Product Info Summary
| SKU: | PA1518 |
|---|---|
| Size: | 100 μg/vial |
| Reactive Species: | Mouse, Rat |
| Host: | Rabbit |
| Application: | Flow Cytometry, IF, IHC, WB |
Customers Who Bought This Also Bought
Product info
Product Name
Anti-Macrosialin CD68 Antibody Picoband®
SKU/Catalog Number
PA1518
BA3638 is an alternative SKU for this antibody, used in previous lots.
Size
100 μg/vial
Form
Lyophilized
Description
Superstar antibody: Boster’s anti mouse CD68 antibody (PA1518) is among its top 10 bestselling antibodies, and one of the most cited mouse/rat CD68 antibodies on the market. Because of this antibody’s high demand, we have subsequently developed a rabbit monoclonal anti CD68 antibody (M00602-1) targetting a similar epitope.
High specificity: PA1518 reacts with murine macrosialin/CD68, and has been validated with Western blotting and confirmed its stellar specificity. CD68 is a highly glycosylated protein, and its expected western blot molecular weight is between 80kDa to 110kDa, depending on glycosylation level. PA1518’s observed MW in WB is near 90-100kDa. Testing on negative control tissues showed no significant bands (images available on request) or staining. See more info in the positive and negative control design section. Based on immunogen sequence homology, this antibody is not expected to cross react with other proteins from the LAMP family, which complies with QC testing observations.
Great for CD68 IHC: This antibody produces clean and specific immunobiological stains in both mouse and rat tissues. Our QC team has validated it on spleen and liver tissues of both mice and rats. Click product images for more details on experiment conditions.
More about CD68: CD68 is a cell surface marker often used to identify macrophages and other cell types in the monocyte lineage. It is a transmembrane protein that binds to electins and selectins and plays a role in macrophage homing movement. Check out the CD68 biomarker page for more information on CD68 and view all CD68 antibodies, ELISA kits and proteins.
The brand Picoband indicates this is a premium antibody that guarantees superior quality, high affinity, and strong signals with minimal background in Western blot applications. Only our best-performing antibodies are designated as Picoband, ensuring unmatched performance. CD68 (macrosialin) is a heavily glycosylated glycoprotein enriched in macrophages/mononuclear phagocytes; described roles include tissue macrophage phagocytic activities, intracellular lysosomal metabolism, and extracellular cell–cell/cell–pathogen interactions, with lectin/selectin binding implicated in macrophage subset localization/homing. Assay context: validated for Flow Cytometry/IF/IHC/WB with an observed WB band near 90–100 kDa (glycosylation-dependent). Often profiled alongside T-cell marker CD4 and cytotoxic effector GZMB to contextualize immune infiltrates; for tissue macrophage quantification and co-localization, multiplex IHC can be used with downstream pathology review to interpret staining patterns in histologic context.Storage & Handling
Store at -20˚C for one year from date of receipt. After reconstitution, at 4˚C for one month. It can also be aliquotted and stored frozen at -20˚C for six months. Avoid repeated freeze-thaw cycles.
Cite This Product
Anti-Macrosialin CD68 Antibody Picoband® (Boster Biological Technology, Pleasanton CA, USA, Catalog # PA1518)
Host
Rabbit
Contents
Each vial contains 4mg Trehalose, 0.9mg NaCl and 0.2mg Na2HPO4.
Clonality
Polyclonal
Isotype
Rabbit IgG
Immunogen
A synthetic peptide corresponding to a sequence in the middle region of mouse CD68, different from the related rat sequence by one amino acid.
Cross-reactivity
No cross-reactivity with other proteins
Reactive Species
PA1518 is reactive to CD68 in Mouse, Rat
Observed Molecular Weight
90-100 kDa
Calculated molecular weight
34.8 kDa
Background of CD68
CD68 is a heavily glycosylated glycoprotein that is highly expressed in macrophages and other mononuclear phagocytes. Macrosialin, if not glycosylated, has the main sequence that consists of 354 amino acids with an expected molecular weight of 37.4 kD. CD68, which can be located in the cytoplasmic granules of a variety of blood cells and myocytes, can be detected via immunohistochemistry. It's particularly effective as a marker for monocytes, histiocytes, giant cells, Kupffer cells, and osteoclasts, all of which belong to the macrophage lineage. CD68 is expressed in tissue macrophages and to a lesser extent in dendritic cells. CD68 is involved in tissue macrophage phagocytic activities, as well as intracellular lysosomal metabolism and extracellular cell-cell and cell-pathogen interactions. It works by binding to tissue and organ-specific lectins or selectins, enabling macrophage subsets to be directed to specific locations. Macrophages may be able to crawl over selectin-bearing substrates or other cells if CD68 is rapidly recirculated from endosomes and lysosomes to the plasma membrane.
Antibody Validation
Boster validates all antibodies on WB, IHC, ICC, Immunofluorescence, and ELISA with known positive control and negative samples to ensure specificity and high affinity, including thorough antibody incubations.
Application & Images
Applications
PA1518 is guaranteed for Flow Cytometry, IF, IHC, WB Boster Guarantee
Recommend Dilution
| Application | Dilution | Species |
|---|---|---|
| Western blot | 0.1-0.5μg/ml | Mouse, Rat |
| Immunohistochemistry (Paraffin-embedded Section) | 2-5μg/ml | Mouse, Rat |
| Immunofluorescence | 5μg/ml | Mouse |
| Flow Cytometry(Fixed) | 1-3 μg/1x106 cells | Mouse |
Tested application
Suggested blocking solution with 5% non-fat milk or BSA; (*)Recommended protein loading: 20-40 µg per lane
Use TE buffer pH 9.0 for antigen retrieval; (*) citrate buffer pH 6.0 is an alternative.
Validation Images & Assay Conditions

Click image to see more details
Western blot analysis of CD68 using anti-CD68 antibody (PA1518).
Electrophoresis was performed on a 5-20% SDS-PAGE gel at 70V (Stacking gel) / 90V (Resolving gel) for 2-3 hours. The sample well of each lane was loaded with 30 ug of sample under reducing conditions.
Lane 1: rat spleen tissue lysates,
Lane 2: mouse spleen tissue lysates,
Lane 3: mouse RAW264.7 whole cell lysates.
After electrophoresis, proteins were transferred to a nitrocellulose membrane at 150 mA for 50-90 minutes. Blocked the membrane with 5% non-fat milk/TBS for 1.5 hour at RT. The membrane was incubated with rabbit anti-CD68 antigen affinity purified polyclonal antibody (Catalog # PA1518) at 0.5 μg/mL overnight at 4°C, then washed with TBS-0.1%Tween 3 times with 5 minutes each and probed with a goat anti-rabbit IgG-HRP secondary antibody at a dilution of 1:5000 for 1.5 hour at RT. The signal is developed using an Enhanced Chemiluminescent detection (ECL) kit (Catalog # EK1002) with Tanon 5200 system. A specific band was detected for CD68 at approximately 90-100 kDa. The expected band size for CD68 is at 37 kDa.

Click image to see more details
IHC analysis of CD68 using anti-CD68 antibody (PA1518).
CD68 was detected in a paraffin-embedded section of mouse spleen tissue. Heat mediated antigen retrieval was performed in EDTA buffer (pH 8.0, epitope retrieval solution). The tissue section was blocked with 10% goat serum. The tissue section was then incubated with 2 μg/ml rabbit anti-CD68 Antibody (PA1518) overnight at 4°C. Peroxidase Conjugated Goat Anti-rabbit IgG was used as secondary antibody and incubated for 30 minutes at 37°C. The tissue section was developed using HRP Conjugated Rabbit IgG Super Vision Assay Kit (Catalog # SV0002) with DAB as the chromogen.

Click image to see more details
IHC analysis of CD68 using anti-CD68 antibody (PA1518).
CD68 was detected in a paraffin-embedded section of mouse liver tissue. Heat mediated antigen retrieval was performed in EDTA buffer (pH 8.0, epitope retrieval solution). The tissue section was blocked with 10% goat serum. The tissue section was then incubated with 2 μg/ml rabbit anti-CD68 Antibody (PA1518) overnight at 4°C. Peroxidase Conjugated Goat Anti-rabbit IgG was used as secondary antibody and incubated for 30 minutes at 37°C. The tissue section was developed using HRP Conjugated Rabbit IgG Super Vision Assay Kit (Catalog # SV0002) with DAB as the chromogen.

Click image to see more details
IHC analysis of CD68 using anti-CD68 antibody (PA1518).
CD68 was detected in a paraffin-embedded section of rat spleen tissue. Heat mediated antigen retrieval was performed in EDTA buffer (pH 8.0, epitope retrieval solution). The tissue section was blocked with 10% goat serum. The tissue section was then incubated with 2 μg/ml rabbit anti-CD68 Antibody (PA1518) overnight at 4°C. Peroxidase Conjugated Goat Anti-rabbit IgG was used as secondary antibody and incubated for 30 minutes at 37°C. The tissue section was developed using HRP Conjugated Rabbit IgG Super Vision Assay Kit (Catalog # SV0002) with DAB as the chromogen.

Click image to see more details
IHC analysis of CD68 using anti-CD68 antibody (PA1518).
CD68 was detected in a paraffin-embedded section of rat lung tissue. Heat mediated antigen retrieval was performed in EDTA buffer (pH 8.0, epitope retrieval solution). The tissue section was blocked with 10% goat serum. The tissue section was then incubated with 2 μg/ml rabbit anti-CD68 Antibody (PA1518) overnight at 4°C. Peroxidase Conjugated Goat Anti-rabbit IgG was used as secondary antibody and incubated for 30 minutes at 37°C. The tissue section was developed using HRP Conjugated Rabbit IgG Super Vision Assay Kit (Catalog # SV0002) with DAB as the chromogen.

Click image to see more details
IHC analysis of CD68 using anti-CD68 antibody (PA1518).
CD68 was detected in a paraffin-embedded section of rat lung tissue. Heat mediated antigen retrieval was performed in EDTA buffer (pH 8.0, epitope retrieval solution). The tissue section was blocked with 10% goat serum. The tissue section was then incubated with 2 μg/ml rabbit anti-CD68 Antibody (PA1518) overnight at 4°C. Peroxidase Conjugated Goat Anti-rabbit IgG was used as secondary antibody and incubated for 30 minutes at 37°C. The tissue section was developed using HRP Conjugated Rabbit IgG Super Vision Assay Kit (Catalog # SV0002) with DAB as the chromogen.

Click image to see more details
IHC analysis of CD68 using anti-CD68 antibody (PA1518).
CD68 was detected in a paraffin-embedded section of rat liver tissue. Heat mediated antigen retrieval was performed in EDTA buffer (pH 8.0, epitope retrieval solution). The tissue section was blocked with 10% goat serum. The tissue section was then incubated with 2 μg/ml rabbit anti-CD68 Antibody (PA1518) overnight at 4°C. Peroxidase Conjugated Goat Anti-rabbit IgG was used as secondary antibody and incubated for 30 minutes at 37°C. The tissue section was developed using HRP Conjugated Rabbit IgG Super Vision Assay Kit (Catalog # SV0002) with DAB as the chromogen.

Click image to see more details
IF analysis of CD68 using anti-CD68 antibody (PA1518).
CD68 was detected in a paraffin-embedded section of mouse spleen tissue. Heat mediated antigen retrieval was performed in EDTA buffer (pH 8.0, epitope retrieval solution). The tissue section was blocked with 10% goat serum. The tissue section was then incubated with 5 μg/mL rabbit anti-CD68 Antibody (PA1518) overnight at 4°C. DyLight®488 Conjugated Goat Anti-Rabbit IgG (BA1127)) was used as secondary antibody at 1:500 dilution and incubated for 30 minutes at 37°C. The section was counterstained with DAPI. Visualize using a fluorescence microscope and filter sets appropriate for the label used.

Click image to see more details
Flow Cytometry analysis of RAW264.7 cells using anti-CD68 antibody (PA1518).
Overlay histogram showing RAW264.7 cells stained with PA1518 (Blue line). The cells were fixed with 4% paraformaldehyde and blocked with 10% normal goat serum. And then incubated with rabbit anti-CD68 Antibody (PA1518, 1 μg/1x106 cells) for 30 min at 20°C. DyLight®488 conjugated goat anti-rabbit IgG (BA1127, 5-10 μg/1x106 cells) was used as secondary antibody for 30 minutes at 20°C. Isotype control antibody (Green line) was rabbit IgG (1 μg/1x106) used under the same conditions. Unlabelled sample without incubation with primary antibody and secondary antibody (Red line) was used as a blank control.
Click image to see more details
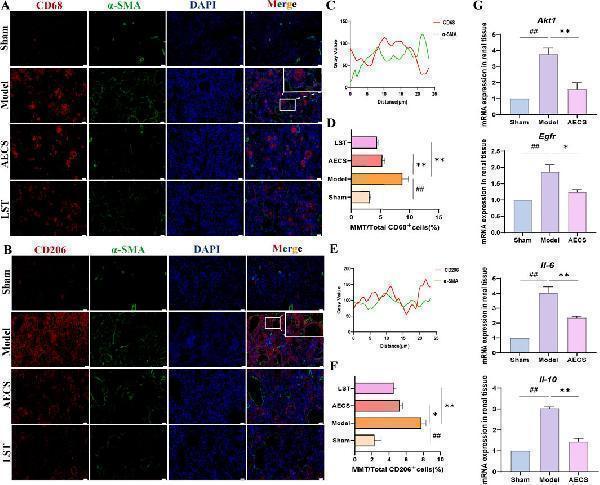
CS attenuates MMT in RIF rats. (A) Co-staining of the α-SMA (green) with CD68 (red) by Double Immunofluorescence. (B) Co-localization analysis of CD68+α-SMA + MMT cells. (C) Staining intensity of CD68+α-SMA + MMT cells was quantified. (D) Co-staining of the α-SMA (green) with CD206 (red) by Double Immunofluorescence. (E) Co-localization analysis of CD206+α-SMA + MMT cells. (F) Staining intensity of CD206+α-SMA + MMT cells was quantified (n = 4). (G) mRNA Expression of AKT1, EGFR, IL-6, and IL-10 in RIF rats kidney (n = 3). ## p < 0.01 vs. the sham group; * p < 0.05 vs. the model group; ** p < 0.01 vs. the model group. Scale bars (40 μm, 10 μm).
