Product Info Summary
| SKU: | M02690 |
|---|---|
| Size: | 100 μg/vial |
| Reactive Species: | Human |
| Host: | Mouse |
| Application: | Flow Cytometry, IHC, WB |
Customers Who Bought This Also Bought
Product info
Product Name
Anti-PPT1 Antibody Picoband® (monoclonal, 10F3)
SKU/Catalog Number
M02690
Size
100 μg/vial
Form
Lyophilized
Description
Boster Bio Anti-PPT1 Antibody Picoband® (monoclonal, 10F3) catalog # M02690. Tested in Flow Cytometry, IHC, WB applications. This antibody reacts with Human. The brand Picoband indicates this is a premium antibody that guarantees superior quality, high affinity, and strong signals with minimal background in Western blot applications. Only our best-performing antibodies are designated as Picoband, ensuring unmatched performance.
Storage & Handling
Store at -20˚C for one year from date of receipt. After reconstitution, at 4˚C for one month. It can also be aliquotted and stored frozen at -20˚C for six months. Avoid repeated freeze-thaw cycles.
Cite This Product
Anti-PPT1 Antibody Picoband® (monoclonal, 10F3) (Boster Biological Technology, Pleasanton CA, USA, Catalog # M02690)
Host
Mouse
Contents
Each vial contains 4mg Trehalose, 0.9mg NaCl, 0.2mg Na2HPO4, 0.05mg NaN3.
Clonality
Monoclonal
Clone Number
10F3
Isotype
Mouse IgG2b
Immunogen
A synthetic peptide corresponding to a sequence at the C-terminus of human PPT1, different from the related mouse and rat sequences by four amino acids.
Cross-reactivity
No cross-reactivity with other proteins.
Reactive Species
M02690 is reactive to PPT1 in Human
Observed Molecular Weight
34 kDa
Calculated molecular weight
34.2 kDa
Background of PPT1
Palmitoyl-protein thioesterase 1 (PPT-1), also known as palmitoyl-protein hydrolase 1, is an enzyme that in humans is encoded by the PPT1 gene. PPT-1 is a member of the palmitoyl protein thioesterase family. The protein encoded by this gene is a small glycoprotein involved in the catabolism of lipid-modified proteins during lysosomal degradation. The encoded enzyme removes thioester-linked fatty acyl groups such as palmitate from cysteine residues. Defects in this gene are a cause of infantile neuronal ceroid lipofuscinosis 1 (CLN1, or INCL) and neuronal ceroid lipofuscinosis 4 (CLN4). Two transcript variants encoding different isoforms have been found for this gene.
Antibody Validation
Boster validates all antibodies on WB, IHC, ICC, Immunofluorescence, and ELISA with known positive control and negative samples to ensure specificity and high affinity, including thorough antibody incubations.
Application & Images
Applications
M02690 is guaranteed for Flow Cytometry, IHC, WB Boster Guarantee
Assay Dilutions Recommendation
The recommendations below provide a starting point for assay optimization. The actual working concentration varies and should be decided by the user.
Western blot, 0.1-0.5μg/ml
Immunohistochemistry (Paraffin-embedded Section), 0.5-1μg/ml
Flow Cytometry (Fixed), 1-3μg/1x106 cells
Positive Control
Suggested blocking solution with 5% non-fat milk or BSA; (*)Recommended protein loading: 20-40 µg per lane
Use TE buffer pH 9.0 for antigen retrieval; (*) citrate buffer pH 6.0 is an alternative.
Validation Images & Assay Conditions

Click image to see more details
Western blot analysis of PPT1 using anti-PPT1 antibody (M02690).
Electrophoresis was performed on a 5-20% SDS-PAGE gel at 70V (Stacking gel) / 90V (Resolving gel) for 2-3 hours. The sample well of each lane was loaded with 50ug of sample under reducing conditions.
Lane 1: human K562 whole cell lysates,
Lane 2: human HepG2 whole cell lysates,
Lane 3: human Hela whole cell lysates.
