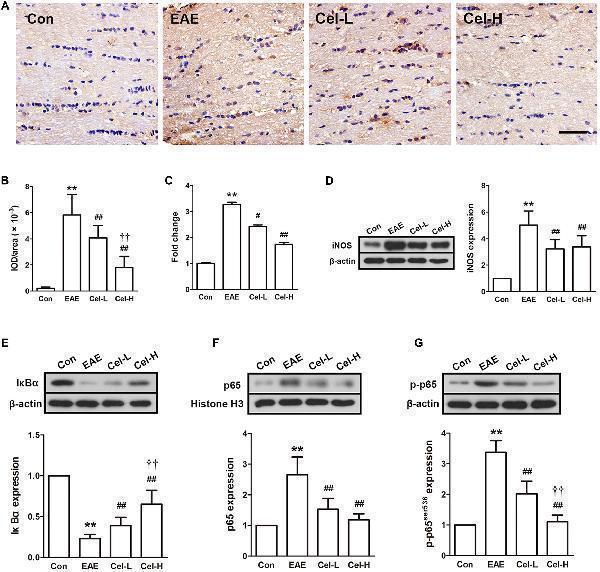
Celastrol inhibited inducible nitric oxide synthase (iNOS) expression and activation of NF-κB in optic nerve in EAE rats. (A) IHC staining of iNOS in optic nerve. (B) Quantification of iNOS-positive areas. (C) Quantitative real-time PCR analysis of iNOS expression. (D) Western blot analysis of iNOS expression. (E–G) Western blot analysis of IκBα, p65 and p-p65 expression, respectively. Treatment of celastrol reduced expression of iNOS and inhibited the activation of NF-κB in optic nerve in EAE rats. Scale bar: 100 μm. Data were shown as mean ± SD, n = 5. ∗∗ P < 0.01 versus control group, ## P < 0.01 versus EAE group, †† P < 0.01 versus low dosage of celastrol group.
Index in PubMed under a CC BY license. PMID: 28239352
Click image to see more details
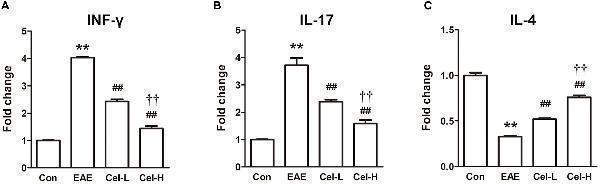
Celastrol regulated production of cytokines and activation of NF-κB in spinal cords of EAE rats. Celastrol dose-dependently downregulated the mRNA expression of INF-γ (A) and IL-17 (B) but upregulated IL-4 (C) in spinal cord of EAE rats. Data were shown as mean ± SD, n = 5. ∗∗ P < 0.01 versus control group, ## P < 0.01 versus EAE group, †† P < 0.01 versus low dosage of celastrol group.
Index in PubMed under a CC BY license. PMID: 28239352
Click image to see more details
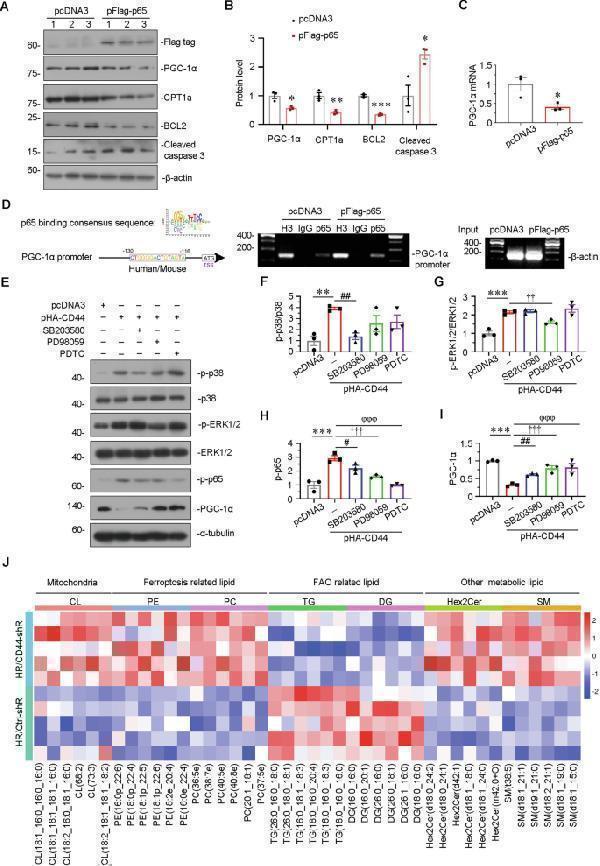
CD44 induces tubular cell injury through MAPK-NF-κB p65-silenced PGC-1α signaling. A and B HKC-8 was transfected with pcDNA3 or p-Flag-p65 overexpression plasmid for 24 h. Representative western blot ( A ) and graphical presentations of PGC-1α, CPT1a, BCL2, and cleaved caspase 3 protein expression levels are shown. * P < 0.05, ** P < 0.01, *** P < 0.001 versus pcDNA3 groups ( n = 3). C Quantitative PCR result showing relative mRNA level of PGC-1α. * P < 0.05 versus pcDNA3 groups ( n = 3). D Representative ChIP assay results showing the binding of p65 to PGC-1α gene promoter region. HKC-8 cells were transfected with pcDNA3 or p-Flag-p65 for 24 h. Cell lysates were precipitated with an antibody against p65, histone H3, or nonimmune IgG, and ChIP assay was performed for PGC-1α gene promoters. Total diluted lysate was used as total genomic input DNA. E–I HKC-8 was pre-treated with SB203580, PD98059 or PDTC at 1 h before transfection with pcDNA3 or p-HA-CD44 plasmid for 24 h. Representative western blot ( E ) and graphical presentations of F p-p38/p38, G p-ERK1/2/ERK1/2, H p-p65, and I PGC-1α protein expression levels are shown. ** P < 0.01, *** P < 0.001 versus pcDNA3 group; # P < 0.05, ## P < 0.01 versus pHA-CD44 group; †† P < 0.01, ††† P < 0.001 versus pHA-CD44 group; φφφ P < 0.001 versus pHA-CD44 group ( n = 3). J The heatmap exhibiting differentiated lipids of lipidomics sequencing between H/R with Crtl-shR group and H/R with CD44-shR group.