Index in PubMed under a CC BY license. PMID: 41142245
Click image to see more details
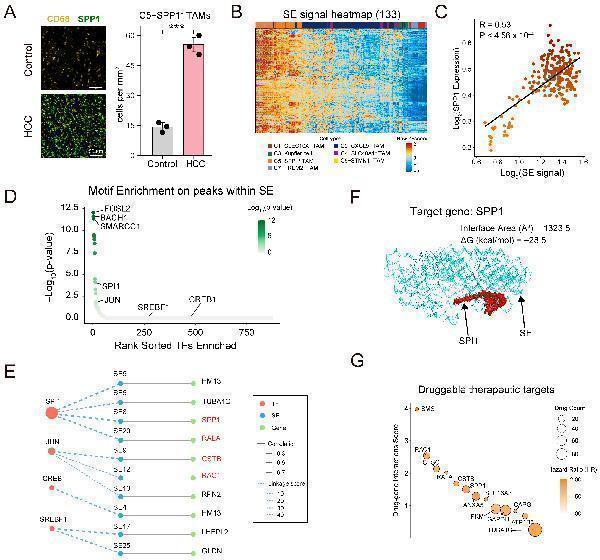
Super-enhancer analysis reveals potential targets in SPP1+ TAMs. (A) Immunofluorescence staining of CD68 (yellow) and SPP1 (green) in liver tissues from control and HCC mice (n = 3 per group). Quantification shows significant increase in SPP1+ TAMs in HCC. Data are shown as mean ± SD; *** p < 0.001. (B) Heatmap showing normalized signal intensity of 133 super-enhancers (rows) across seven macrophage subtypes (columns), revealing subtype-specific enrichment in SPP1+ TAMs. Each row represents one SE region, and each column corresponds to one TAM subtype. (C) Correlation between SE signal and SPP1 expression. R and p-values were calculated by Pearson correlation. (D) Motif enrichment analysis of TFs on peaks within SE region. The TFs were ranked based on statistical significance using hypergeometric testing. (E) TF–SE–target gene regulatory network in SPP1+ TAMs. Nodes in red represent TFs, nodes in blue represent SEs, and nodes in green represent SE target genes. Node size of TFs corresponds to the significance of motif enrichment within SE-associated peaks, measured as −log10(P) from hypergeometric testing. Gene nodes labeled in red indicate known markers of SPP1+ TAMs. Inner edges represent regulatory association between each TF and SE calculated by linking scores. Outer edges reflect the correlation between chromatin accessibility and expression. (F) Structural modeling of the SPI1 protein bound to the SE region that regulates SPP1 gene, as predicted by AlphaFold3 and PDBePISA. (G) Drug–gene interaction analysis of SE target genes in (E). The y-axis indicates druggability score from DGIdb. Dot size reflects the number of drug interactions, and dot color indicates hazard ratio in HCC.
Index in PubMed under a CC BY license. PMID: 40725473
Click image to see more details
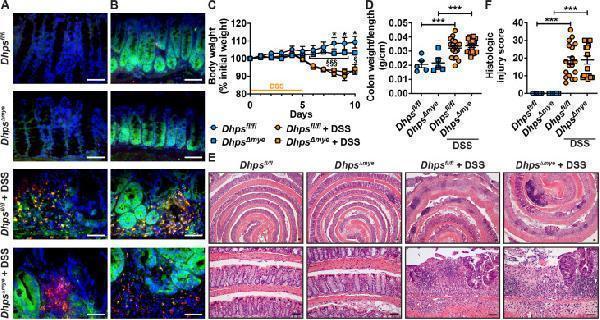
DSS-induced colitis in Dhpsfl/fl and DhpsΔmye mice. Animals were treated or not with 4 % DSS. Expression of CD68 (red) and DHPS (green) (A) or CD68 (red) and EIF5AHyp (green) (B) was assessed in the colon by IF; merged, yellow; nucleus, blue. These pictures are representative of staining performed on 3 control mice and 4–5 DSS-treated mice per genotype. Body weights were measured daily and are shown as percentage of initial body weight (C); *P < 0.05 versus Dhpsfl/fl mice + DSS and §P < 0.05, §§§P < 0.001 compared to DSS-treated DhpsΔmye mice, by two-way ANOVA and Tukey test. At sacrifice, colons were dissected, washed, weighed, and measured (D). Swiss-rolls were stained with H&E (E; scale bars, 50 μm) and analyzed to determine the histologic injury score (F). In (D) and (F), ***P < 0.001, ****P < 0.0001 by one-way ANOVA and Tukey test; Dhpsfl/fl, n = 5; DhpsΔmye, n = 4; Dhpsfl/fl + DSS, n = 18; DhpsΔmye + DSS, n = 18.
Index in PubMed under a CC BY license. PMID: 39027559
Click image to see more details
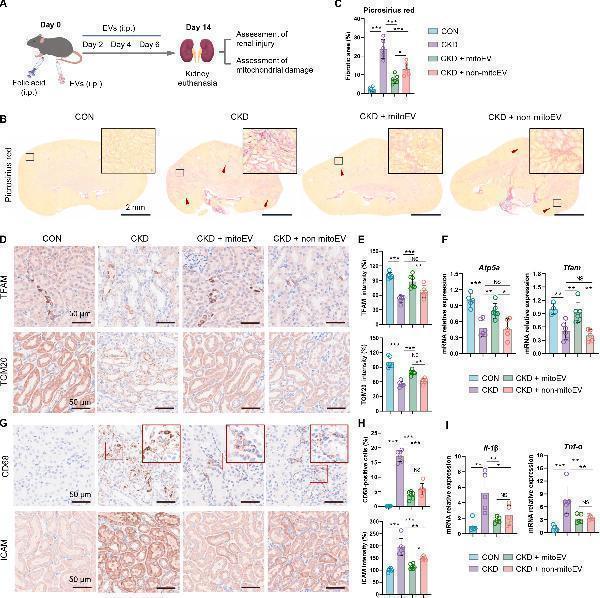
Evaluating the repair efficacy of mitoEVs in CKD. (A) Experimental scheme of the CKD study. (B) Representative picrosirius red stained images of kidneys on day 14 after CKD (scale bar = 2 mm). (C) Quantitative analysis of the renal fibrosis area (n = 6). (D) IHC staining images for TFAM and TOM20 on day 14 after CKD (scale bar = 50 μm). (E) Quantitative analysis of TFAM and TOM20 protein expression (n = 6). (F) qPCR analysis of mitochondrial-related gene expression in the kidneys of the mice (n = 6). (G) Images of IHC staining for CD68 and ICAM on day 14 after CKD induction (scale bar = 50 μm). (H) Quantitative analysis of CD68 and ICAM protein expression (n = 6). (I) qPCR analysis of inflammation-related gene expression in the kidneys of the mice (n = 6). *P < 0.05, **P < 0.01, ***P < 0.001, and NS = not significant.
Index in PubMed under a CC BY license. PMID: 40668934
Click image to see more details
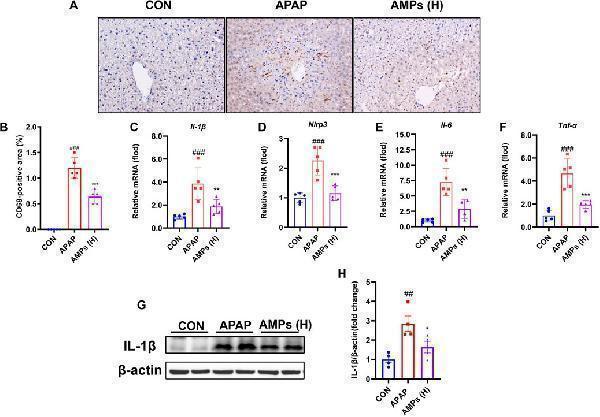
AMPs improves APAP-induced liver inflammation in mice. (A) Immunohistochemical staining results of CD68 in mouse liver tissue. (B) Statistical results of CD68-positive area in mouse liver tissue. (C–F) The IL-1β mRNA, Nlrp3 mRNA, IL-6 mRNA and TNF-α mRNA expression levels in mouse liver tissue (n = 5). (G) Protein expression level of IL-1β in mouse liver tissue. (H) Fold change of IL-1β protein. Data of biochemical indexes were presented as mean ± SD (n = 5). # represents compared with CON group; ## p < 0.01, ### p < 0.001; * indicates compared with APAP group; * p < 0.05, ** p < 0.01, *** p < 0.001.
Index in PubMed under a CC BY license. PMID: 40612747
Click image to see more details
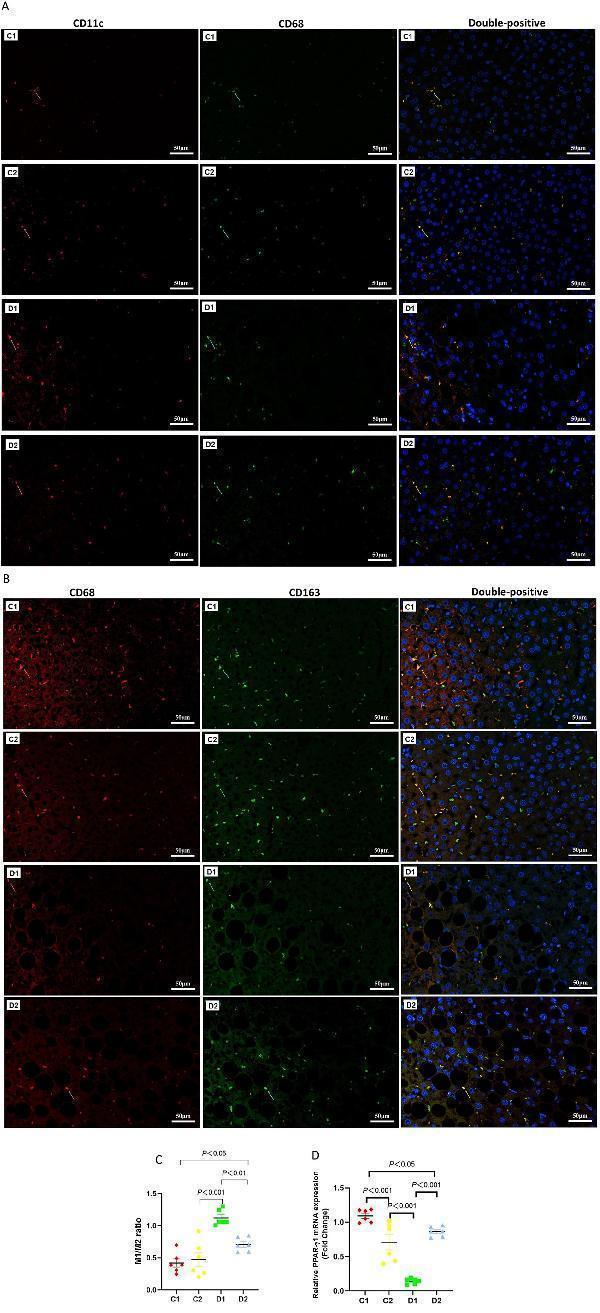
Effects of n-6 PUFA on liver macrophage phenotype in rats with NASH induced by a choline-deficient diet. (A) M1-type Kupffer cells (KCs) identified by double staining: red arrows show CD11c-positive cells, green arrows show CD68-positive cells, and yellow arrows highlight CD11c and CD68 double-positive M1-type KCs (Scale bar – 50 μM). (B) M2-type KCs identified similarly, with red arrows indicating CD163-positive cells, green arrows showing CD68-positive cells, and yellow arrows marking CD163 and CD68 double-positive M2-type KCs (Scale bar – 50 μM). For (A,B) (see ) for full-size photomicrographs. (C) M1/M2 phenotype ratio (unitless), calculated as the proportion of CD68 + CD11c + to CD68 + CD163 + cells. (D) Relative PPAR-γ2 mRNA expression (fold change normalized to GAPDH) in the liver, which is linked to macrophage polarization and inflammation. Data are expressed as mean ±SEM; n = 6/group.
Index in PubMed under a CC BY license. PMID: 40626231
Click image to see more details
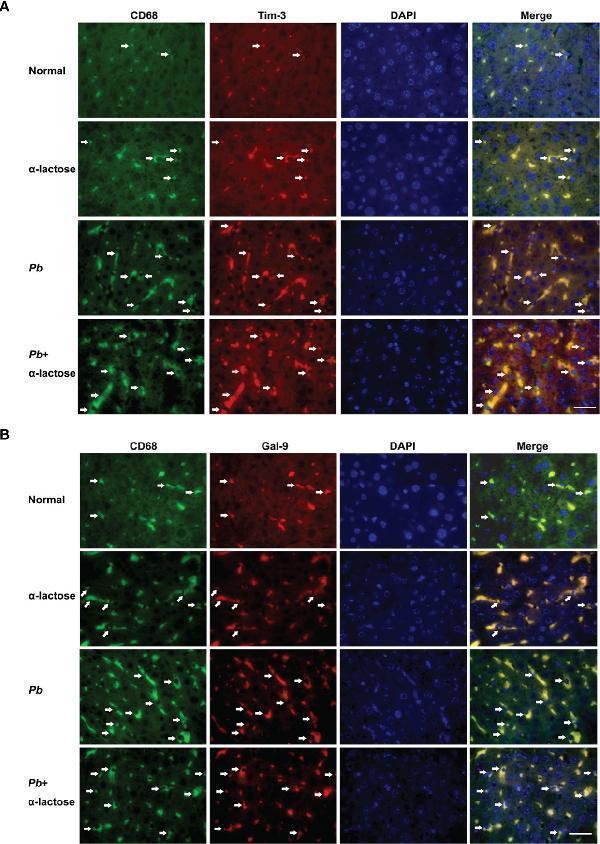
Immunofluorescence staining for CD68 + -Gal-9 + and CD68 + -Tim-3 + macrophages in the liver tissues of Pb ANKA-infected mice with or without α-lactose treatment. Double immunofluorescence showed expression of Gal-9 or Tim-3 on CD68 + macrophages. (A) CD68 + -Gal-9 + macrophages in the liver tissues of malarial mice and malarial mice with α-lactose treatment on day 7 p.i. (B) CD68 + -Tim-3 + macrophages in the liver tissues of malarial mice and malarial mice with α-lactose treatment on day 7 p.i. in comparison of naive and α-lactose-control. Positive cells were indicated with white arrow heads. Original magnification × 1,000.
Index in PubMed under a CC BY license. PMID: 34899708
Click image to see more details
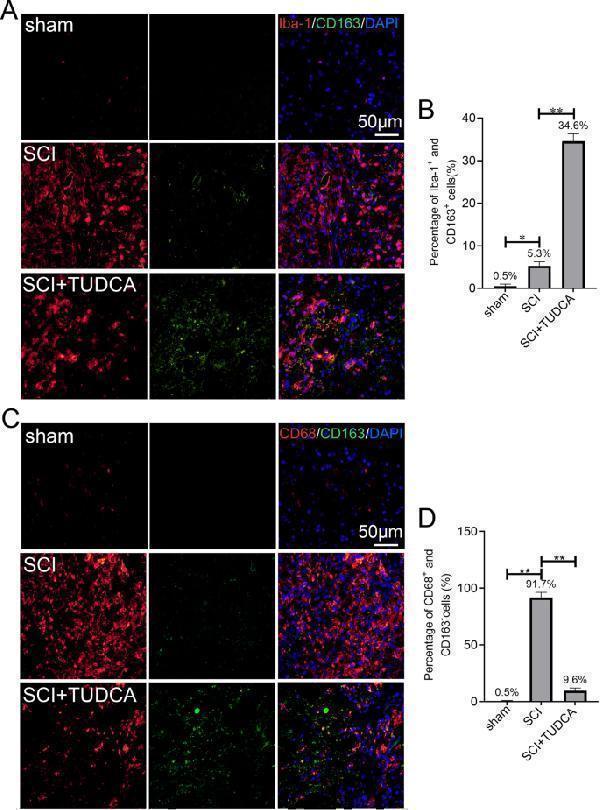
TUDCA treatment promotes microglia polarization toward the M2 phenotype. A , C Immunofluorescent staining of Iba-1(red) /CD163(green) or CD68(red)/CD163(green) in the lesion site of the spinal cord 14 days after SCI. B , D Quantification the number of Iba-1 + /CD163 + or CD68 + /CD163 − cells in spinal cord. All experiments were performed in triplicated and data were presented as means ± SD, n =3 per group. * P < 0.05, ** P < 0.01
Index in PubMed under a CC BY license. PMID: 34544428
Click image to see more details
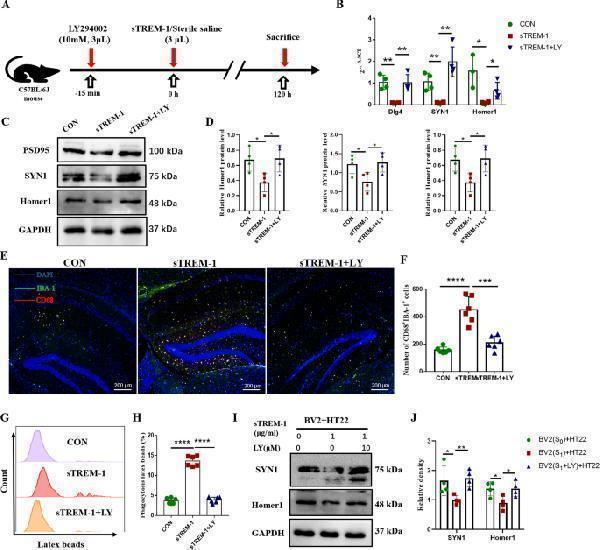
Inhibition of the PI3K–AKT pathway attenuates the phagocytic activation of microglia induced by sTREM-1. (A) The experimental plan was visualized. (B) The mRNA expression levels of Dlg4 , SYN1, and Homer1 in mouse hippocampus detected by Q-PCR, n = 4. (C) Representative western blot images of Dlg4, SYN1, and Homer1 protein expression in mouse hippocampus. (D) Quantitative analysis of PSD95, SYN1, and Homer1 protein level in mouse hippocampus, n = 4. (E) Representative image of IBA-1 and CD68 immunofluorescence double staining of the hippocampus in per group mice, scale bar is 200 μm. (F) Quantitative analysis of IBA-1 and CD68 double-positive cells, n = 6. (G) Representative histograms of BV2 cells phagocytosis of latex beads in different treatment groups detected by flow cytometry. (H) Phagocytosis in different BV2 cells groups detected by flow cytometry. (I) Representative western blot images of SYN1 and Homer1 protein expression in different HT22 cells groups. (J) Quantitative analysis of SYN1 and Homer1 protein level in different HT22 cells groups. Data are presented as means ± SEM of at least three separate experiments, * P < 0.05, ** P < 0.01, *** P < 0.001, **** P < 0.0001.