After Electrophoresis, proteins were transferred to a Nitrocellulose membrane at 150mA for 50-90 minutes. Blocked the membrane with 5% Non-fat Milk/ TBS for 1.5 hour at RT. The membrane was incubated with mouse anti-PPT1 antigen affinity purified monoclonal antibody (Catalog # M02690) at 0.5 μg/mL overnight at 4°C, then washed with TBS-0.1%Tween 3 times with 5 minutes each and probed with a goat anti-mouse IgG-HRP secondary antibody at a dilution of 1:10000 for 1.5 hour at RT. The signal is developed using an Enhanced Chemiluminescent detection (ECL) kit (Catalog # EK1001) with Tanon 5200 system. A specific band was detected for PPT1 at approximately 34KD. The expected band size for PPT1 is at 34KD.
Click image to see more details
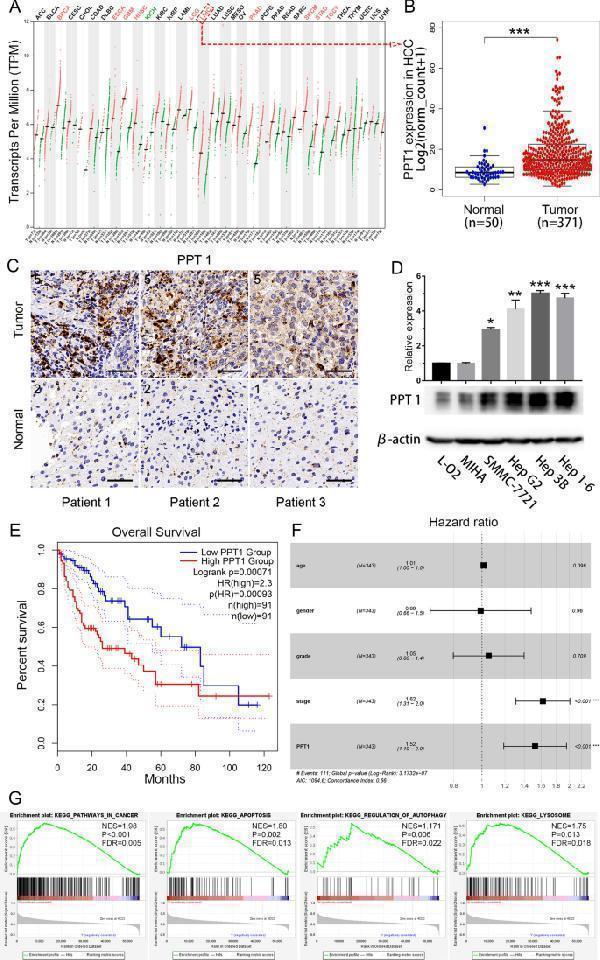
Role of PPT1 in HCC. A PPT1 mRNA expression in various normal human tissues and tumor tissues. B TCGA database-based comparison of PPT1 mRNA expression in HCC tissues (n = 371) and normal liver tissues (n = 50); *** P < 0.001. C Representative immunohistochemical results of PPT1 in patient-derived HCC tissue and normal liver tissue. Scale bar, 50 μm. D Expression of PPT1 in a panel of normal liver cell lines (L-O2 and MIHA) and HCC cell lines. On Western blot analysis, PPT1 was found to be preferentially expressed in HCC cell lines, Hep 3B and Hep 1-6. E Kaplan–Meier curves of overall survival (OS). High PPT1 expression was correlated with poor OS in HCC patients. F Cox proportional hazards regression model analysis of OS. G Gene set enrichment analysis (GSEA) using the TCGA dataset. PPT1 overexpression was significantly correlated with “KEGG_PATHWAYS_IN_CANCER”, “KEGG_APOPTOSIS”, “KEGG_REGULATION_OF_AUTOPHAGY”, and “KEGG_LYSOSOME” pathways. NES: normalized enrichmentscore
Index in PubMed under a CC BY license. PMID: 35277179
Click image to see more details