Index in PubMed under a CC BY license. PMID: 39979265
Click image to see more details
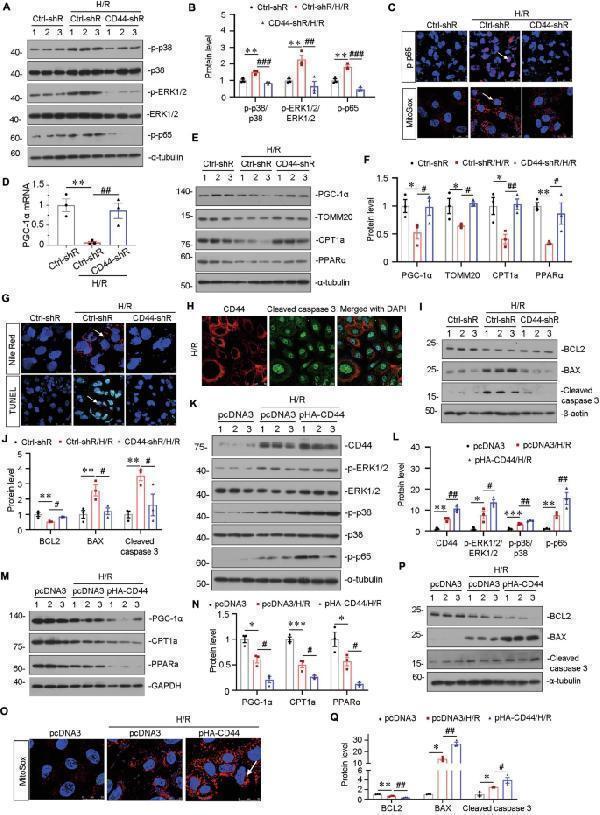
CD44 aggravates mitochondrial and FAO dysfunction and drives cell apoptosis through MAPK and NF-κB p65 signaling in vitro. A and B HKC-8 was transfected with Ctrl-shR or CD44-shR and then were incubated in basal culture medium in a 1% O 2 environment for 24 h and then were reoxygenated in normal O 2 for 6 h. Representative western blot ( A ) and graphical presentations of p-p38/p38, p-ERK1/2/ERK1/2, and p-p65 protein expression levels are shown. ** P < 0.01 versus Crtl-shR group; ## P < 0.01, ### P < 0.001 versus H/R with Crtl-shR group ( n = 3). C Representative micrographs show the expression of p-p65 and MitoSox staining in different groups, as indicated. Arrows indicate positive staining. Cells cultured on coverslips were stained with an antibody against p-p65 or were stained with MitoSox. Scale bar, 25 or 50 μm. D Graphic presentation shows the relative mRNA levels of PGC-1α in different groups as indicated. ** P < 0.01 versus Crtl-shR group; ## P < 0.01 versus H/R with Crtl-shR group ( n = 3). E and F Representative western blot ( E ) and graphical representations of PGC-1α, TOMM20, CPT1a, and PPARα protein expression levels are shown. * P < 0.05, ** P < 0.01 versus Crtl-shR group; # P < 0.05, ## P < 0.01 versus H/R with Crtl-shR group ( n = 3). G Representative micrographs show Nile Red staining and TUNEL assay in different groups, as indicated. Arrows indicate positive staining. Cells cultured on coverships were stained with Nile Red or TUNEL assay. Scale bar, 25 or 50 μm. H Co-localization of CD44 and cleaved caspase 3 in HKC-8 after H/R treatment. Cells cultured on coverships were subjected to immunostaining of CD44 (red) and cleaved caspase 3 (green). Scale bar, 50 μm. I and J Representative western blot ( I ) and graphical representations of BCL2, BAX, and cleaved caspase 3 protein expression levels are shown. ** P < 0.01 versus Crtl-shR group; # P < 0.05 versus H/R with Crtl-shR group ( n = 3). K and L HKC-8 was transfected with pcDNA3 or p-HA-CD44 overexpression plasmid and then were incubated in basal culture medium in a 1% O 2 environment for 24 h and then were reoxygenated in normal O 2 for 6 h. Representative western blot ( K ) and graphical representations of CD44, p-ERK1/2/ERK1/2, p-p38/p38 and p-p65 protein expression levels are shown. * P < 0.05, ** P < 0.01, *** P < 0.001 versus pcDNA3 group; # P < 0.05, ## P < 0.01 versus H/R with pcDNA3 group ( n = 3). M and N Representative western blot ( M ) and graphical presentations of PGC-1α, CPT1a, and PPARα protein expression levels are shown. * P < 0.05, *** P < 0.001 versus pcDNA3 group; # P < 0.05 versus H/R with pcDNA3 group ( n = 3). O Representative micrographs show MitoSox staining in different groups, as indicated. Arrow indicates positive staining. Cells cultured on coverships were stained with MitoSox. Scale bar, 25 μm. P and Q Representative western blot ( P ) and graphical representations of BCL2, BAX, and cleaved caspase 3 protein expression levels are shown. * P < 0.05, ** P < 0.01 versus pcDNA3 group; # P < 0.05; ## P < 0.01 versus H/R with pcDNA3 group ( n = 3).
Index in PubMed under a CC BY license. PMID: 39979265
Click image to see more details
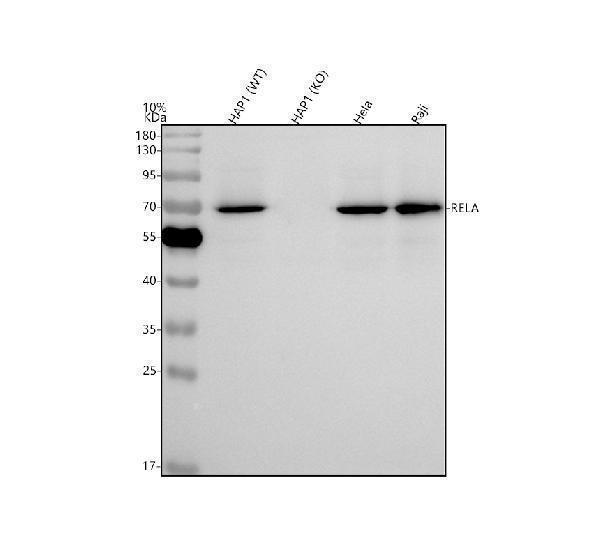
Western blot analysis of NF-kB p65 using anti-NF-kB p65 antibody (A00284-1).
Electrophoresis was performed on a 5-20% SDS-PAGE gel at 70V (Stacking gel) / 90V (Resolving gel) for 2-3 hours. The sample well of each lane was loaded with 30 ug of sample under reducing conditions.
Lane 1: human HAP1-WT whole cell lysates,
Lane 2: human HAP1-NF-kB p65 KO whole cell lysates,
Lane 3: human Hela whole cell lysates,
Lane 4: human Raji whole cell lysates.