Index in PubMed under a CC BY license. PMID: 35487953
Click image to see more details
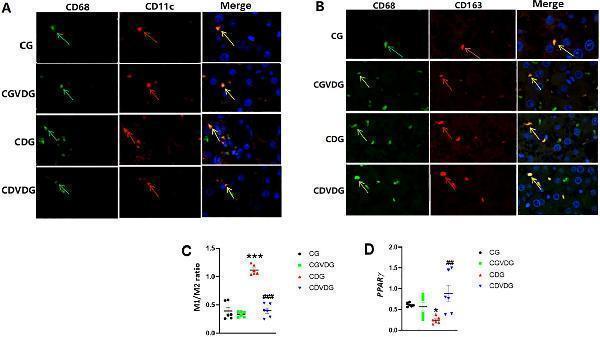
Effects of 1,25 VD3 supplementation on macrophage polarization and PPARγ expression in NASH rats. (A) M1-type KCs identified by double immunofluorescence staining; green arrows indicate CD68-positive cells, red arrows indicate CD11c-positive cells, and yellow arrows indicate CD11c/CD68 double-positive cells (M1KCs). (B) M2-type KCs identified by double immunofluorescence staining; green arrows indicate CD68-positive cells, red arrows indicate CD163-positive cells, and yellow arrows indicate CD163/CD68 double-positive cells (M2KCs). Antibodies were diluted as follows: anti-CD68 (1:100), anti-CD11c (1:100), and anti-CD163 (1:200). (C) The M1/M2 macrophage ratio was determined by quantifying the proportion of M1 and M2 KCs in liver tissue, as identified through double immunofluorescence staining. (D) PPARγ levels in liver tissue of the indicated groups were determined by qPCR and expressed as relative mRNA levels following normalization to GAPDH mRNA levels. Data are presented as mean ± SEM, n=6. *p < 0.05 and ***p < 0.001 vs CG; ##p < 0.01 and ###p < 0.001 vs CDG.
Index in PubMed under a CC BY license. PMID: 40190400
Click image to see more details
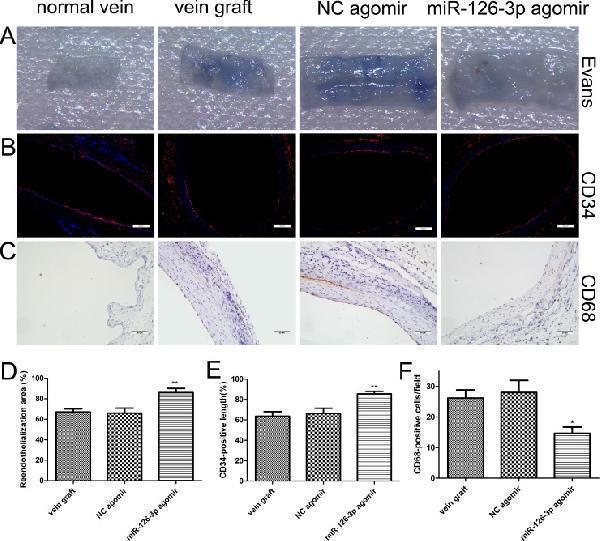
miR-126-3p agomir promoted reendothelialization in vivo . ( A ) Representative Evans Blue staining photomicrographs of vessel wall harvested from normal and vein arterialization rats at 14 days after surgery. ( B ) Immunofluorescence staining for vascular endothelial cell marker CD34 (red) was used to assess endothelial recovery of the vein graft for each group. Nuclei were stained with DAPI (blue). Scale bars represent 200 μm. ( C ) Representative staining of CD68 within the intima layer 28 days after surgery is shown. Scale bars represent 50 μm. ( D ) The ratio of non-stained area (white) to the total en face area of the vein graft was used to evaluate reendothelialization. ( E ) Quantification of the reendothelialized areas was performed as a percentage of the CD34-positive surface to the total luminal surface ( F ) Quantitative evaluation of the number of accumulating CD68-positive monocytes/macrophages within the vascular wall. Values are expressed as the mean ± SEM. n =6 per group. * P < 0.05, ** P < 0.01vs NC agomir and vein graft groups.
Index in PubMed under a CC BY license. PMID: 29290989
Click image to see more details
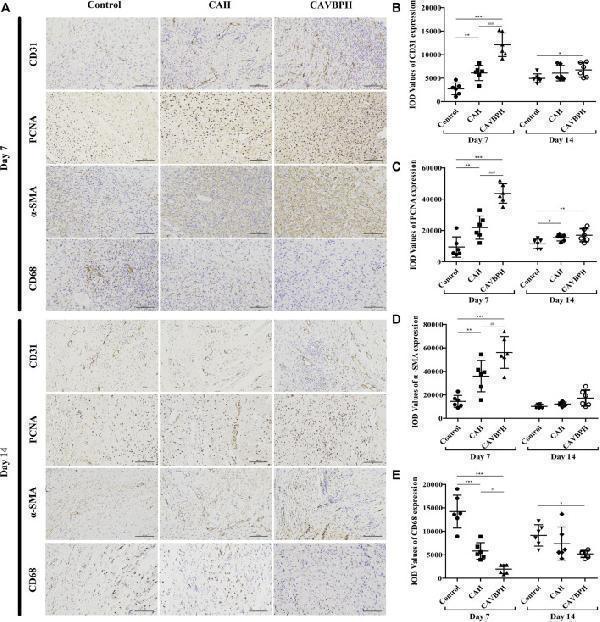
Immunohistochemistry staining of wound tissues on days 7 and 14. (A) Representative images for CD31, PCNA, α-SMA, and CD68 staining (scale bar = 100 μm). (B–E) Quantification of CD31, PCNA, α-SMA, and CD68 protein expressions, respectively. * p < 0.05, ** p < 0.01, *** p < 0.001 vs . control group; # p < 0.05, ## p < 0.01, ### p < 0.001 vs . CAH group as statistically significant.
Index in PubMed under a CC BY license. PMID: 35784748
Click image to see more details
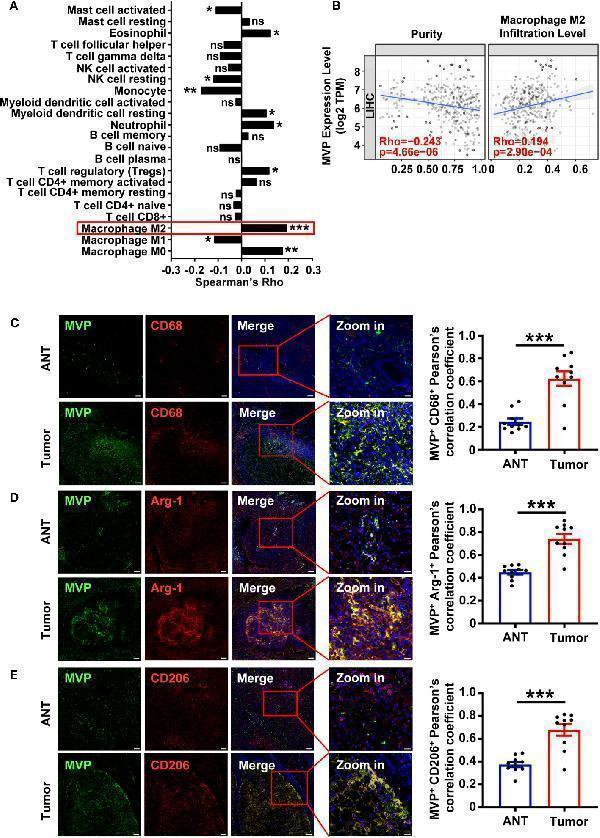
MVP positively correlates with CD68 + /Arg-1 + /CD206 + TAMs in HCC samples. (A) The histogram of Rho between MVP expression and 22 types of immune cell infiltration levels in LIHC from the TIMER2.0 database. The red box highlights the immune cells with the highest positive correlation. (B) The scatter plot of the correlation between MVP expression level with tumor purity and M2 macrophage infiltration level determined by the TIMER2.0. TPM: Transcripts Per Kilobase of exon model per Million mapped reads. (C–E) Immunofluorescence representative images and co-localization quantitative analysis of MVP (green) with CD68 (red) (C) , MVP (green) with Arg-1 (red) (D) , and MVP (green) with CD206 (red) (E) in adjacent nontumorous tissues (ANT) and HCC tumor tissues (n=10). Scale bar, 100 µm (MVP/CD68/Arg-1/CD206/Merge) and 20 µm (Zoom in). All levels of co-localization are indicated by Pearson’s correlation coefficients calculated using Image J. Data are expressed as means ± SEM, two-tailed Student’s t-test, (***P < 0.001, **P < 0.01, *P < 0.05, ns, no significance). See also and .
Index in PubMed under a CC BY license. PMID: 38264642
Click image to see more details
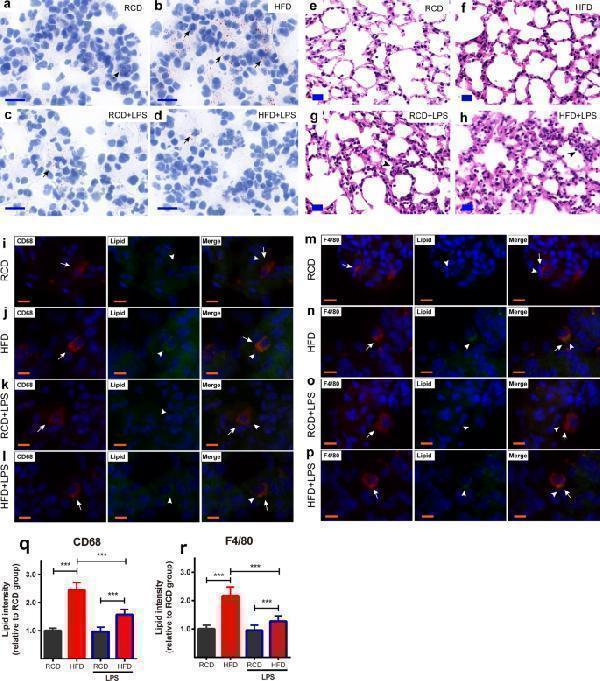
Obesity-induced pathological changes in lungs. ( a–d ) Oil Red O staining of lung in RCD and HFD mice. Less fats accumulated in the lung tissue (arrows pointing) in RCD mice ( a ), RCD + LPS mice ( c ). A large quantity of lipid droplets (arrows pointing) were accumulated in lung tissue in HFD mice ( b ) and less fat deposits in HFD + LPS mice ( d ). ( a – d ) scale bars represent 20 μm, the nuclei were re-stained blue with hematoxylin. ( e–h ) HE staining of lungs in RCD and HFD mice. HE staining of RCD mice 24 h after intratracheal administration of saline ( e ). HE staining of HFD mice 24 h after intratracheal administration of saline ( f ). HE staining of RCD mice 24 h after intratracheal administration of LPS ( g ). HE staining of HFD mice 24 h after intratracheal administration of LPS, arrows designate the alveolar inflammatory infiltration and alveolar wall thickening after LPS challenge ( h ). ( e–h ) scale bars represent 20 μm. ( i–l ) Lung immunofluorescence staining of CD68 + macrophages in RCD and HFD mice. CD68 + macrophages were stained red (arrow pointing). Lipid droplets were stained green (arrow head pointing) by Bodipy. Representative image of CD68 + macrophages observed in HFD ( j ) mice, the merged images showed the lipid droplets were located in the macrophages of the HFD mice ( j ) and the lipid droplets decreased in macrophages of HFD + LPS ( l ) mice. Representative images of CD68 + macrophages observed in RCD ( i ) and RCD + LPS ( k ) mice, the merged images indicated there were less lipid droplets in macrophages. ( i–l ) scale bars represent 10 μm. ( m–p ) Lung immunofluorescence staining of F4/80 + macrophages in RCD and HFD mice. F4/80 + macrophages were stained red (arrow pointing). Lipid droplets were stained green (arrow head pointing) by Bodipy. Representative image of F4/80 + macrophages were observed in HFD ( n ) mice, the merged images showed the lipid droplets located in the F4/80 + macrophages in HFD ( n ) mice and the lipid droplets decreased in macrophages of HFD + LPS ( p ) mice. Representative images of F4/80 + macrophages were observed in RCD ( m ), RCD + LPS ( o ) mice, the merged images indicated there were less lipid droplets in F4/80 + macrophages. ( m–p) scale bars represent 10 μm. ( q–r ) Quantification analysis of lipid accumulation in each group (n = 240 cells/12 slices/4 mice for each group). Data were normalized to the averaged signal intensity from RCD mice for each group. ( q ) Lipid intensity in CD68 + macrophages, RCD: 1.0 ± 0.09; HFD: 2.44 ± 0.27; RCD + LPS: 0.97 ± 0.16; HFD + LPS: 1.57 ± 0.19. ( r ) Lipid intensity in F4/80 + macrophages, RCD: 1.0 ± 0.13; HFD: 2.17 ± 0.31; RCD + LPS: 0.95 ± 0.18; HFD + LPS: 1.27 ± 0.19. *** P < 0.001, significant differences between each group (ANOVA, LSD-t post hoc).