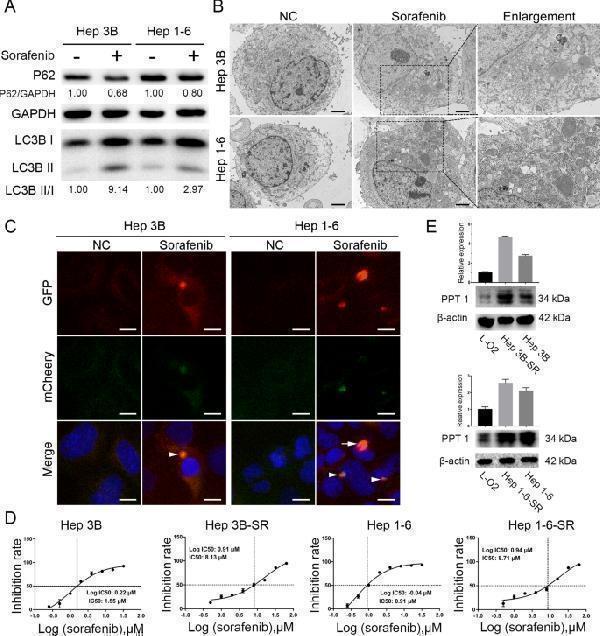
Sorafenib induced autophagy and upregulated the expression of PPT1 in sorafenib-resistant HCC cells. A Western blot showing increase in LC3-II in HCC cells after treatment with sorafenib (10 µΜ, 24 h). B Photographs from transmission electron microscopy showing an increase in autophagosomes in HCC cells after treatment with sorafenib (10 µΜ, 24 h). Scale bar, 2 μm. C Fluorescence microscopic images showing punctate fluorescence from transfected mCherry-GFP-LC3 constructs in HCC cells treated with sorafenib (10 µΜ, 24 h); nuclei are labeled with Hoechst 33,258. Arrowheads indicate typical examples of co-localized particles of GFP and mCherry signal, while the arrow points to a typical example of a particle with an mCherry signal but without GFP signal. Scale bar, 10 μm. D IC50 values of sorafenib (48 h) for Hep 3B, Hep 3B-SR, Hep 1 – 6, and Hep 1-6-SR cells determined by CCK-8 assay. The data shown are from three independent experiments. E Confirmation of the upregulation of PPT1 protein in sorafenib-resistant cells derived from Hep 3B and Hep 1 – 6 by Western blot analysis
Index in PubMed under a CC BY license. PMID: 35277179
Click image to see more details
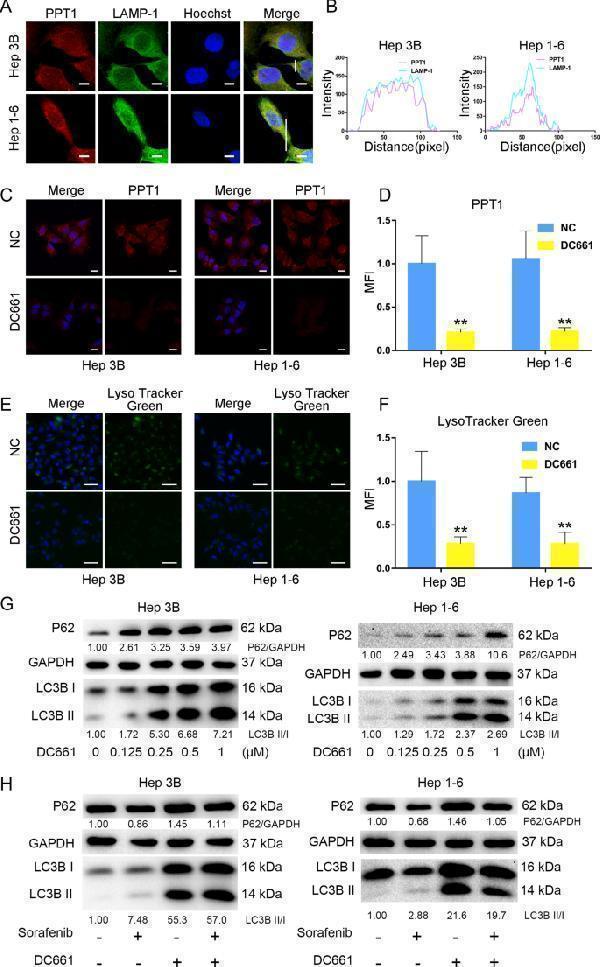
PPT1 inhibitor DC661 inhibited autophagy by inhibiting lysosomes. A Confocal laser-scanning microscopy images of intracellular co-localization between PPT1 and lysosomes. LAMP-1 staining indicates the lysosomes, and Hoechst 33,258 indicates the nucleus. Co-localization is visualized by color and area overlap (red + green = yellow). Scale bar, 10 μm. B Fluorescence co-localization analysis according to ( A ) was performed by ImageJ software. C Immunofluorescence staining of PPT1 in HCC cells after treatment with DC661 (3 µM, 6 h). Scale bar, 20 μm. D Semiquantitative analysis of the mean fluorescence intensity (MFI) in the HCC cells according to ( C ) was performed by using ImageJ software (n = 6). Data represent mean ± SD; ** P < 0.01. E Fluorescence images of LysoTracker Green (lysosome probe) in HCC cells after treatment with DC661 (3 µM, 6 h). Hoechst 33,258 was used to stain the nucleus. Scale bar, 50 μm. F Semiquantitative analysis of the MFI of LysoTracker Green in HCC cells according to ( E ) was performed by ImageJ software (n = 6). Data represent mean ± SD; ** P < 0.01. G Western blot showing an increase in LC3-II and P62 in HCC cells treated with DC661 (3 µM, 6 h). H Western blot showing P62 degradation and LC3 lipidation in HCC cells treated with sorafenib and/or DC661. HCC cells were treated with or without 10 µM sorafenib in the presence or absence of 1 µM DC661 for 24 h
Index in PubMed under a CC BY license. PMID: 35277179
Click image to see more details
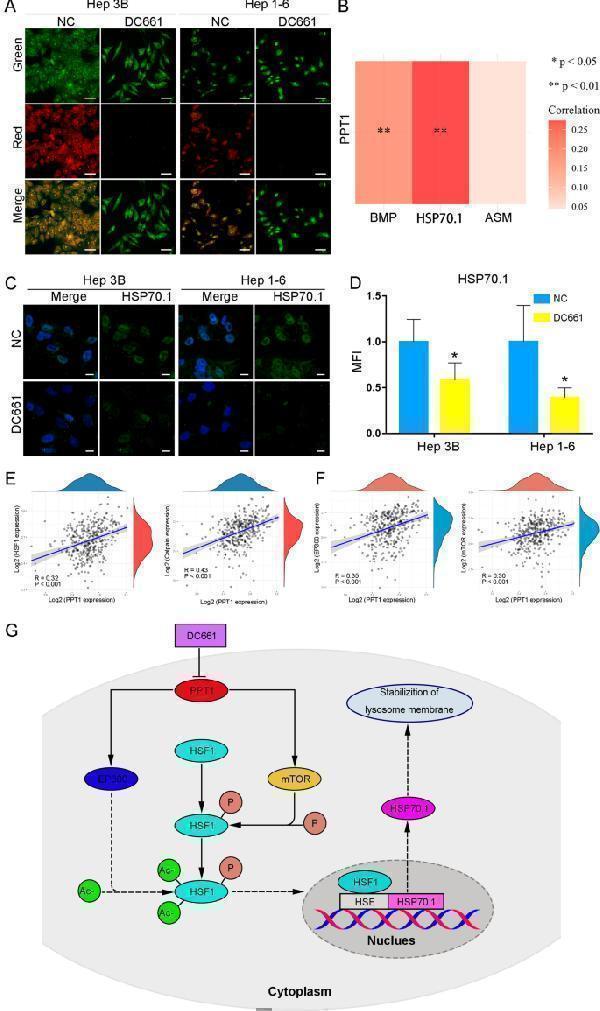
Potential molecular mechanism of lysosomal membrane hyperpermeability induced by PPT1 inhibitor. A Fluorescence images of acridine orange staining in HCC cells treated with DC661 (3 µM, 6 h). Hoechst 33,258 staining marks the nuclei. Scale bar, 50 μm. B Gene correlation analysis using TCGA dataset. Heat map of the correlation between multiple genes and one gene. PPT1 expression was positively correlated with HSP70.1 and BMP expression in HCC (linear regression); ** P < 0.01. C Immunofluorescent staining of HSP70.1 in HCC cells treated with DC661 (3 µM, 6 h). Scale bar, 10 μm. D Semiquantitative analysis of the mean fluorescence intensity (MFI) in HCC cells according to ( C ) was performed by using ImageJ software (n = 6). Data represent mean ± SD; * P < 0.05. E Gene correlation analysis using TCGA dataset. PPT1 expression was positively correlated with HSF1 and calpain expression in HCC (linear regression); P < 0.001. F Gene