After electrophoresis, proteins were transferred to a nitrocellulose membrane at 150 mA for 50-90 minutes. Blocked the membrane with 5% non-fat milk/TBS for 1.5 hour at RT. The membrane was incubated with rabbit anti-NF-kB p65 antigen affinity purified polyclonal antibody (Catalog # A00284-1) at 0.5 μg/mL overnight at 4°C, then washed with TBS-0.1%Tween 3 times with 5 minutes each and probed with a goat anti-rabbit IgG-HRP secondary antibody at a dilution of 1:5000 for 1.5 hour at RT. The signal is developed using an Enhanced Chemiluminescent detection (ECL) kit (Catalog # EK1002) with Tanon 5200 system. A specific band was detected for NF-kB p65 at approximately 70 kDa. The expected band size for NF-kB p65 is at 60 kDa.
Click image to see more details
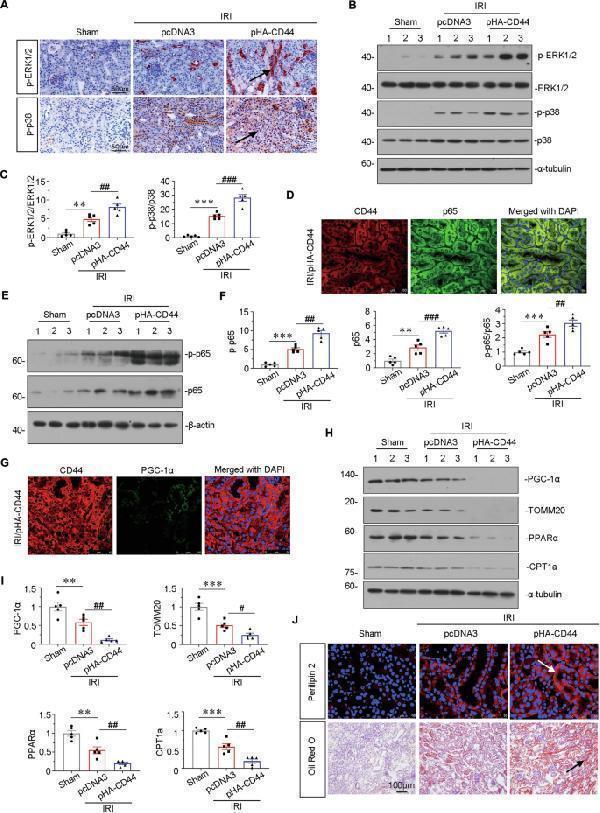
Ectopic expression of CD44 impairs mitochondrial function and FAO through activating MAPK and NF-κB p65 signaling. A Representative micrographs show the expression of p-ERK1/2 and p-p38 in different groups, as indicated. Paraffin sections were stained with antibodies against p-ERK1/2 and p-p38. Arrows indicate positive staining. Scale bar, 50 μm. B and C Representative western blot ( B ) and graphical presentations of p-ERK1/2/ERK1/2 and p-p38/p38 protein levels are shown. ** P < 0.01, *** P < 0.001 versus sham group; ## P < 0.01, ### P < 0.001 versus IRI group injected with pcDNA3 ( n = 5). D Co-localization of CD44 and p65 in CD44 overexpression mice upon IRI. Frozen renal sections were subjected to immunostaining of CD44 (red) and p65 (green). Scale bar, 50 μm. E and F Representative western blot ( E ) and graphical presentations of p-p65, p65, and p-p65/p65 protein levels are shown. ** P < 0.01, *** P < 0.001 versus sham group; ## P < 0.01, ### P < 0.001 versus IRI group injected with pcDNA3 ( n = 5). G Co-localization of CD44 and PGC-1α in CD44 overexpression mice upon IRI. Frozen renal sections were subjected to immunostaining of CD44 (red) and PGC-1α (green). Scale bar, 50 μm. H and I Representative western blot ( H ) and graphical presentations of PGC-1α, TOMM20, PPARα, and CPT1a protein expression levels are shown. ** P < 0.01, *** P < 0.001 versus sham group; # P < 0.05, ## P < 0.01 versus IRI group injected with pcDNA3 ( n = 5). J Representative micrographs show the expression of perilipin 2, and LDs via Oil Red O staining in different groups, as indicated. Frozen kidney sections were subjected to Oil Red O staining or stained with an antibody against perilipin 2. Arrows indicate positive staining. Scale bar, 50 or 100 μm.
Index in PubMed under a CC BY license. PMID: 39979265
Click image to see more details
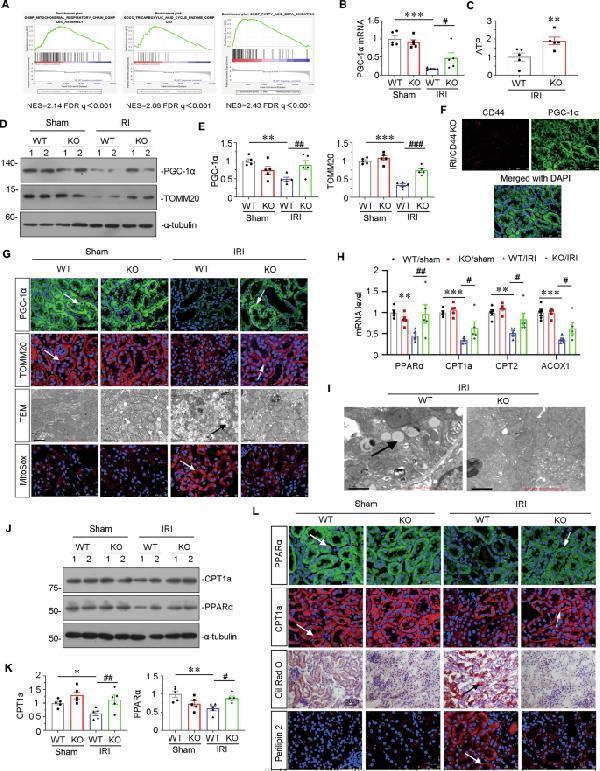
CD44 knockout ameliorates mitochondrial dysfunction and FAO deficiency in IRI mice. A GSEA shows that positive regulation of mitochondrial function and FAO was enriched in CD44 knockout mice versus wild-type mice upon IRI. NES, normalized enrichment score; FDR q -value < 0.25. B Graphic presentation shows the relative levels of renal expression of PGC-1α mRNA in different groups as indicated. *** P < 0.001 versus wild-type mice upon sham group; # P < 0.05 versus wild-type mice upon IRI group ( n = 5). C Graphic presentation shows the relative levels of ATP production in 2 groups as indicated. ** P < 0.01 versus wild-type upon IRI group ( n = 5). D and E Representative western blot ( D ) and graphical presentations of PGC-1α and TOMM20 protein expression levels are shown. ** P < 0.01, *** P < 0.001 versus wild-type mice upon sham group; ## P < 0.01, ### P < 0.001 versus wild-type mice upon IRI group ( n = 5). F Co-localization of CD44 and PGC-1α in CD44 gene ablation mice upon IRI. Frozen kidney sections were subjected to immunostaining of CD44 (red) and PGC-1α (green). Scale bar, 50 μm. G Representative micrographs show the expression of PGC-1α, TOMM20, mitochondrial morphology via TEM, and mitochondrial ROS via MitoSox staining in different groups, as indicated. Arrows indicate positive staining or impaired mitochondria. Scale bar, 50 or 1 μm, as indicated. H Graphic presentation shows the relative mRNA levels of renal expression of PPARα, CPT1, CPT2, and ACOX1 mRNA in different groups as indicated. ** P < 0.01, *** P < 0.001 versus wild-type mice upon sham group; # P < 0.05, ## P < 0.01 versus wild-type mice upon IRI group ( n = 5). I Representative micrographs show the abundance of LDs via TEM in 2 groups, as indicated. Arrows indicate LDs. Scale bar, 1 μm. J and K Representative western blot ( J ) and graphical presentations of CPT1a and PPARα protein expression levels are shown. * P < 0.05, ** P < 0.01 versus wild-type mice upon sham group; # P < 0.05, ## P < 0.01 versus wild-type mice upon IRI group ( n = 5). L Representative micrographs show the expression of PPARα, CPT1a, perilipin 2, and LDs via Oil Red O staining in different groups, as indicated. Arrows indicate positive staining. Frozen kidney sections were subjected to Oil Red O staining or stained with antibodies against PPARα, CPT1a, and perilipin 2. Scale bar, 50 μm.