Index in PubMed under a CC BY license. PMID: 33110180
Click image to see more details
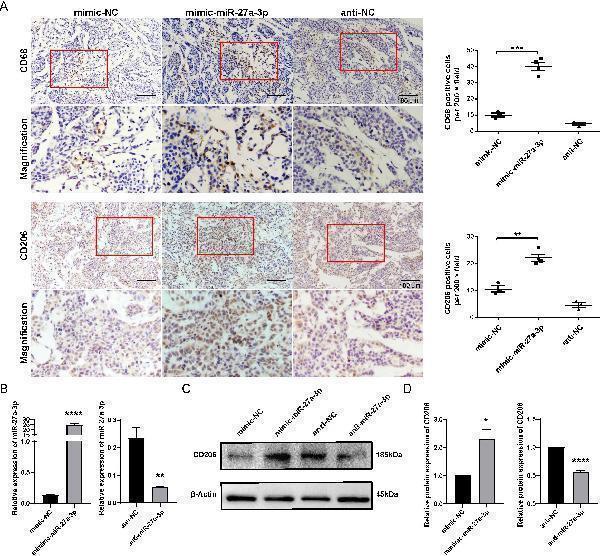
miR-27a-3p promotes the M2 polarization of macrophage. (A) Representative images for the immunohistochemical staining of macrophage maker CD68 and CD206 in nude mice xenografts, bar = 100 μm; and the number of CD68, CD206 positive cells per 200 × field for each group (mean ± SEM). **P < 0.01, ***P < 0.001 vs control. Below each group of images is an image showing the magnification of the corresponding box area above. (B) The expression of miR-27a-3p in mimic-miR-27a-3p or anti-miR-27a-3p treated THP1 cells was evaluated by qRT-PCR 24 h after transfection, normalized to U6 snRNA (mean ± SEM). **P < 0.01, ****P < 0.0001 vs. control. (C, D) The expression of CD206 was evaluated in mimic-miR-27a-3p or anti-miR-27a-3p treated THP1 cells by western blotting normalized to β-actin (mean ± SEM). *P < 0.05, ****P < 0.0001 vs. control. The data were from three independent experiments.
Index in PubMed under a CC BY license. PMID: 39742261
Click image to see more details
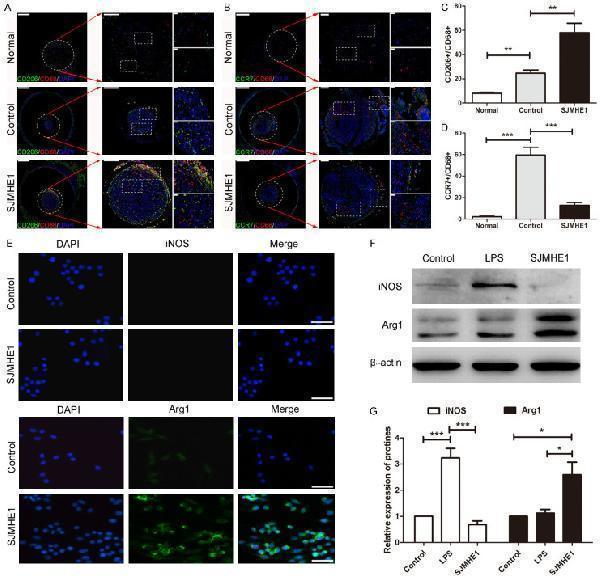
SJMHE1 induced M2 polarization of macrophages. A, B. Images of double immunofluorescence staining of CD206 + (green)/CD68 + (red) and CCR7 + (green)/CD68 + (red) macrophages in cross-sections of the regenerated nerve region at the midpoint of the nerve conduit. Scale bar (left) = 500 μm. The round and boxed areas are higher-magnification images. Scale bar = 200 μm and 20 μm. DAPI (blue)-stained nuclei. C, D. Quantitative analysis of the number of CD206 + /CD68 + and CCR7 + /CD68 + macrophages in different groups. The ratio of CD206 + cells to CD68 + cells was calculated as the ratio of M2 macrophages. The ratio of CCR7 + cells to CD68 + cells was calculated as the ratio of M1 macrophages. The SJMHE1 group vs the control group; n = 6 for each group. E. Immunofluorescence staining of macrophages with the phenotypic markers iNOS and Arg1. DAPI (blue)-stained nuclei. Scale bar = 100 μm. n = 3. F. Western blot analysis of iNOS and Arg1 expression in different groups. The LPS group represents the positive control group. G. Statistical analysis of the relative expression of iNOS and Arg1 proteins. The SJMHE1 group vs the control group or LPS group; n = 3 for each group; * P <0.05, ** P <0.01, *** P <0.001 (ANOVA). The data are presented as the mean ± SEM.
Index in PubMed under a CC BY license. PMID: 33841657
Click image to see more details
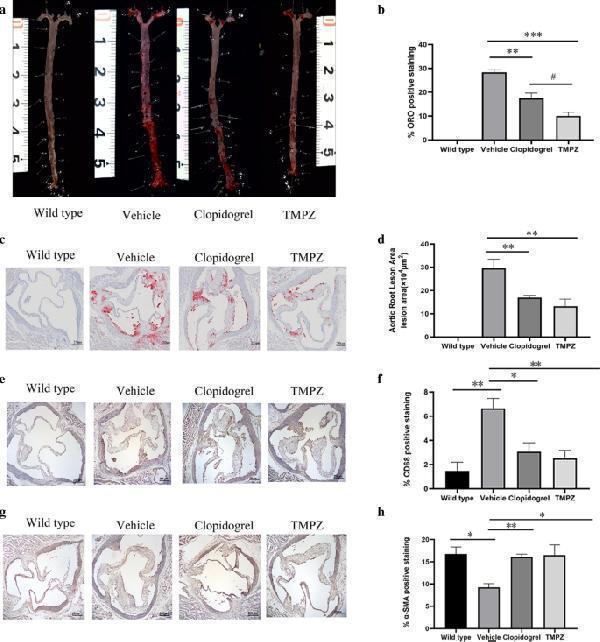
Histological analysis. a Representative images of en face analysis. n = 6. b Quantitative analysis of lesion areas in whole aortas. Differences were assessed by unpaired students’ t -test. c Representative images of Oil Red O staining of aortic root sections. d Quantitative analysis of lesion areas in aortic root sections. e Representative images of macrophage (CD68) analysis ( b ) Quantitative analysis of lesions area in macrophage analysis. f Representative images of SMC (SMA) analysis ( g ) Quantitative analysis of lesions area in SMC. Differences were assessed by unpaired students’ t -test. * p < 0.05 vs Vehicle, ** p < 0.01 vs Vehicle. # p < 0.05 vs clopidogrel. Scale bar = 250 μm. n = 3.
Index in PubMed under a CC BY license. PMID: 33187537
Click image to see more details
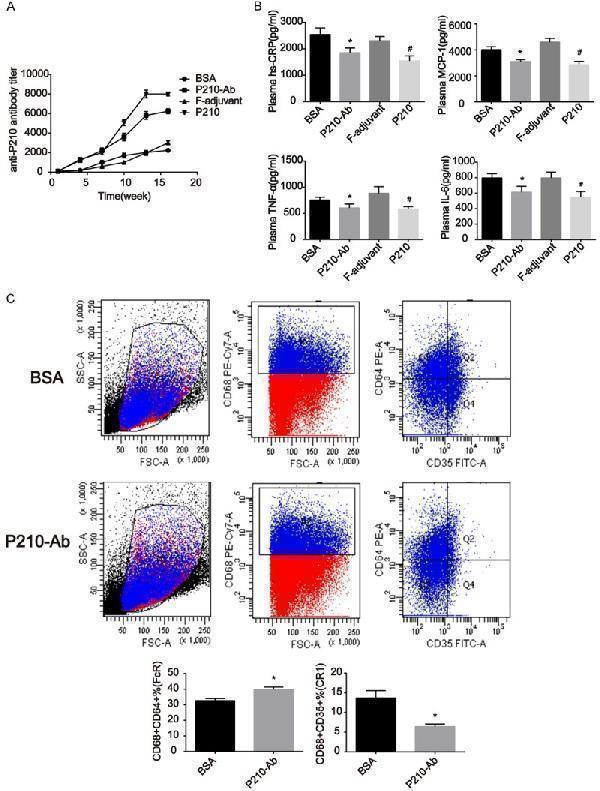
P210-Ab reduced plasma levels of hsCRP, MCP-1, TNF-α, and IL-6, and affected the expression of FcR (CD64) and CR1 (CD35) on PBMM. A. The titer of antibodies against P210 changes over time in ApoE -/- mice. The titer of P210-Ab started to increase significantly after the first injection and continued to increase gradually with the following booster immunization in the P210 and P210-Ab groups but increased slowly in the BSA and F-adjuvant groups. B. P210-Ab affected the plasma levels of hsCRP, MCP-1, TNF-α, and IL-6 in ApoE -/- mice at the age of 23 weeks. The plasma levels of hsCRP, MCP-1, TNF-α, and IL-6 were determined by ELISA. C. Immunization had an impact on PBMM of ApoE -/- mice. The number of CD68 + CD64 + and CD68 + CD35 + cells in P210-Ab group and BSA group (control group), respectively, are shown. Data are expressed as mean ± SEM. *P<0.05 versus BSA group; #P<0.05 versus F-adjuvant group.
Index in PubMed under a CC BY license. PMID: 30018722
Click image to see more details
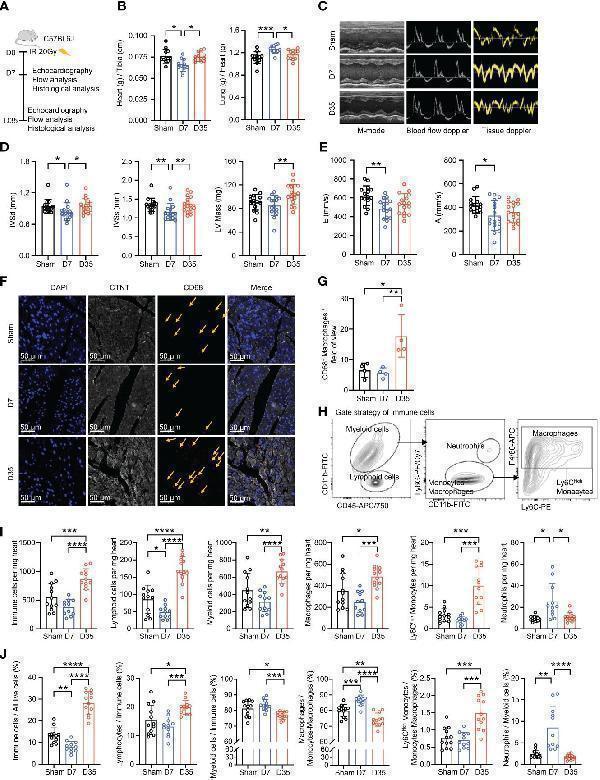
Transient changes in cardiac function are accompanied by changes in the number of cardiac macrophages. (A) Schematic diagram of animal modeling and experimental procedures. (B) Comparisons on Heart (g) to Tibia (cm) ratio a and Lung (g) to Heart (g) ratio b (Sham, n=12; D7, n=11; D35, n=12). Data were pooled from three independent experiments. (C) Representative images of echocardiography in cardiac function. M-mode, blood flow doppler and tissue doppler images views. (Sham, n=15; D7, n=16; D35, n=16). Data were pooled from three independent experiments. (D) Echocardiographic analysis of systolic function by M-mode: interventricular septum in diastole (IVSd b ), interventricular septum in systole (IVSs b ) and left ventricular (LV) Mass (corrected) a . (E) Echocardiographic analysis of diastolic function by blood flow doppler and tissue doppler: the speed E peak a (left ventricular early-diastolic fast filling) and the speed A peak a (left ventricular late-diastolic filling). (F) Representative images of immunofluorescent staining: CD68+ macrophages (Red), CTNT+ cardiomyocytes (Gray), DAPI+ nucleus (Blue). The yellow arrows point to macrophages. Scale bar, 50 μm. (n=4 in each group). (G) The statistical plot a of panel (F) . (H) Flow cytometry gating scheme used to identify cardiac immune cells. (I) Comparison on number of immune cells in per mg heart tissue by flow analysis. Ly6C high Monocytes c ; Neutrophils c ; the other cell types a . (J) Comparison on percentage of each cell type by flow analysis. Immune cells/All live cells (%) c ; Ly6C high Monocytes/Myeloid cells (%) c ; Neutrophils/Myeloid cells (%) b ; the other parameters a . Results of (I, J) were from three independent flow analyses. (Sham: n=12; D7: n=11; D35: n=12). “a”: Ordinary one-way ANOVA test; “b”: Kruskal-Wallis test; “c”: Welch ANOVA test. * P <0.05, ** P <0.01, *** P <0.001 and **** P <0.0001.
Index in PubMed under a CC BY license. PMID: 38601160
Click image to see more details
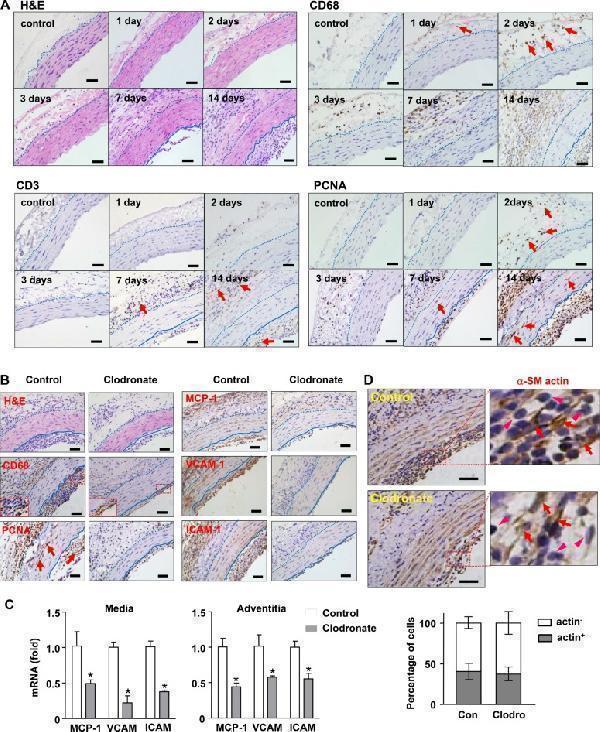
Effects of macrophage depletion with clodronate liposomes on transplantation-induced arterial remodeling and inflammation. ( A ) Time course of early adventitial macrophage infiltration and subsequent neointimal hyperplasia revealed by hematoxylin and eosin (H&E) staining and immunohistochemistry. Macrophages were stained with anti-CD68; T cells stained with anti-CD3; and proliferating cells stained with anti-PCNA. ( B ) Histology images showing that macrophage depletion inhibited the growth of neointima, diminished infiltrating macrophages in the adventitia, reduced the number of PCNA + cells, and reduced inflammatory responses (samples of day 14). ( C ) Effects of clodronate on the expression levels of various inflammatory molecules measured with qPCR. ( D ) Proportion of smooth muscle cells (stained with anti-a-SM actin, indicated by arrows) and non-muscle cells (arrowheads) in the neointima of untreated and clodronate-treated vessels. Data are mean ± SD. * P < 0.05 versus control, unpaired t-test , n = 6. Dashed lines indicate the border of adventitia and media; the solid lines indicate the border between media and neointima. Bar represents 50 μm.