correlation analysis using TCGA dataset. PPT1 expression was positively correlated with EP300 and mTOR expression in HCC (linear regression); P < 0.001. G Schematic illustration of a potential molecular mechanism of lysosomal membrane hyperpermeability induced by PPT1 inhibitor. HSE: heat shock element
Index in PubMed under a CC BY license. PMID: 35277179
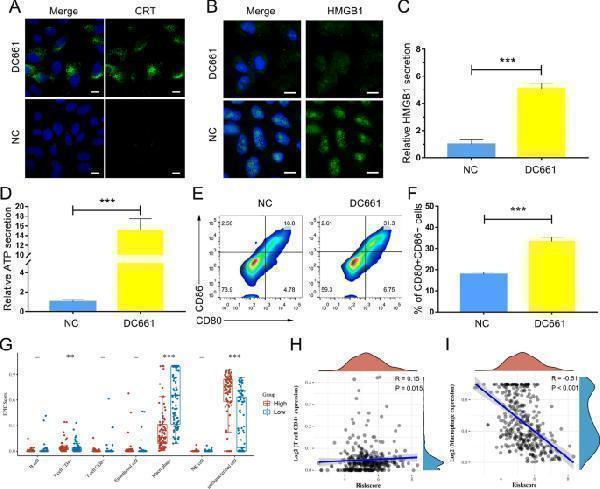
Click image to see more details
PPT1 inhibitor DC661-induced immunogenic cell death promoted the maturation of DCs and the activation of CD8 + T cells. A Immunofluorescent imaging of CRT expression on the cell surface of Hep 1-6 cells treated with DC661 (3 µM, 6 h). Nuclei are stained with Hoechst 33,258. Scale bars, 10 μm. B Immunofluorescent imaging of HMGB-1 release by Hep 1-6 cells treated with DC661 (3 µM, 6 h). Nuclei are stained with Hoechst 33,258. Scale bars, 10 μm. C ELISA detection of HMGB1 release into cell culture medium. Cells were incubated with DC661 at 3 µM for 6 h. Data represent mean ± SD; *** P < 0.001. D ELISA detection of ATP release into the cell culture medium. Cells were incubated with DC661 at 3 µM for 6 h. Data represent mean ± SD; *** P < 0.001. E , F Flow cytometric analysis (left) and quantification (right) of mature DC cells (CD11c + CD80 + CD86 + ) from the spleens of DC661- or vehicle-treated Hep 1-6 tumor-bearing mice. Data represent mean ± SD; *** P < 0.001. G–I The expression level of PPT1 was associated with the immune infiltration in the HCC tumor microenvironment. Scatter plots ( G ) and correlation diagrams ( H , I ) showing the difference of CD4 + T cells and macrophages infiltration level between PPT1-high and -low groups in TCGA-LIHC. ** P < 0.01; *** P < 0.001
Index in PubMed under a CC BY license. PMID: 35277179

Click image to see more details
IHC analysis of PPT1 using anti PPT1 antibody (M02690).