Index in PubMed under a CC BY license. PMID: 39979265
Click image to see more details
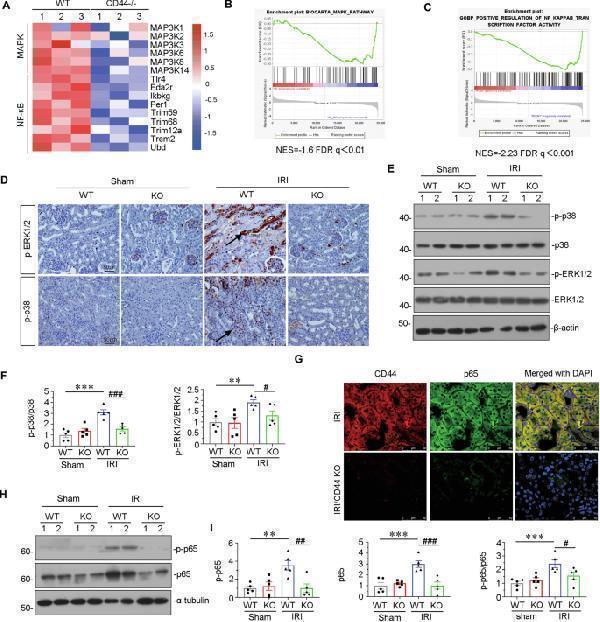
CD44 promotes AKI progression through inducing MAPK and NF-κB p65 signaling. A Representative heatmap gene expression of RNA sequencing analysis shows that CD44 is involved with MAPK and NF-κB signaling pathway. B and C GSEA shows that negative regulation of MAPK and NF-κB pathway was enriched in CD44 knockout mice versus wild-type mice upon IRI. NES, normalized enrichment score; FDR q -value < 0.25. D Representative micrographs show the expression of p-ERK1/2 and p-p38 in different groups, as indicated. Paraffin sections were stained with antibodies against p-ERK1/2 and p-p38. Arrows indicate positive staining. Scale bar, 50 μm. E and F Representative western blot ( E ) and graphical presentations of p-p38/p38 and p-ERK1/2/ERK1/2 protein levels are shown. ** P < 0.01, *** P < 0.001 versus wild-type mice upon sham group; # P < 0.05, ### P < 0.001 versus wild-type mice upon IRI group ( n = 5). G Co-localization of CD44 and p65 in 2 groups, as indicated. Frozen kidney sections were subjected to immunostaining of CD44 (red) and p65 (green). Scale bar, 50 μm. H and I Representative western blot ( H ) and graphical presentations of p-p65, p65, and p-p65/p65 protein levels are shown. ** P < 0.01, *** P < 0.001 versus wild-type mice upon sham group; # P < 0.05, ## P < 0.01, ### P < 0.001 versus wild-type mice upon IRI group ( n = 5).
Index in PubMed under a CC BY license. PMID: 39979265

Click image to see more details
Western blot analysis of NF-kB p65 using anti-NF-kB p65 antibody (A00284-1).
Electrophoresis was performed on a 5-20% SDS-PAGE gel at 80V (Stacking gel) / 120V (Resolving gel) for 2 hours. The sample well of each lane was loaded with 30 ug of sample under reducing conditions.
Lane 1-5: mouse tissue,
After electrophoresis, proteins were transferred to a nitrocellulose membrane at 150 mA for 50-90 minutes. Blocked the membrane with 5% non-fat milk/TBS for 1.5 hour at RT. The membrane was incubated with rabbit anti-NF-kB p65antigen affinity purified polyclonal antibody (A00284-1) at 1:2000 overnight at 4°C, then washed with TBS-0.1%Tween 3 times with 5 minutes each and probed with a goat anti-rabbit IgG-HRP secondary antibody (Catalog # BA1054) at a dilution of 1:5000 for 1.5 hour at RT. The signal is developed using an ECL Plus Western Blotting Substrate (Catalog # AR1196-200) with Tanon 5200 system. A specific band was detected for NF-kB p65 at approximately 70 kDa. The expected band size for NF-kB p65 is at 70 kDa.