Index in PubMed under a CC BY license. PMID: 28415796
Click image to see more details
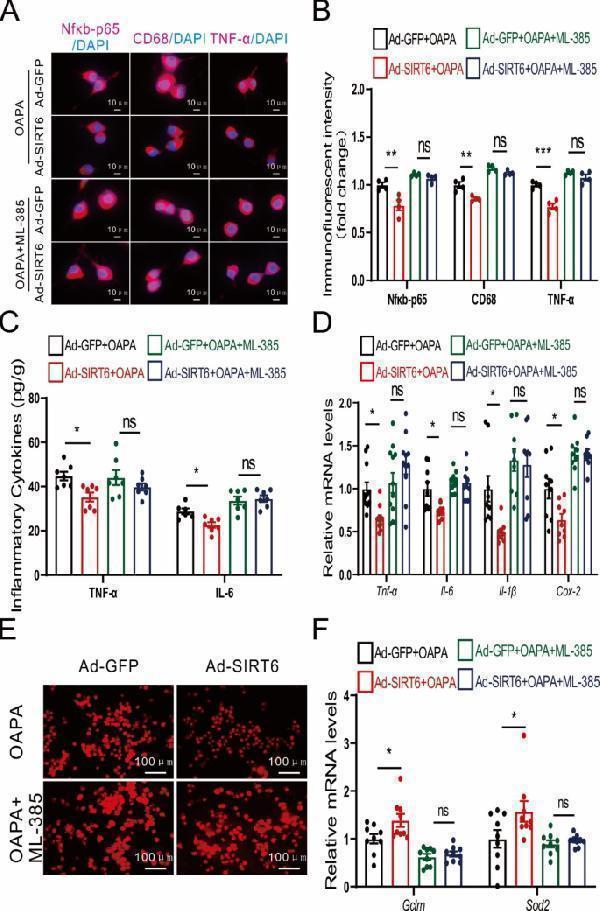
NRF2 antagonists counteract the ameliorating effects of Sirt6 on inflammation and oxidative stress. A Immunofluorescence images of Nfκb-p65, CD68, and TNF-α in BV2 cells with combined Sirt6 overexpression and NRF2 antagonist treatment. Scale bars: 10 μm. B Quantitative analysis of immunofluorescent intensity of Nfκb-p65, CD68, and TNF-α in BV2 cells with combined Sirt6 overexpression and NRF2 antagonist treatment. n = 4/group. C Expression of inflammatory cytokines TNF-α and IL-6. n = 6–7/group. D mRNA expression of Tnf-α , Il-6 , Il-1β , and Cox-2 . n = 9–12/group. E Immunofluorescence images of ROS. Scale bars: 100 μm. F mRNA expression levels of Gclm and Sod2 . n = 9/group. B-D, F Two-way ANOVA analysis followed by the Tukey post hoc test were performed. ns: not significant. Data are presented as mean ± SEM, * p < 0.05, ** p < 0.01, *** p < 0.001
Index in PubMed under a CC BY license. PMID: 37582706
Click image to see more details
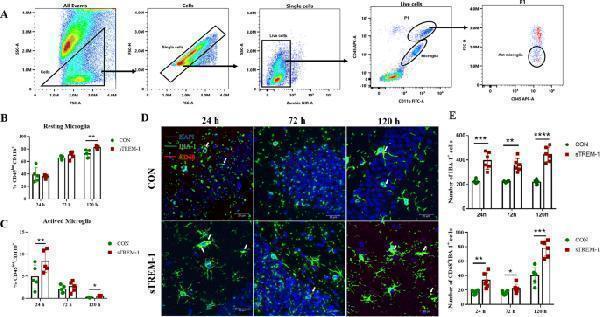
sTREM-1 promotes the activation of microglia in the hippocampus of mice. (A) The gating strategy of resting and activating microglia in the mouse hippocampus. (B) Quantitative analysis of the proportion of resting microglia in the hippocampus of per group mice, n = 5. (C) Quantitative analysis of the proportion of activated microglia in the hippocampus of each group mice, n = 4. (D) Representative image of IBA-1 and CD68 immunofluorescence double staining of the hippocampus in per group mice, where the white arrows indicated CD68 + IBA-1 + cells, scale bar is 200 μm. (E) Quantitative analysis of IBA-1 + , IBA-1 and CD68 double-positive cells, n = 6; Data are presented as means ± SEM of at least three separate experiments, * P < 0.05, ** P < 0.01, *** P < 0.001, **** P < 0.0001.
Index in PubMed under a CC BY license. PMID: 35487953
Click image to see more details
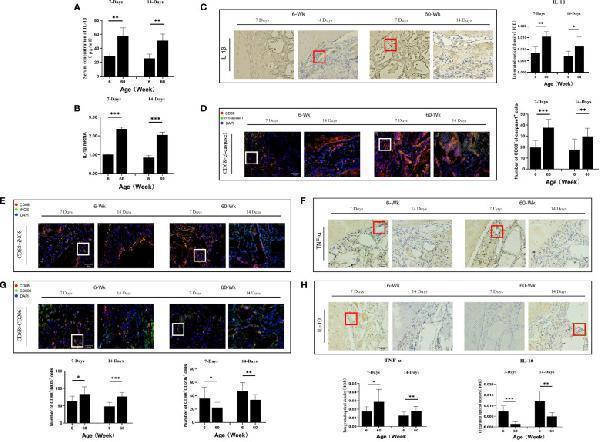
(A) ELISA determination of IL-1β levels in the serum of young and aged SD rats. (B) Local IL-1β gene expression in young and aged SD rats. (C) Immunohistochemical detection of IL-1β secretion in Bio-Oss® and its surrounding tissues on days 7 and 14 to assess differences in the local inflammatory microenvironment of defects in young and aged rats; Red square: positive cells. (D) CD68 (red) and cl-caspase-1 (green) co-immunolabeling to evaluate NLRP3 inflammasome expression in macrophages; nuclei were stained with DAPI; White square: double-positive cells. (E, G) CD68 (red), i-NOS (green), CD68 (red) and CD206 (green) co-immunolabeling was used to evaluate macrophage phenotypes; nuclei were stained with DAPI; White square: double-positive cells. (F, H) TNF-α and IL-10 were used to assess changes in the local inflammatory microenvironment; Red square: positive cells. Aged rats showed a significant increase in M1 macrophages and a significant decrease in M2 macrophages in the local microenvironment at the sites of bone defects, suggesting a delayed local inflammatory phase, elevated systemic and local IL-1β expression, and increased NLRP3 inflammasome activation in macrophages in aged rats. *P<0.05, **P<0.01, ***P<0.001.
Index in PubMed under a CC BY license. PMID: 37153554
Click image to see more details
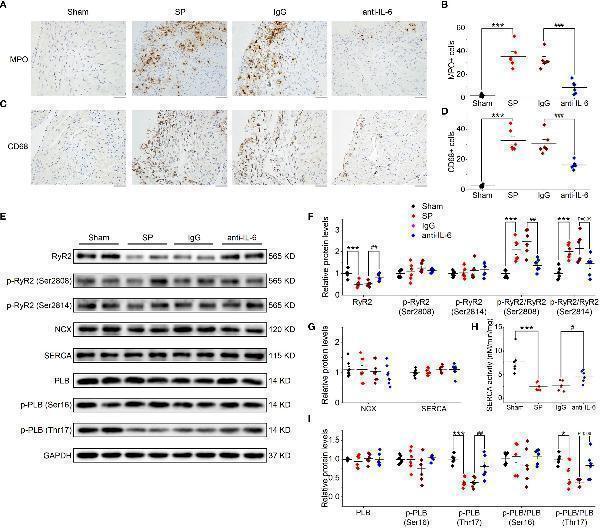
IL-6 neutralization prevents atrial inflammation and the alterations in Ca 2+ handling proteins in SP rats. (A, B) Representative images of atrial MPO staining (A) and quantification (B) in the 4 indicated groups. Five visual fields are taken in each sample, and the number of immune cells from these fields is quantified with ImagePro 6.0 software and is averaged to make a statistical analysis. n = 6-7/group. Scale bar: 50 µm. (C, D) Representative images of atrial CD68 staining (C) and quantification (D) . n = 6-7/group. Scale bar: 50 µm. (E–I) Original Western blot (E) and quantification of the expression of RyR2, p-RyR2 (Ser2808), p-RyR2 (Ser2814) (F) , the expression of SERCA and NCX (G) , SERCA activity (H) , the expression of PLB, p-PLB (Ser16), and p-PLB (Thr17) (I) , in atrial tissue of 4 indicated groups. n =6-8/group. * P < 0.05, *** P < 0.001 vs . Sham; # P < 0.05, ## P < 0.01, ### P < 0.001 vs . IgG, determined by Student t-test or one-way ANOVA with Bonferroni’s post-hoc test. (F–H) .
Index in PubMed under a CC BY license. PMID: 34975847
Click image to see more details
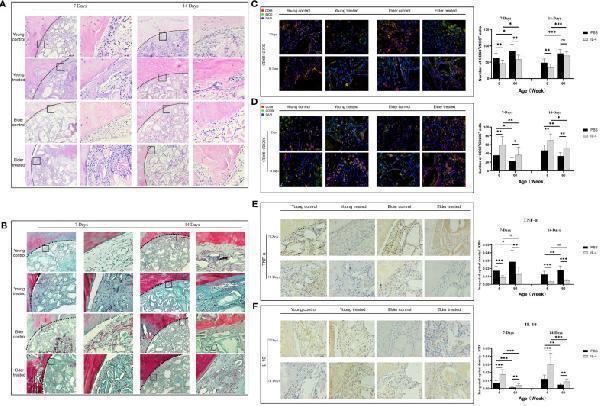
Local immunomodulation by IL-4 (A) H&E and (B) Masson’s trichrome staining. (C, D) CD68 (red), i-NOS (green), CD68 (red), and CD206 (green) co-immunolabeling was used to evaluate macrophage phenotypes; nuclei were stained with DAPI. (E, F) TNF-α and IL-10 were used to assess changes in the local inflammatory microenvironment. IL-4 recruited more macrophages at the sites of bone defects in aged rats and altered the M1/M2 ratio to form a promoted healing microenvironment by promoting the polarization of M1-type macrophages to M2-type macrophages. *P<0.05, **P<0.01, ***P<0.001. ns, no significance.
Index in PubMed under a CC BY license. PMID: 37153554
Click image to see more details
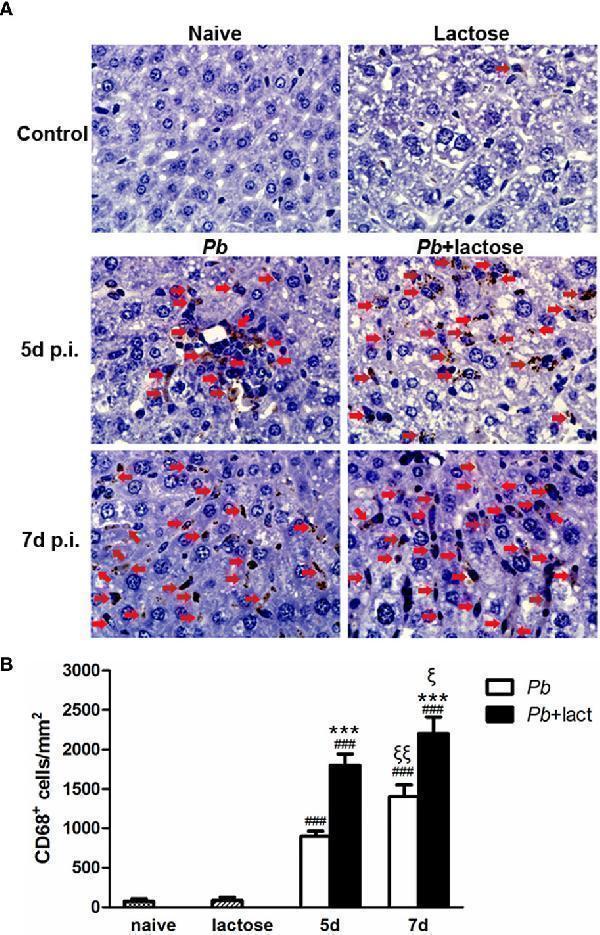
Immunohistochemical staining for CD68 + macrophages in the livers of Pb ANKA-infected mice with or without α-lactose treatment. (A) Immunohistochemical staining of CD68 + macrophages in the livers of uninfected mice, uninfected mice with α-lactose treatment, malarial mice, and malarial mice with α-lactose treatment on day 7 p.i. Positive cells were indicated with red arrow heads. Original magnification × 1,000. (B) Morphometric analysis of liver tissues. Shown are CD68 + macrophages per square millimeter. Data are presented as means ± SD; experiments were performed with three mice per group. ### P < 0.001, malarial mice on day 5 p.i., malarial mice with α-lactose treatment on day 5 p.i., malarial mice on day 7 p.i., and malarial mice with α-lactose treatment on day 7 p.i. vs . naive group; *** P < 0.001, malarial mice with α-lactose treatment on day 5 p.i. vs . malarial mice on day 5 p.i.; *** P < 0.001, malarial mice with α-lactose treatment on day 7 p.i. vs . malarial mice on day 7 p.i.; ξ P < 0.05, malarial mice with α-lactose treatment on day 7 p.i. vs . malarial mice with α-lactose treatment on day 5 p.i.; ξξ P < 0.01, malarial mice on day 7 p.i. vs . malarial mice on day 5 p.i.
Index in PubMed under a CC BY license. PMID: 34899708
Click image to see more details
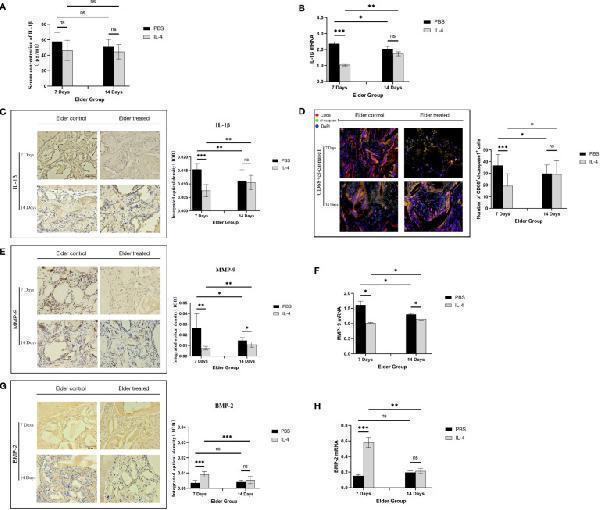
Local immunomodulation by IL-4 (A) ELISA detection of IL-1β in the serum of aged SD rats. (B) Changes in local IL-1β gene expression in aged SD rats. (C) Immunohistochemical detection of IL-1β secretion in Bio-Oss® and its surrounding tissues on days 7 and 14 to assess differences in the local inflammatory microenvironment of defects in aged rats. (D) CD68 (red) and cl-caspase-1 (green) co-immunolabeling was used to evaluate NLRP3 inflammasome expression in macrophages; nuclei were stained with DAPI. (E, G) Immunohistochemical detection of BMP-2 and MMP-9 secretion in Bio-Oss® and its surrounding tissues on day 7 and day 14. (F, H) Local BMP-2 and MMP-9 gene expression in aged SD rats was used to assess the dynamic balance of local bone regeneration and bone resorption in the defects of aged rats. Local immunomodulation in aged rats resulted in reduced M1 numbers, inhibition of NLRP3 inflammasome activation, downregulation of IL-1β expression, reduced osteoclast fractionation, changes in the M1/M2 ratio, and increased BMP-2 expression, thereby promoting bone regeneration. *P<0.05, **P<0.01, ***P<0.001. ns, no significance.