PPT1 was detected in paraffin-embedded section of human intestinal cancer tissue. Heat mediated antigen retrieval was performed in EDTA buffer (pH8.0, epitope retrieval solution). The tissue section was blocked with 10% goat serum. The tissue section was then incubated with 1μg/ml mouse anti-PPT1 Antibody (M02690) overnight at 4°C. Biotinylated goat anti-mouse IgG was used as secondary antibody and incubated for 30 minutes at 37°C. The tissue section was developed using Strepavidin-Biotin-Complex (SABC) (Catalog # SA1021) with DAB as the chromogen.

Click image to see more details
IHC analysis of PPT1 using anti PPT1 antibody (M02690).
PPT1 was detected in paraffin-embedded section of human lung cancer tissue. Heat mediated antigen retrieval was performed in EDTA buffer (pH8.0, epitope retrieval solution). The tissue section was blocked with 10% goat serum. The tissue section was then incubated with 1μg/ml mouse anti-PPT1 Antibody (M02690) overnight at 4°C. Biotinylated goat anti-mouse IgG was used as secondary antibody and incubated for 30 minutes at 37°C. The tissue section was developed using Strepavidin-Biotin-Complex (SABC) (Catalog # SA1021) with DAB as the chromogen.

Click image to see more details
IHC analysis of PPT1 using anti PPT1 antibody (M02690).
PPT1 was detected in paraffin-embedded section of human mammary cancer tissue. Heat mediated antigen retrieval was performed in EDTA buffer (pH8.0, epitope retrieval solution). The tissue section was blocked with 10% goat serum. The tissue section was then incubated with 1μg/ml mouse anti-PPT1 Antibody (M02690) overnight at 4°C. Biotinylated goat anti-mouse IgG was used as secondary antibody and incubated for 30 minutes at 37°C. The tissue section was developed using Strepavidin-Biotin-Complex (SABC) (Catalog # SA1021) with DAB as the chromogen.

Click image to see more details
Flow Cytometry analysis of THP-1 cells using anti- PPT1 antibody (M02690).
Overlay histogram showing THP-1 cells stained with M02690 (Blue line). To facilitate intracellular staining, cells were fixed with 4% paraformaldehyde and permeabilized with permeabilization buffer. The cells were blocked with 10% normal goat serum. And then incubated with mouse anti-PPT1 Antibody (M02690, 1μg/1x106 cells) for 30 min at 20°C. DyLight®488 conjugated goat anti-mouse IgG (BA1126, 5-10μg/1x106 cells) was used as secondary antibody for 30 minutes at 20°C. Isotype control antibody (Green line) was mouse IgG (1μg/1x106) used under the same conditions. Unlabelled sample without incubation with primary antibody and secondary antibody (Red line) was used as a blank control.
Specific Publications For Anti-PPT1 Antibody Picoband® (monoclonal, 10F3) (M02690)
Loading publications
Recommended Resources
Here are featured tools and databases that you might find useful.
- Boster's Pathways Library
- Protein Databases
- Bioscience Research Protocol Resources
- Data Processing & Analysis Software
- Photo Editing Software
- Scientific Literature Resources
- Research Paper Management Tools
- Molecular Biology Software
- Primer Design Tools
- Bioinformatics Tools
- Phylogenetic Tree Analysis
Customer Reviews
Have you used Anti-PPT1 Antibody Picoband® (monoclonal, 10F3)?
Share your experimental results or join a short interview to earn up to $1,000 in product credits or other rewards.
0 Reviews For Anti-PPT1 Antibody Picoband® (monoclonal, 10F3)
Customer Q&As
Have a question?
Find answers in Q&As, reviews.
Can't find your answer?
Submit your question