Click image to see more details
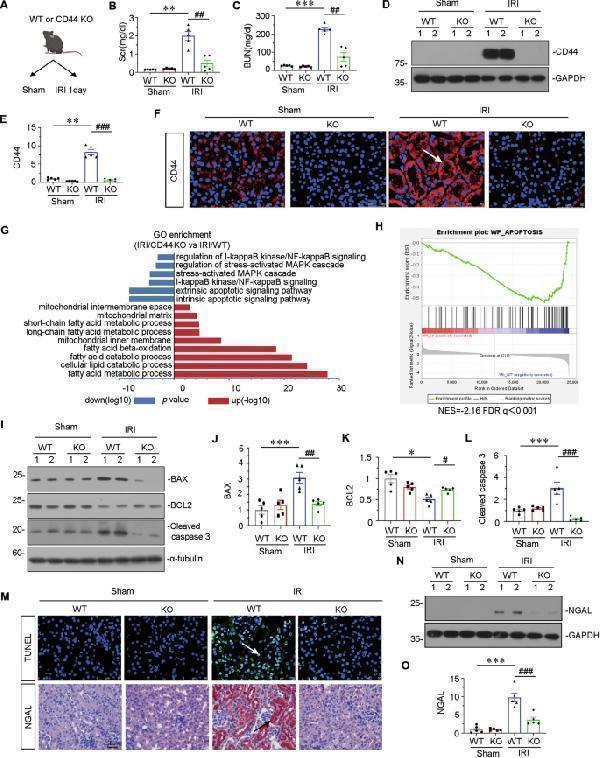
Gene ablation of CD44 attenuates renal tubular cell apoptosis and kidney injury upon IRI. A Experimental design: Wild-type mice and CD44 conventional knockout mice were subjected to IRI or sham, respectively, and euthanized 24 h after IRI. B Scr levels in four groups, as indicated. Scr was expressed as milligrams per deciliter. ** P < 0.01 versus wild-type mice upon sham group; ## P < 0.01 versus wild-type mice upon IRI group ( n = 5). C BUN levels in four groups, as indicated. BUN was expressed as milligrams per deciliter. *** P < 0.001 versus wild-type mice upon sham group; ## P < 0.01 versus wild-type mice upon IRI group ( n = 5). D and E Representative western blot of CD44 ( D ) and graphical presentations ( E ) of protein expressional levels are shown. ** P < 0.01 versus wild-type mice upon sham group; ### P < 0.001 versus wild-type mice upon IRI group ( n = 5). F Representative micrographs show the expression of CD44 in different groups, as indicated. Frozen kidney sections were stained with an antibody against CD44. Arrow indicates positive staining. Scale bar, 50 μm. G GO analysis shows CD44 is involved with several important pathways, including MAPK, NF-κB, apoptosis, mitochondria and FAO. H GSEA shows that negative regulation of apoptosis was enriched in CD44 knockout mice versus wild-type mice upon IRI. NES, normalized enrichment score; FDR q-value < 0.25. I–L Representative western blot ( I ) and graphical presentations of J BAX, K BCL2, and L cleaved caspase 3 protein expressional levels are shown. * P < 0.05, *** P < 0.001 versus wild-type mice upon sham group; # P < 0.05, ## P < 0.01, ### P < 0.001 versus wild-type mice upon IRI group ( n = 5). M Representative micrographs show TUNEL assay in different groups, as indicated. Frozen kidney sections were subjected to TUNEL assay. Parrafin sections were performed by immunohistochemistry staining of NGAL. Arrow indicates positive staining. Scale bar, 50 μm. N and O Representative western blot of NGAL ( N ) and graphical presentations ( O ) of protein expressional levels are shown. *** P < 0.001 versus wild-type mice upon sham group; ### P < 0.001 versus wild-type mice upon IRI group ( n = 5).
Index in PubMed under a CC BY license. PMID: 39979265
Click image to see more details
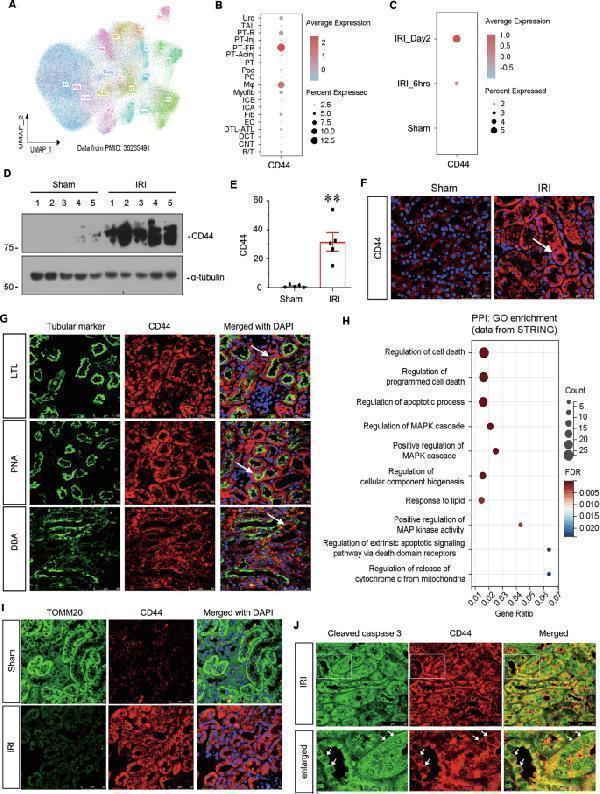
CD44 is upregulated in TECs and associated with mitochondrial dysfunction and apoptosis. A UMAP shows cell population in kidneys of sham and IRI at 6 h and day 2. PT proximal tubule, PT-Inj injured PT, PT-R repairing PT, FR-PTC failed repair PT cell, PT-AcInj acute injury PT, DTL descending limb of loop of Henle (LoH), ATL thin ascending limb of LoH, TAL thick ascending limb of LoH, DCT distal convoluted tubule, CNT connecting tubule, PC principal cell of collecting duct, ICA type A intercalated cell of collecting duct, ICB type B intercalated cell of collecting duct, Pod podocyte, EC endothelial cell, Fib fibroblast, Myofib myofibroblast, Ma macrophage (Mφ), B/T immune cell, Uro urothelium. Data from PMID: 36265491. B Graphic presentation of single-cell sequencing analysis shows the expression of CD44 in different cell populations. C Graphic presentation of single-cell sequencing analysis shows the expression of CD44 at different time point. D and E Representative western blot of CD44 ( D ) and graphical presentations ( E ) of protein expressional levels are shown. ** P < 0.01 versus sham group ( n = 5). F Representative micrographs show the expression of CD44 in sham and IRI groups, as indicated. Frozen kidney sections were stained with an antibody against CD44. Arrow indicates positive staining. Scale bar, 50 μm. G Co-localization staining of CD44 and various segment-specific tubular markers in the kidneys of the IRI model. Frozen kidney sections were collected from the mice 1 day after IRI. CD44 (red) and various segment-specific tubular markers (green), including LTL, PNA, and DBA, were detected by immunofluorescence. Arrows indicate positive staining. Scale bar, 50 μm. H GO analysis of CD44-related pathway through STRING ( ). I Co-localization of CD44 and TOMM20 in tubules. Frozen renal sections were subjected to immunostaining of CD44 (red) and TOMM20 (green). Scale bar, 50 μm. J Co-localization of CD44 and cleaved caspase 3 in IRI group. Frozen kidney sections were subjected to immunostaining of CD44 (red) and cleaved caspase 3 (green). Scale bar, 50 μm.