Index in PubMed under a CC BY license. PMID: 37153554
Click image to see more details
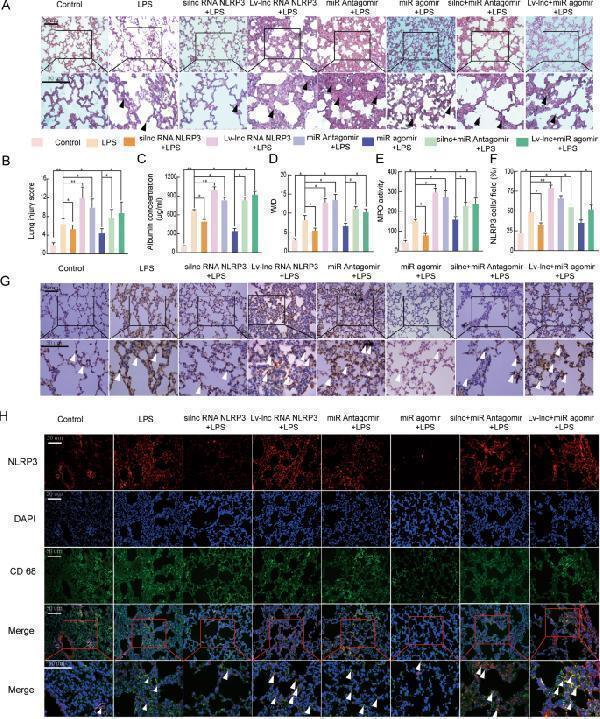
LncRNA NLRP3/miR-138-5p/NLRP3 functions via the ceRNET during the NLRP3-triggered inflammatory response in vivo. Rat lungs were injected with PBS in the control group and LPS-treated rats were further treated with si-r-lncRNA NLRP3, Lv-lncRNA NLRP3, agomiR-138-5p, antagomiR-138-5p, Lv-lncRNA NLRP3 + agomiR-138-5p, and si-r-lncRNA NLRP3 + antagomiR-138-5p. A Lung tissue samples were collected 6 h after establishing LPS-induced ALI to analyse the histopathological changes (×200, ×400). The black arrow indicates neutrophil infiltration, pulmonary oedema, alveolar wall thickening, and alveolar haemorrhage. B The lung injury score was determined via H&E staining, a representative histological analysis ( n = 6 animals per group). C ELISA was used to measure the BALF albumin content. D Detection of the lung W/D ratio in rats. E MPO activity in the lung tissues of rats. F , G Immunohistochemical detection of the NLRP3 contents in rat lung tissues (×200, ×400). H The inflammatory response in NR8383 AM cells was suppressed by si-r-lncRNA NLRP3 and miR-138-5p mimics alone or in combination, as shown by the decreased number of cells colabeled with CD68 (green) and NLRP3 (red). LncRNA NLRP3 overexpression, miR-138-5p inhibition, and NLRP3 augmented the inflammatory response in LPS-induced ALI with more NLRP3 and CD68 anchored in the plasma membrane of the AM cells. The data are presented as mean ± SE ( n = 6). * P < 0.05; ** P < 0.01; *** P < 0.001; NS, no statistically significant difference.
Index in PubMed under a CC BY license. PMID: 34599154
Click image to see more details
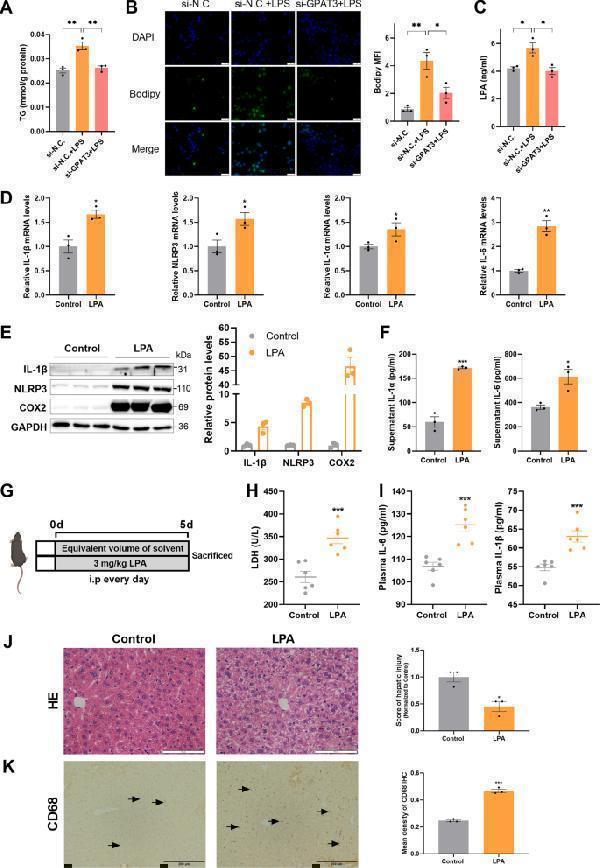
Exogenous LPA enhanced inflammatory response in vitro and vivo. A Effect of si-GPAT3 or si-N.C. on TG content in KCs stimulated with LPS (1 μg/ml) for 24 h ( n = 3). B Bodipy 493/503 fluorescence visualized by fluorescence microscopy (scale bars represent 50 μm) ( n = 3). C LPA concentrations in the supernatant of KCs after LPS (100 ng/ml 12 h) stimulated and with or without si-GPAT3 ( n = 3). D The mRNA expression of IL-1β, NLRP3, IL-1α and IL-6 in KCs treated with LPA (30 μM 12 h) ( n = 3). E, F The effects of exogenous LPA (30 μM 12 h) on the protein expression of inflammatory factors IL-1β, NLRP3, COX2, IL-1α and IL-6 in KCs ( n = 3). G Experimental design of mice treated with LPA. H, I The contents of LDH, IL-6 and IL-1βin plasma of LPA treated with mice ( n = 6). J, K Representative images and quantification of HE staining (Scale bars represent 100 μm) and CD68 IHC (Scale bars represent 200 μm) after LPA treatment in liver of mice ( n = 3). Data represents mean ± SEM. * P < 0.05, ** P < 0.01, *** P < 0.001.
Index in PubMed under a CC BY license. PMID: 36964139
Click image to see more details
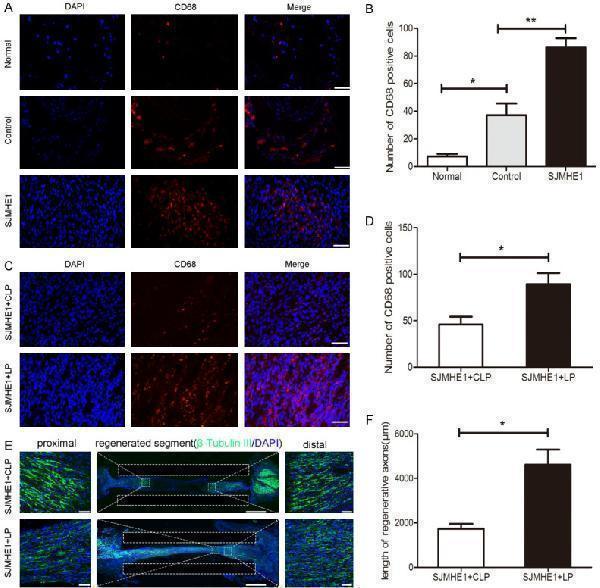
Chlorophosphate liposomes reduced the therapeutic effect of SJMHE1 in an in vivo model. A. Immunofluorescence staining of cross-sections of the regenerated nerve for CD68 (red) 35 days after nerve injury. n = 3. Scale bar = 50 μm. B. Quantitative analysis of the number of CD68-positive cells in different groups. C. Immunofluorescence staining for CD68 (red) in longitudinal sections of the regenerated nerve 10 days after nerve injury. CLP: chlorophosphate liposome. LP: liposome (negative control). n = 6. Scale bar = 50 μm. D. Quantitative analysis of the number of CD68-positive cells in different groups. E. Immunofluorescence staining for β-tubulin III (green) in longitudinal sections of the regenerated nerve. The length of regenerative axons from the proximal end was measured for different groups. Scale bar = 2000 μm. The boxed areas are higher-magnification images. Scale bar = 500 μm. DAPI (blue)-stained nuclei. F. Statistical analysis of the length of regenerative axons. SJMHE1+CLP vs SJMHE1+LP; n = 6 for each group; * P <0.05, ** P <0.01 (Student’s t test; ANOVA). The data are presented as the mean ± SEM.
Index in PubMed under a CC BY license. PMID: 33841657
Click image to see more details
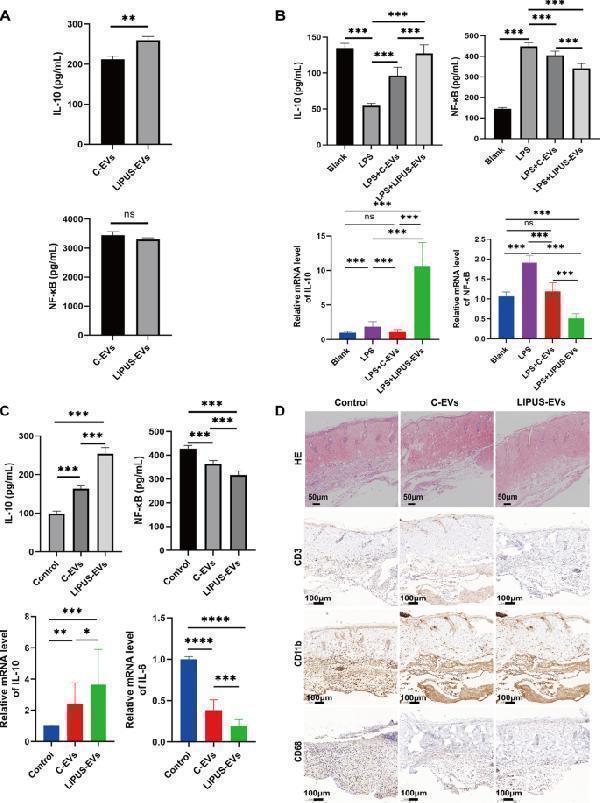
LIPUS strengthened the anti-inflammatory effect of BMSCs-derived EVs. A The protein expression levels of pro-inflammatory cytokines (IL-10 and NF-κB) in both C-EVs and LIPUS-EVs detected by ELISA assay ( n = 6 per group). B The protein expression of IL-10 and NF-κB in RAW264.7 cells measured by ELISA. And the mRNA level of IL-10 and NF-κB in RAW264.7 cells by qRT-PCR. ( n = 6 per group). C The protein expression of IL-10 and NF-κB in skin allografts measured by ELISA. The mRNA expression of IL-10 and IL-6 in skin allografts measured by qRT-PCR. ( n = 4 per group). D H&E-stained sections of skin allograft. Scale bar = 50 μm. Immunohistochemistry staining of the skin allograft in control group contained massive infiltrates of CD3 + , CD11b + and CD68 + cells in comparison to the C-EVs and LIPUS-EVs group. (Representative images from 3 different mice per group). Scale bar = 100 μm. Error bars represent mean ± SD. ns p > 0.05, * p < 0.05, ** p < 0.01, *** p < 0.001. Statistical significance assessed by unpaired two-tailed t -test. EVs, extracellular vesicles
Index in PubMed under a CC BY license. PMID: 36717768
Click image to see more details
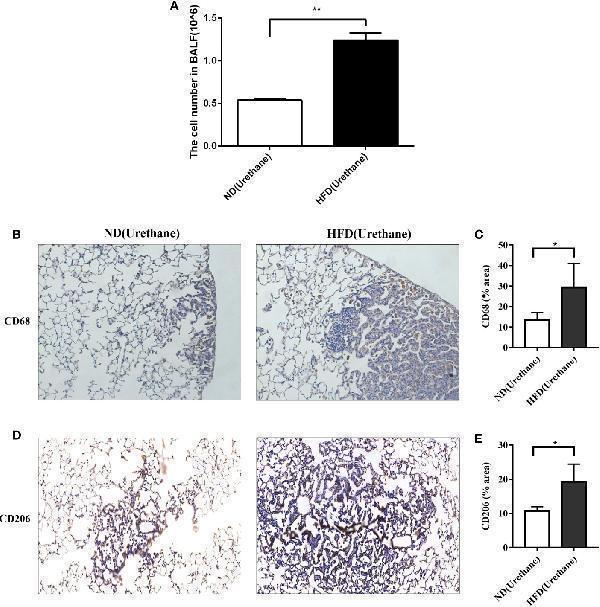
Infiltration of immune cells in bronchoalveolar lavage fluids and lung tissue. (A) Increased BALF cell count in urethane treated high-fat diet fed mice compared to urethane treated control diet fed mice (n=3 mice/group). **P < 0.01. (B, C) Increased expression of CD68 in lung tissue of urethane treated HFD-fed mice compared to urethane treated ND-fed mice. (B) Representative images of lung sections in urethane treated mice. Brown dots indicate the CD68 positive macrophages. (Original magnification: ×200). (C) Quantification analysis of immunohistochemistry related to CD68 between two diet groups by ImageJ. (n=4 mice/ group). *P < 0.05. (D, E) Increased expression of CD206 in lung tissue of urethane treated HFD-fed mice compared to urethane treated ND-fed mice. (D) Representative images of lung sections in urethane treated mice. Brown dots indicate the CD206 positive macrophages. (Original magnification: ×200). (E) Quantification analysis of immunohistochemistry related to CD206 between two diet groups by ImageJ. (n =4-5 mice/ group). *P < 0.05.
Index in PubMed under a CC BY license. PMID: 33708630
Click image to see more details
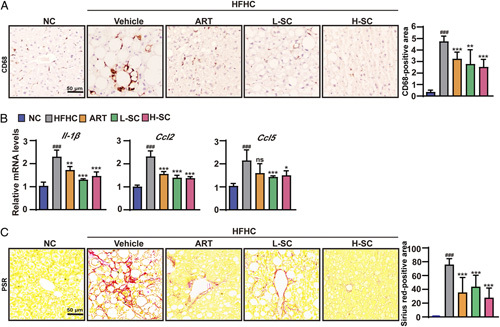
Sodium cholate suppresses hepatic inflammation and fibrosis in mice fed an HFHC diet. (A) Immunohistochemical staining of CD68 in the livers of the indicated mice fed the normal chow or HFHC diet for 20 weeks. n=5 per group. Scale bar, 50 μm. (B) Quantitative real-time PCR analysis of the transcript levels of genes related to inflammation ( Il-1β, Ccl2 , and Ccl5 ). n=5 per group. (C) Representative images showing PSR staining in the livers of the indicated mice fed the normal chow or HFHC diet for 20 weeks. n=5 per group. Scale bar, 50 μm. The data are presented as the mean±SD. #indicates a significant difference between the NC group and the HFHC group ( t test); *indicates a significant difference between the L-SC (Low dose-Sodium Cholate: 90 mg/kg)/(High dose-Sodium Cholate: 180 mg/kg)/ART group and the HFHC group (one-way ANOVA). ### p <0.001 versus NC mice; * p <0.05, ** p <0.01, *** p <0.001 versus mice fed by HFHC. Abbreviations: ART, atorvastatin; CCL2, C-C motif chemokine ligand 2; CCL5, C-C motif chemokine ligand 5; CD68, fatty acid translocase CD68; HFHC, high-fat and high-cholesterol; NC, normal chow; NS, no significance; PSR, picrosirius red.