Index in PubMed under a CC BY license. PMID: 39979265
Click image to see more details
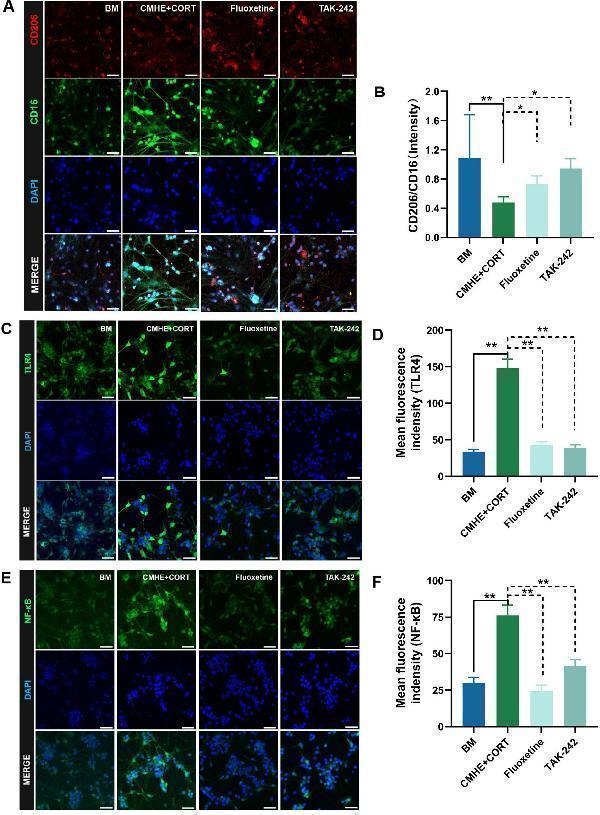
Activation of TLR4/NF-κB signaling and increased microglial M1-type polarization induces neuronal damage in the tri-culture model. (A) Fluorescence images of CD206 amd CD16 proteins in the microglia in the different groups. (B) Ratio of the relative fluorescence intensities between CD206 amd CD16 proteins in microglia. (C) Fluorescence images of TLR4 protein in neurons in the different groups. (D) Changes in fluorescence intensity of TLR4 protein in neurons in the different groups. (E) Fluorescence images of NF-κB protein in neurons in the different groups. (F) Changes in fluorescence intensity of NF-κB in neurons in the different groups. Data are presented as the mean ± SD ( n = 3). * P < 0.05, ** P < 0.01 vs. BM group (one-way ANOVA followed by LSD or Tamhane’s T2). Scale bar = 50 μm. BM: blank media; CHME: conditioned media of hypertensive environment; CORT: corticosterone; TLR4: toll-like receptor 4; NF-κB: nuclear factor-κB.
Index in PubMed under a CC BY license. PMID: 40230380
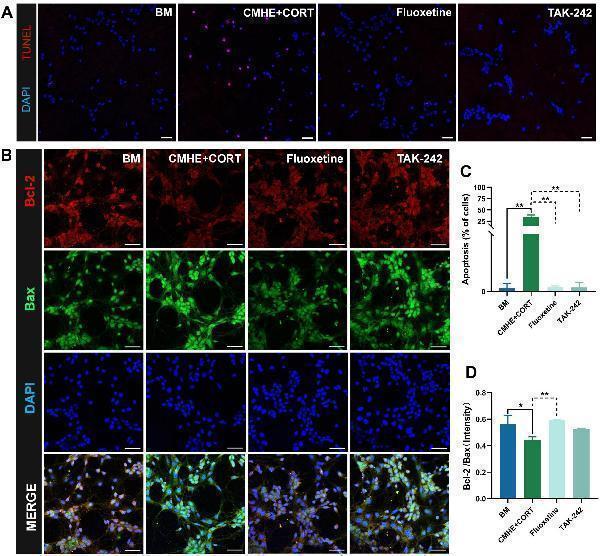
Click image to see more details
TLR4 inhibitors suppress neuronal apoptosis. (A) Fluorescence images of TUNEL staining in the different groups (red, apoptotic cells; blue, nucleus). (B) The number of apoptotic cells in the different groups. (C) Fluorescence images of apoptosis-related proteins Bcl-2 amd Bax in the different groups. (D) Ratio of the relative fluorescence intensity between Bcl-2 and Bax proteins in the different groups. Data are presented as the mean ± SD ( n = 3). * P < 0.05, ** P < 0.01 vs. BM group (one-way ANOVA followed by LSD or Tamhane’s T2). Scale bar = 50 μm. BM: blank media; CHME: conditioned media of hypertensive environment; CORT: corticosterone; Bcl-2: B-Cell Lymphoma 2; Bax: Bcl 2-Associated X.
Index in PubMed under a CC BY license. PMID: 40230380

Click image to see more details
Effects of HMGB1 down-regulation on NF-κB/p65, IκBα and VEGF-C in T24 cells. A1, A2 and A3: The expression of NF-κB/p65 and VEGF-C in the shRNA group was lower than in the other two groups transfected with shNC plasmids or the untransfected controls (all P < 0.05). On the contrary, the expression of IκBα showed an opposite tendency, while no significant differences in the expression of NF-κB/p65, IκBα and VEGF-C in T24 cells were found between the CON and NC groups (all P > 0.05). The display of cropped gels is used to improve the clarity and conciseness of the presentation and all the cropped gels have been run under the same experimental conditions. : The blue areas indicate nuclei stained using 4, 6-diamidino-2-phenylindole (DAPI) and the green areas indicate the nuclear translocation of NF-κB/p65 in T24 cells transfected with shNC plasmids or untransfected and cytoplasmic localization of NF-κB/p65 in cells transfected with shRNA plasmids. The results showed that knockdown of HMGB1 expression inhibited the translocation of NF-κB/p65 from the cytoplasm to the nucleus. : EMSA revealed that the DNA-binding activity of NF-κB/p65 in T24 cells was decreased by HMGB1 knockdown.
Index in PubMed under a CC BY license. PMID: 26239046
Click image to see more details
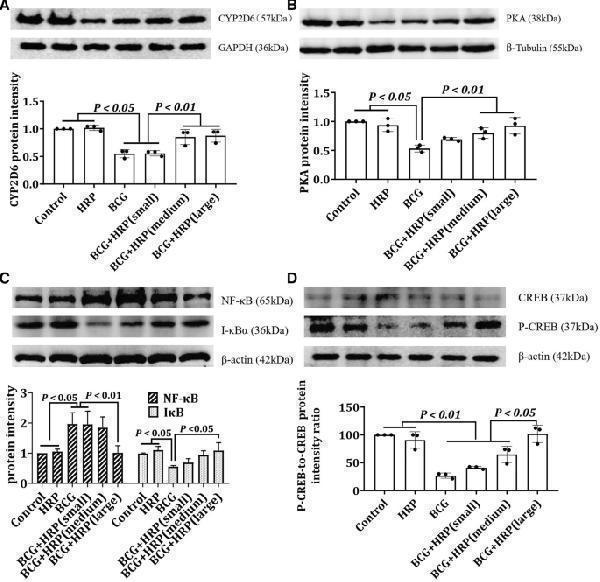
Effect of HRP on the expression of CYP2D6, PKA, CREB, PCREB, IκB, and NF-κB in rats with BCG-induced immune-mediated liver injury. Rats were administered BCG (single intravenous dose of 125 mg/kg BCG) or BCG + HRP (50, 100, or 200 mg ⋅ kg −1 ⋅ d −1 orally for 7 d). Liver proteins were extracted to determine the expression levels of CYP2D6, PKA, CREB, PCREB, IκB, and NF-κB. SDS-PAGE was performed using equal amounts (30 μg) of protein, and western blotting was performed using antibodies against CYP2D6, PKA, CREB, PCREB, i-κB, and NF-κB. The results were normalized to tubulin, GAPDH, or β-actin. The protein expression levels of CYP2D6 ( a ), PKA ( b ), IκB and NF-κB ( c ), CREB, and P-CREB ( d ) in the rat liver were measured by western blotting. The expression levels of CYP2D6, PKA, IκB, NF-κB, CREB, and P-CREB were quantified using the ImageJ software (NIH, Maryland, USA). The data represent the mean ± standard deviation (SD) of three independent experiments. BCG Bacille Calmette-Guerin; HRP Hippophae rhamnoides.
Index in PubMed under a CC BY license. PMID: 37833431
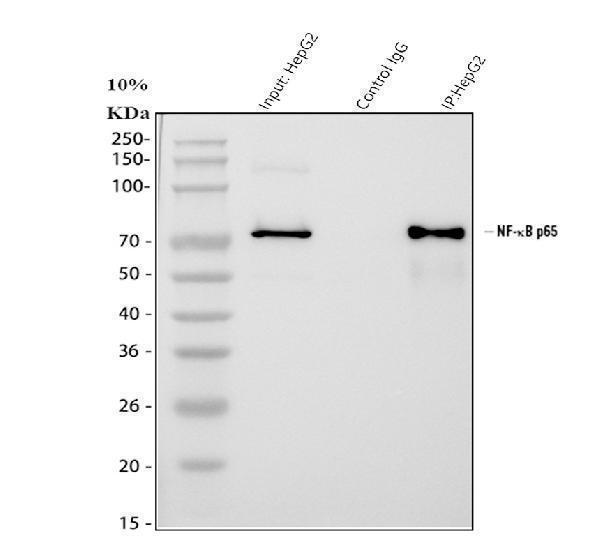
Click image to see more details
Immunoprecipitating NF-kB p65 in HepG2 whole cell lysate.
Western blot analysis of NF-kB p65 using anti-NF-kB p65 antibody (A00284-1).
Lane 1: HepG2 whole cell lysates (30ug),
Lane 2: Rabbit control IgG instead of anti-NF-kB p65 antibody in HepG2 whole cell lysate,
Lane 3: anti-NF-kB p65 antibody (2μg) + HepG2 whole cell lysate (500μg).
After electrophoresis, proteins were transferred to a membrane. Then the membrane was incubated with rabbit anti-NF-kB p65 antigen affinity purified polyclonal antibody (A00284-1) at a dilution of 0.5 μg/mL and probed with a goat anti-rabbit IgG-HRP secondary antibody (Catalog # BA1054). The signal is developed using ECL Plus Western Blotting Substrate (Catalog # AR1196-200). A specific band was detected for NF-kB p65 at approximately 65-70 kDa. The expected band size for NF-kB p65 is at 60 kDa.
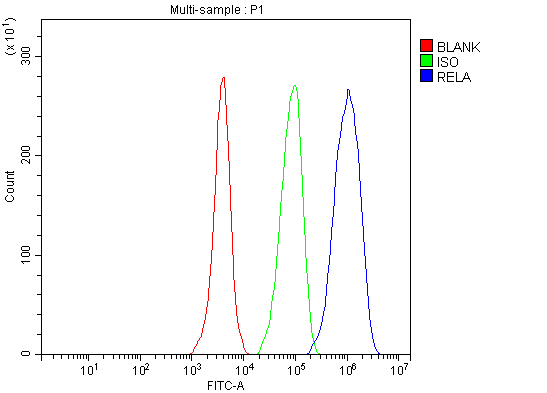
Click image to see more details
Flow Cytometry analysis of Hela cells using anti-NF-kB p65 antibody (A00284-1).
Overlay histogram showing Hela cells stained with A00284-1 (Blue line). To facilitate intracellular staining, cells were fixed with 4% paraformaldehyde and permeabilized with permeabilization buffer. The cells were blocked with 10% normal goat serum. And then incubated with rabbit anti-NF-kB p65 Antibody (A00284-1, 1 μg/1x106 cells) for 30 min at 20°C. DyLight®488 conjugated goat anti-rabbit IgG (BA1127, 5-10 μg/1x106 cells) was used as secondary antibody for 30 minutes at 20°C. Isotype control antibody (Green line) was rabbit IgG (1 μg/1x106) used under the same conditions. Unlabelled sample without incubation with primary antibody and secondary antibody (Red line) was used as a blank control.
Click image to see more details
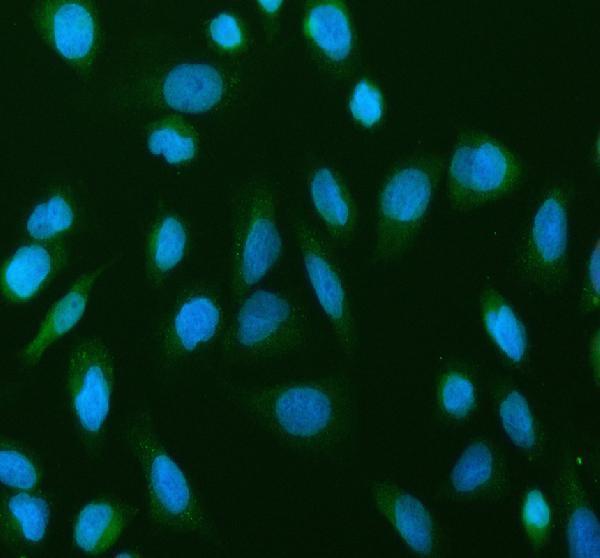
IF analysis of NF-kB p65 using anti-NF-kB p65 antibody (A00284-1).
NF-kB p65 was detected in an immunocytochemical section of U2OS cells. Enzyme antigen retrieval was performed using IHC enzyme antigen retrieval reagent (AR0022) for 15 mins. The cells were blocked with 10% goat serum. And then incubated with 5 μg/mL rabbit anti-NF-kB p65 Antibody (A00284-1) overnight at 4°C. DyLight®488 Conjugated Goat Anti-Rabbit IgG (BA1127) was used as secondary antibody at 1:500 dilution and incubated for 30 minutes at 37°C. The section was counterstained with DAPI. Visualize using a fluorescence microscope and filter sets appropriate for the label used.