Index in PubMed under a CC BY license. PMID: 36706173
Click image to see more details
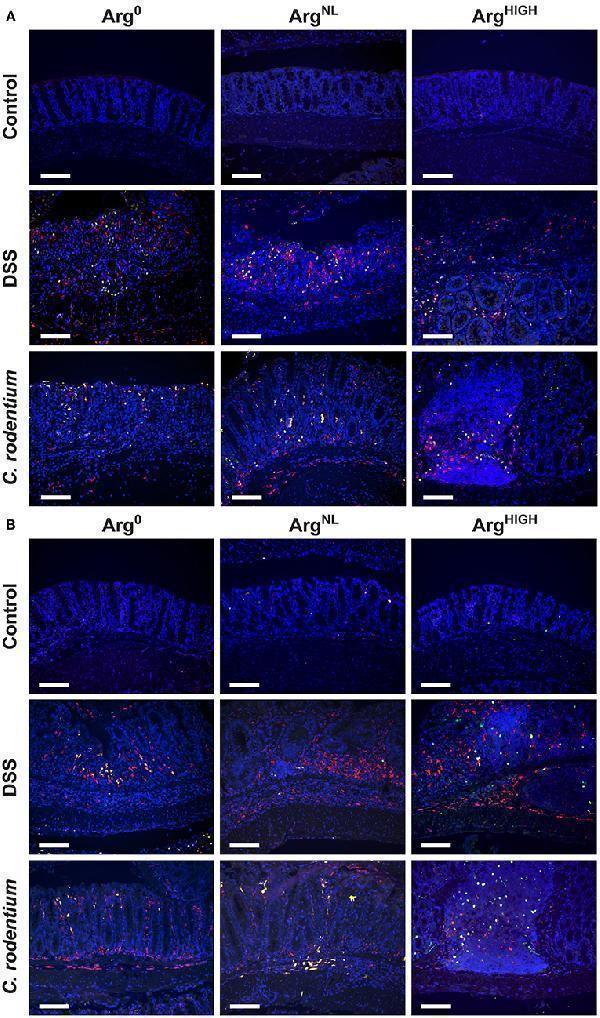
Regulation of NOS2 and arginase-1. NOS2, arginase-1, and CD68 were immunodetected in colonic tissues from C57BL/6 mice ± DSS or C. rodentium ± Arg 0 , Arg NL , or Arg HIGH diets. In each panel, CD68 is depicted in red, NOS2 (A) or arginase-1 (B) in green, and the nuclei in blue; CD68 + NOS2 + and CD68 + ARG1 + cells are shown in yellow. The data shown are representative photomicrographs of at least 3 animals per condition. Scale bar, 50 μm.
Index in PubMed under a CC BY license. PMID: 30972302
Click image to see more details
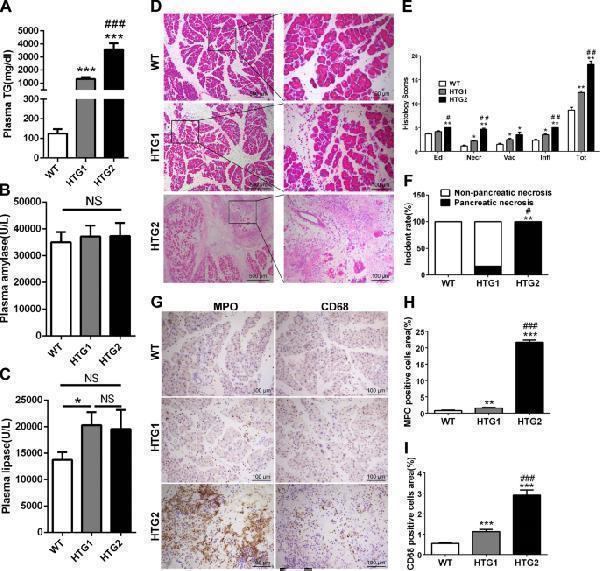
The comparison of pancreatic injury and inflammation between Gpihbp1−/− mice with different HTG severity in caerulein-induced acute pancreatitis. a Triglyceride concentration of plasma before caerulein treatment between wild-type, HTG1 (triglyceride concentration, 1000–2000 mg/dL) and HTG2 mice(triglyceride concentration, å 2000 mg/dL). b , c Plasma amylase and lipase activity at 12 th hour after the first injection of caerulein (50 μg/kg, 10 times, hourly) between these three groups. d Representative photomicrographs of H&E-stained section of pancreas in wild-type, HTG1 and HTG2 mice after acute pancreatitis induction. Scale bar = 500 or 100 μm. e Pathological scores of the pancreas in wild-type, HTG1 and HTG2 mice after acute pancreatitis induction. f Incidence rates of pancreatic necrosis between these three groups after acute pancreatitis induction. Each value was the mean ± SEM for n = 6. g Immunohistochemistry evaluation for myeloperoxidase and CD68 in pancreas between wild-type, HTG1 and HTG2 mice after acute pancreatitis induction. Scale bar = 100 μm. i Semiquantitative results of the area ratio of myeloperoxidase and CD68-positive cells among wild-type, HTG1 and HTG2 mice after acute pancreatitis induction. Each value was the mean ± SEM for n = 3–5. * p < 0.05 or ** p < 0.01 or *** p < 0.001 vs wild-type group. # p < 0.05 or ## p < 0.01 or ### p < 0.001 vs HTG1 group. WT wild-type, TG triglyceride, Ed edema, Necr necrosis, Vac vacuolisation, Infl inflammation, FFAs free fatty acids, MPO myeloperoxidase
Index in PubMed under a CC BY license. PMID: 31570698
Click image to see more details
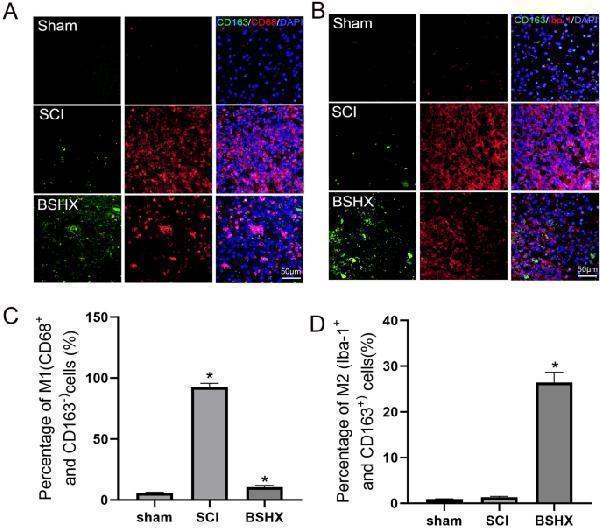
BSHX decoction promoted microglia polarization from M1 to M2. A Immunofluorescent staining of CD68 (red)/CD163 (green) in the lesion site of the spinal cord 14 days after SCI. B Immunofluorescent staining of Iba-1 (red)/CD163 (green) in the lesion site of the spinal cord 14 days after SCI. C , D Quantification the number of M1 (CD68 + /CD163 − ) or M2 (Iba-1 + /CD163 + ) cells in spinal cord. All experiments were performed in triplicated and data were presented as means ± SD, n = 3 per group. * P < 0.05, ** P < 0.01
Index in PubMed under a CC BY license. PMID: 35820953
Click image to see more details
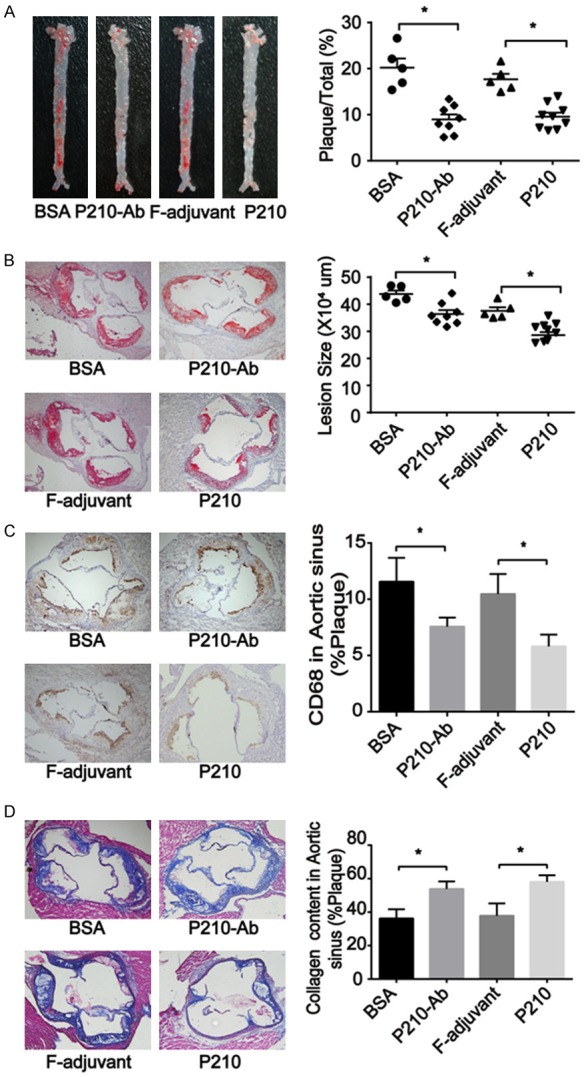
P210-Ab reduced atherosclerotic lesion and affected macrophages and the content of collagen in the aortic sinus. (A, B) The inhibitory effect of P210-Ab on the atherosclerosis of ApoE -/- mice at the age of 23 weeks. Representative images of en face (A) and aortic sinus sections (B) by Oil-red-O staining. Quantitative analyses of en face and aortic sinus sections (100 ×) located in the right column. (C, D) Representative images of aortic sinus section after immunohistochemistry for CD68 (C) (100 ×) and Masson staining (D) (100 ×). Quantitative analyses of CD68 and collagen in the aortic sinus. Data are expressed as mean ± SEM; n = 5, 8, or 9; * P <0.05.
Index in PubMed under a CC BY license. PMID: 30018722
Click image to see more details
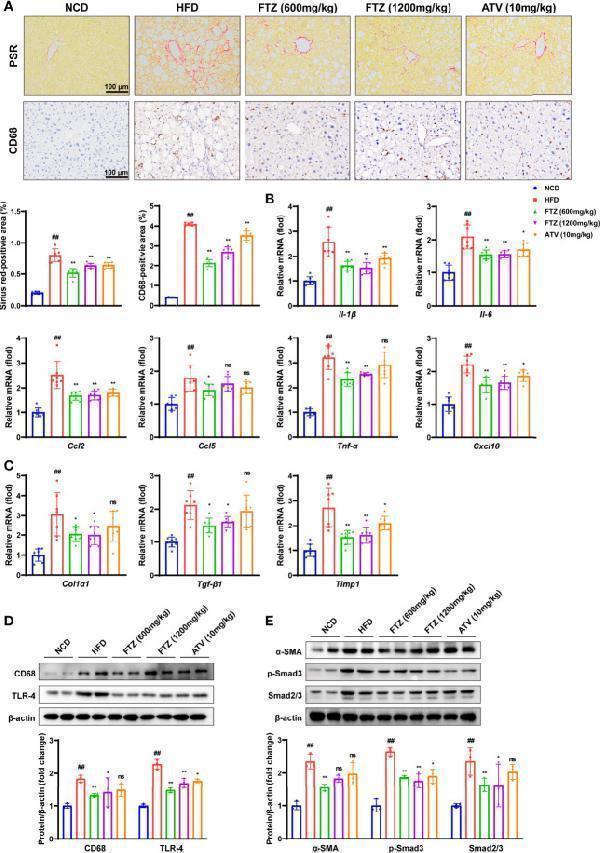
Effects of FTZ on metabolic inflammation and fibrosis in the livers of mice. (A) Representative images showing PSR and immunohistochemical staining of CD68 staining in the livers of the indicated mice (n=6). (B) Relative mRNA levels of inflammatory genes in the livers of the indicated mice (n=7–8). (C) Relative mRNA levels of profibrotic genes in the livers of the indicated mice (n=7-8). (D) The expression of CD68, TLR-4, and β-actin was analyzed by Western blotting. β-actin served as a loading control (n=3). (E) The expression of α-SMA, p-Smad3, Smad2/3, and β-actin was analyzed by Western blotting. β-actin served as a loading control (n=3). Data are represented as means ± SEM. # indicates a significant difference between the NCD group and the HFD group (t-test); * indicates a significant difference between the FTZ (600 mg/kg)/FTZ (1,200mg/kg)/ATV (10 mg/kg) group and the HFD group (one-way ANOVA). ## P < 0.01 versus NCD mice; * P < 0.05, ** P < 0.01 versus mice fed by HFD. ns indicates no significance.
Index in PubMed under a CC BY license. PMID: 35784533
Click image to see more details
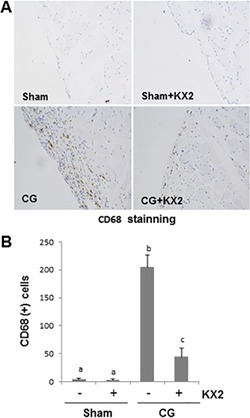
KX2-391 inhibits peritoneal macrophage infiltration in the injured peritoneum. Peritoneal membrane was collected at 21 days after CG injury with or without administration of KX2-391(KX2) ( A , B ). (A) Photomicrographs illustrate immunohistochemical staining of CD68 in the submesothelial compact zone. (B) The number of CD68-positive cells (yellow) was calculated from 10 random fields (200 X) of six rat peritoneal samples. Data are means ± S.E.M. ( n = 6). Bars with different letters (a–c) are significantly different from one another ( P < 0.05).