Click image to see more details
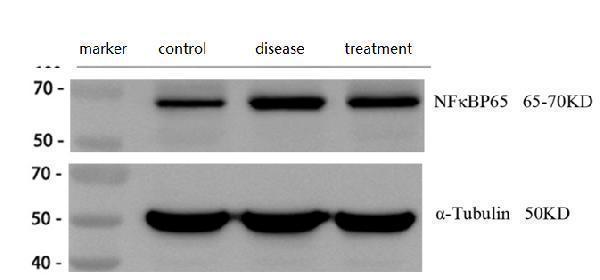
Western blot analysis of NF-kB p65 using anti-NF-kB p65 antibody (A00284-1).
Electrophoresis was performed on a 5-20% SDS-PAGE gel at 80V (Stacking gel) / 120V (Resolving gel) for 2 hours. The sample well of each lane was loaded with 30 ug of sample under reducing conditions.
Lane 1: control group-normal mouse hippocampal tissue lysates,
Lane 2: hippocampal tissue from Alzheimer’s disease model mouse,
Lane 3: hippocampal tissue from Alzheimer’s disease model mouse treated with a self-developed drug.
After electrophoresis, proteins were transferred to a nitrocellulose membrane at 150 mA for 50-90 minutes. Blocked the membrane with 5% non-fat milk/TBS for 1.5 hour at RT. The membrane was incubated with rabbit anti-NF-kB p65antigen affinity purified polyclonal antibody (A00284-1) at 1:2000 overnight at 4°C, then washed with TBS-0.1%Tween 3 times with 5 minutes each and probed with a goat anti-rabbit IgG-HRP secondary antibody (Catalog # BA1054) at a dilution of 1:10000 for 1 hour at RT. The signal is developed using an ECL Plus Western Blotting Substrate (Catalog # AR1196-200) with ChemiDoc MP system. A specific band was detected for NF-kB p65 at approximately 65-70 kDa. The expected band size for NF-kB p65 is at 70 kDa.
Specific Publications For Anti-NF-kB p65/RELA Antibody Picoband® (A00284-1)
Loading publications
Recommended Resources
Here are featured tools and databases that you might find useful.
- Boster's Pathways Library
- Protein Databases
- Bioscience Research Protocol Resources
- Data Processing & Analysis Software
- Photo Editing Software
- Scientific Literature Resources
- Research Paper Management Tools
- Molecular Biology Software
- Primer Design Tools
- Bioinformatics Tools
- Phylogenetic Tree Analysis
Customer Reviews
Have you used Anti-NF-kB p65/RELA Antibody Picoband®?
Share your experimental results or join a short interview to earn up to $1,000 in product credits or other rewards.
2 Reviews For Anti-NF-kB p65/RELA Antibody Picoband®
P65 Antibody (A00284-1) shows clear, specific bands with low background in mouse brain tissues by WB, with elevated expression in Alzheimer’s disease models and reduced levels after treatment, consistent with the expected results.
Excellent
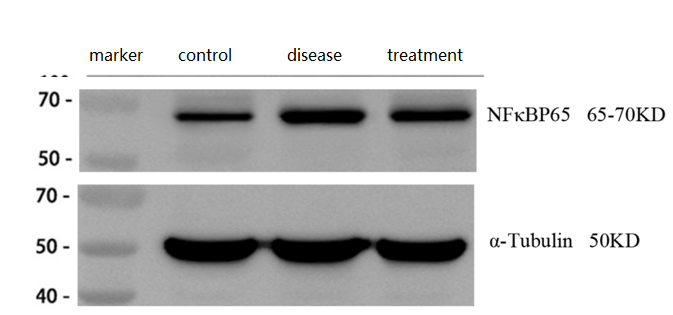
| SKU | A00284-1 |
|---|---|
| Application | Western Blot |
| Sample | mouse brain tissue |
| Sample Processing Description | ① Normal mouse hippocampal tissue, ② Hippocampal tissue from Alzheimer’s disease model mouse, ③ Hippocampal tissue from Alzheimer’s disease model mouse treated with a self-developed drug. Total protein was extracted from all samples. |
| Other Reagents | RIPA lysis buffer, Protease inhibitor, Running buffer, Transfer buffer, Blocking buffer |
| Primary Antibody | NF-kB p65/RELA Antibody Picoband® |
| Primary Incubation | 1:2000, overnight at 4 ℃ |
| Secondary Antibody | HRP Conjugated AffiniPure Goat Anti-Rabbit IgG (H+L) (BA1054) |
| Secondary Incubation | 1:10000, 1 h in RT |
| Detection | Substrate: ECL substrate, Image system: ChemiDoc MP |
| Results Summary | NF-κB p65 (RelA) is one of the most classical and essential subunits of the NF-κB transcription factor family. In Alzheimer’s disease, activation of p65 can initiate neuroprotective programs in neurons. The results showed that p65 expression was significantly increased in the brains of Alzheimer’s disease model mice and decreased after treatment, consistent with the expected outcomes. |
Huili Yin, Shandong First Medical University
Verified customer
Submitted 2026-03-27
This antibody is highly efficient and specific, with virtually no non-specific bands.
Excellent

| SKU | A00284-1 |
|---|---|
| Application | Immunofluorenscence |
| Sample | Mouse cell climbing slides |
| Sample Processing Description | Cells were directly lysed in NP40 lysis buffer, mixed with loading buffer at the appropriate ratio, and denatured by heating at 98°C. The samples were then loaded onto SDS-PAGE, with 20 μl of cell protein sample per lane. |
| Primary Antibody | Anti-NF-kB p65/RELA Antibody Picoband® |
| Primary Incubation | 1:2000, overnight at 4 ℃ |
| Blocking Agent | 5% Non-fat milk |
| Secondary Antibody | HRP-conjugated Goat Anti-Rabbit IgG |
| Secondary Incubation | Incubate at room temperature for 1 hour |
| Detection | Imaging system: Tanon, ECL substrate |
| Results Summary | I will purchase Boster products again and recommend them to my classmates and colleagues. |
Guangyu Mei, Tongji University
Verified customer
Submitted 2025-10-17
Customer Q&As
Have a question?
Find answers in Q&As, reviews.
Can't find your answer?
Submit your question
1 Customer Q&As for Anti-NF-kB p65/RELA Antibody Picoband®
Question
Can A00284-1 be used in both membrane type pvdf and nitrocellulose?
Verified customer
Asked: 2021-02-03
Answer
The Anti-NF-KB P65/RELA Antibody Picoband™ (A00284-1) is okay to use for pvdf or nitrocellulose membrane.
Boster Scientific Support
Answered: 2021-02-03