Index in PubMed under a CC BY license. PMID: 29137389
Click image to see more details
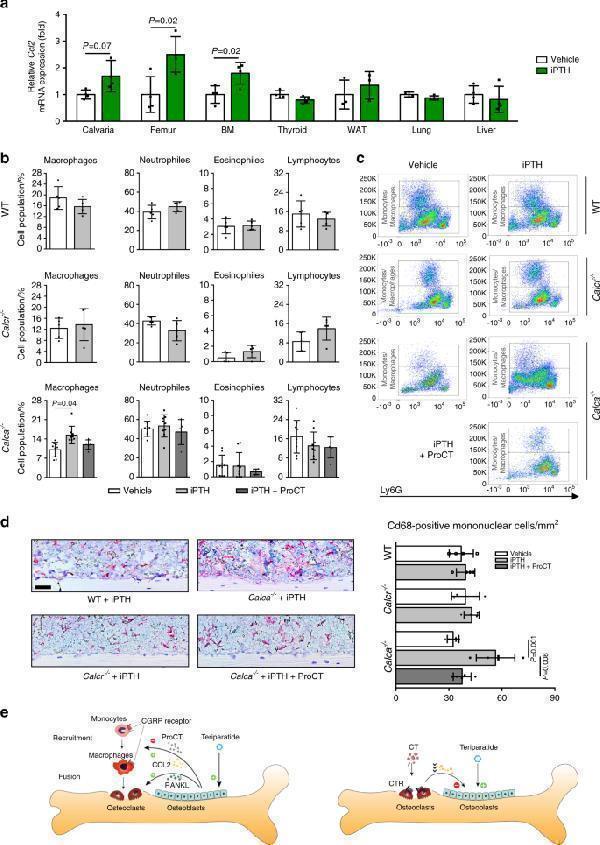
Mice lacking ProCT display increased numbers of macrophages in the bone marrow after iPTH treatment. a Ccl2 expression levels (fold differences) in the indicated tissues following 4 weeks of iPTH treatment (100 μg·kg −1 ). BM = whole bone marrow. n = 3-4 mice per group (one-way ANOVA followed by Tukey’s post hoc test). b Flow cytometric analyses and ( c ) representative flow cytometry plots of the indicated cell populations in flushed bone marrow from mice of the indicated genotypes following 4 weeks of iPTH (100 μg·kg −1 ) and/or ProCT (10 μg·kg −1 ) treatment. Please also refer to Supplementary Fig. for the gating strategy. n = 5–9 mice per group. For Calca −/− mice, the data were pooled from two independent experiments. d Representative images of immunohistochemical staining of tibia sections derived from the same mice using a Cd68-specific antibody. Scale bar = 100 μm. Quantification of intramedullary and endocortical Cd68-positive cells per bone marrow area is displayed on the right. n = 4-5 mice per group. In ( b – d ), one-way ANOVA followed by Tukey’s post hoc test was applied. e Schematic representation of the proposed function of Calca -derived peptides in regulating the therapeutic effect of iPTH. Left: iPTH (teriparatide) induces ProCT expression in osteoblasts, which impairs the recruitment of osteoclast precursors (i.e., monocytes and macrophages) to the bone surface and thus inhibits bone resorption. Right: Circulating CT impairs the osteoanabolic effect of iPTH based on tonic inhibition of bone formation, presumably via osteoclasts
Index in PubMed under a CC BY license. PMID: 35087025

Click image to see more details
Effect of Sirt6 on the inflammation of BV2 cells treated with OA&PA for 24 h. A Expression of M1-type microglia surface markers ( Cd16, Cd86 ) and M2-type microglia surface markers ( Ym1/2 ) in BV2 cells while Sirt6 was knocked down. B Expression of M1-type microglia surface markers ( Cd16, Cd86 ) and M2-type microglia surface markers ( Ym1/2 ) in BV2 cells while Sirt6 was over-expressed. C mRNA levels of Tnf-α , Il-6 , and Il-1β in BV2 cells while Sirt6 was over-expressed. D Inflammatory cytokine (TNF-α, IL-6, and IL-1β) content in culture medium while Sirt6 was over-expressed. E, F Immunofluorescence images of CD68 and TNF-α while Sirt6 was over-expressed. Scale bars: 10 μm. G, H Quantitative analysis of immunofluorescent intensity of CD68 and TNF-α while Sirt6 was over-expressed. A-D, G-H Two-way ANOVA analysis was performed, followed by the Tukey post hoc test. n = 4–8/group. Data are presented as mean ± SEM, * p < 0.05, ** p < 0.01, *** p < 0.001
Index in PubMed under a CC BY license. PMID: 37582706
Click image to see more details
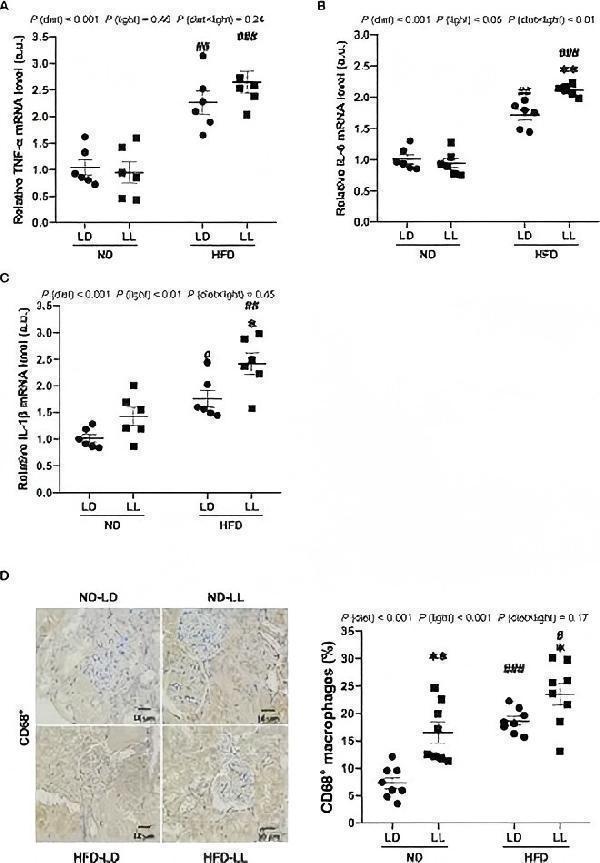
Constant light exposure promotes renal inflammation in HFD-fed rats. (A–C) Renal mRNA expression of TNF-α , IL-6 and IL-1β in rats. (D) IHC staining of CD68 and quantification of CD68 + macrophages. Values represent mean ± SEM (n=6 for A–C , n=8 for D ). Differences were determined using a two-way ANOVA followed by a Bonferroni post hoc analysis. # p <0.05, ## p <0.01, ### p <0.001, vs. ND counterpart. * p <0.05, ** p <0.01, vs. LD counterpart. p (diet), main effect of diet; p (light), main effect of light; p (light × diet), interaction effect of light and diet.
Index in PubMed under a CC BY license. PMID: 35966094
Click image to see more details
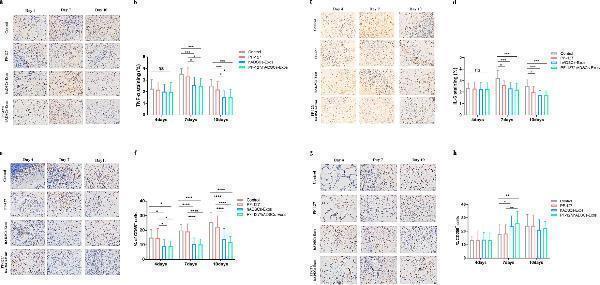
PF-127/hADSCs-Exos complex treatment inhibits inflammatory reaction. a Representative images of TNF-α immunostaining at 4, 7, and 10 days after treatment. Scale bar = 20 µm. b Quantification of TNF-α + IHC stained tissues. c Representative images illustrating IHC results of IL-6 at 4, 7, and 10 days after surgery. Scale bar = 20 µm. d Quantification of IL-6 + IHC stained tissues. e IHC images of wound sections stained with CD68 on days 4, 7, and 10 post-wounding. Scale bar = 20 µm. f Quantification of the number of CD68 positive cells in the wound area on days 4, 7, and 10. g IHC images of wound sections stained with CD206 at days 4, 7, and 10 post-wounding. Scale bar = 20 µm. h Quantification of the number of CD206 positive cells in the wound area on days 4, 7, and 10. In b, d, and f , data are shown as mean ± SEM; n = 6 for each group. * p < 0.05, ** p < 0.01, *** p < 0.001, and **** p < 0.0001 versus vehicle control group
Index in PubMed under a CC BY license. PMID: 35941707
Click image to see more details
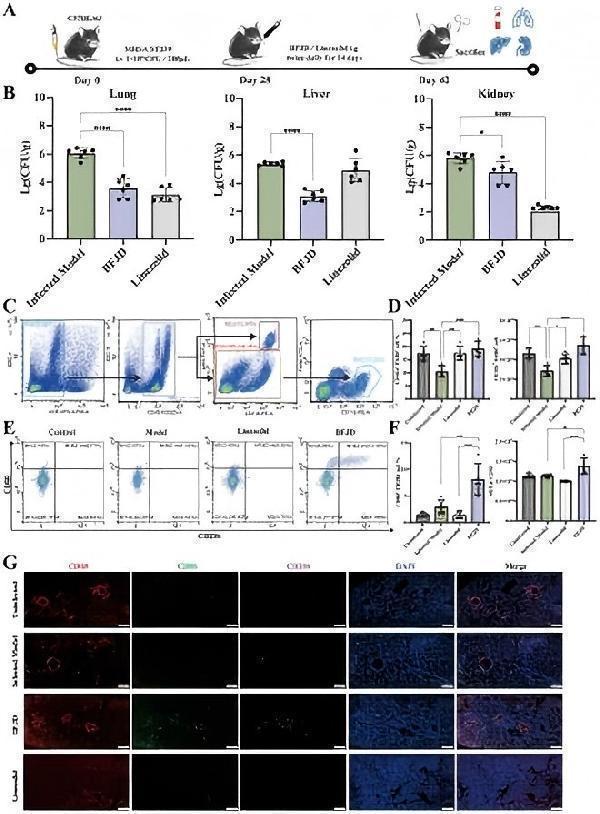
BFJD promotes the clearance of persistent MRSA infection and macrophage M1 polarization in vivo . (A) Schematic illustration of a 42-day long-term persistent MRSA infection model. Mice were inoculated with 1×10 8 CFU of MRSA in 0.2 ml of PBS via a lateral tail vein. At day 28 post- MRSA inoculation, mice were administered BFJD and linezolid by oral gavage intervention, and after 14 days of treatment, the mice were euthanized for sampling. (B) Bacterial burdens in the lungs, liver, and kidneys of mice (n=6). (C) The gating strategy of macrophages (CD11b+F4/80+) in mouse lung tissue flow cytometry. (D) The percentage of macrophages in the Ly6G - cell population and relative count in total harvested cells (n=5). (E) Representative FACS plots of macrophage polarization. (F) The percentage of M1- type (CD80+CD206-) macrophages and MFI of CD80 (n=5). (G) Representative immunofluorescence micrograph of lung tissue stained with indicated antibodies for CD68, CD86 and CD206. Nuclei were revealed by DAPI staining. Scale bars, 200 μm. Data are presented as the mean ± SD. Differences were analyzed applying ordinary one-way ANOVA followed by Dunnett´s multiple comparisons test (comparison with the Infected Model). * P < 0.05, ** P < 0.01, *** P < 0.001, **** P < 0.0001.
Index in PubMed under a CC BY license. PMID: 40746551
Click image to see more details
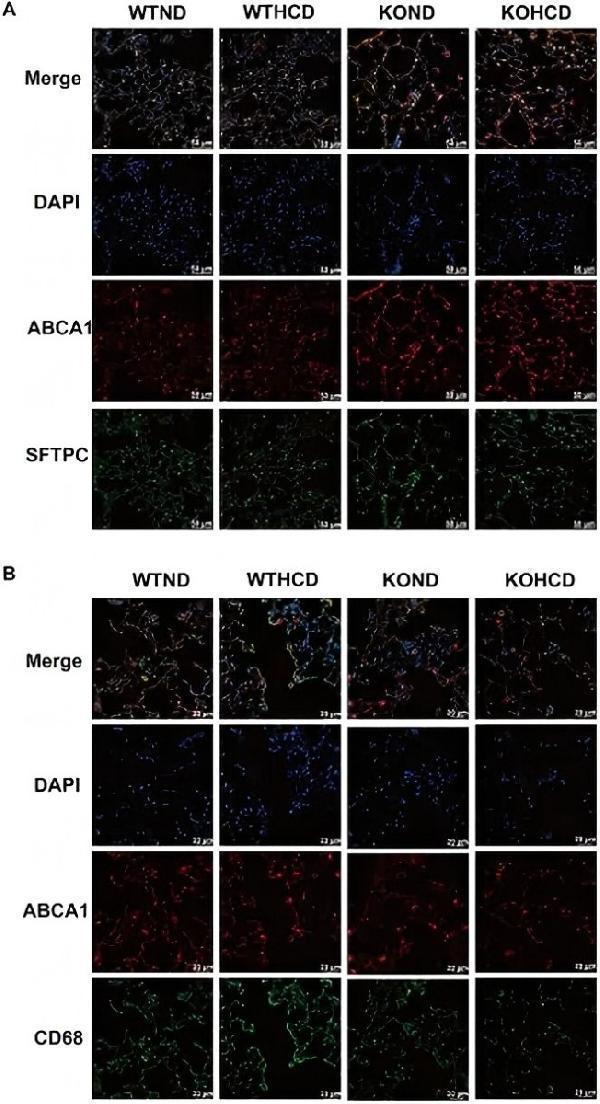
Subcellular localization of ABCA1 in lung tissue. (A) Co-staining of ABCA1 (red) and SFTPC (green, alveolar type II marker); nuclei stained with DAPI (blue). (B) Co-staining of ABCA1 (red) and CD68 (green, macrophage marker) to determine cell-type-specific expression. n = 3–5 mice/group.
Index in PubMed under a CC BY license. PMID: 40808841

Click image to see more details
BSHX decoction promoted remyelination activity. Immunofluorescent staining of activated microglia (CD68 positive, red) and oligodendrocytes (MBP positive, green) in the lesion site of the spinal cord 14 days after SCI
Index in PubMed under a CC BY license. PMID: 35820953
Specific Publications For Anti-Macrosialin CD68 Antibody Picoband® (PA1518)
Loading publications
Recommended Resources
Here are featured tools and databases that you might find useful.
- Boster's Pathways Library
- Protein Databases
- Bioscience Research Protocol Resources
- Data Processing & Analysis Software
- Photo Editing Software
- Scientific Literature Resources
- Research Paper Management Tools
- Molecular Biology Software
- Primer Design Tools
- Bioinformatics Tools
- Phylogenetic Tree Analysis
Customer Reviews
Have you used Anti-Macrosialin CD68 Antibody Picoband®?
Share your experimental results or join a short interview to earn up to $1,000 in product credits or other rewards.
0 Reviews For Anti-Macrosialin CD68 Antibody Picoband®
Customer Q&As
Have a question?
Find answers in Q&As, reviews.
Can't find your answer?
Submit your question
10 Customer Q&As for Anti-Macrosialin CD68 Antibody Picoband®
Question
What Are CD68 Macrophages?
Verified customer
Asked: 2021-10-20
Answer
CD68 is a protein highly expressed by cells in the monocyte lineage by circulating macrophages, and by tissue macrophages such as Kupffer cells, microglia.
Boster Scientific Support
Answered: 2021-10-20
Question
Our team were content with the WB result of your anti-CD68 antibody. However we have seen positive staining in leukocyte isoform short: cell membrane using this antibody. Is that expected? Could you tell me where is CD68 supposed to be expressed?
Verified Customer
Verified customer
Asked: 2020-02-10
Answer
According to literature, leukocyte does express CD68. Generally CD68 expresses in isoform short: cell membrane. Regarding which tissues have CD68 expression, here are a few articles citing expression in various tissues:
Adipose tissue, Pubmed ID: 16625196
Liver, Pubmed ID: 19159218
Platelet, Pubmed ID: 16263699
Skin, Pubmed ID: 15489334
Spleen, and Umbilical cord blood, Pubmed ID: 14702039
Boster Scientific Support
Answered: 2020-02-10
Question
What tissue types is PA1518 known to stain for?
Verified customer
Asked: 2019-12-04
Answer
The Anti-Macrosialin CD68 Antibody (PA1518) stains the following tissue types: Macrophage Dendritic cells Brain Spleen Mammary gland
Boster Scientific Support
Answered: 2019-12-04
Question
We have seen staining in mouse adipose tissue. What should we do? Is anti-CD68 antibody supposed to stain adipose tissue positively?
Verified Customer
Verified customer
Asked: 2019-12-03
Answer
From what I have seen in literature adipose tissue does express CD68. From what I have seen in Uniprot.org, CD68 is expressed in leukocyte, spleen umbilical cord blood, adipose tissue, skin, platelet, liver, among other tissues. Regarding which tissues have CD68 expression, here are a few articles citing expression in various tissues:
Adipose tissue, Pubmed ID: 16625196
Liver, Pubmed ID: 19159218
Platelet, Pubmed ID: 16263699
Skin, Pubmed ID: 15489334
Spleen, and Umbilical cord blood, Pubmed ID: 14702039
Boster Scientific Support
Answered: 2019-12-03
Question
Which amino acid range is the immunogen sequence of PA1518 from?
Verified customer
Asked: 2019-11-29
Answer
For the Anti-Macrosialin CD68 Antibody (PA1518), a synthetic peptide corresponding to a sequence in the middle region of mouse CD68 (312-326aa AFCITRRRQSTYQPL), different from the related rat sequence by one amino acid.
Boster Scientific Support
Answered: 2019-12-02
Question
Has this antibody been tested for cross-reactivity with human samples in IHC or Western Blot?
Verified Customer
Verified customer
Asked: 2019-11-29
Answer
We tested human samples and got negative results in IHC and WB.
Boster Scientific Support
Answered: 2019-11-29
Question
Do you have an IHC application on Ventana machine for PA1518?
Verified customer
Asked: 2019-07-03
Answer
Our lab hasn't used the Ventana machine for our antibodies. We suggest to watch some on-line vedio or read some related articles. http://www.path.queensu.ca/qlmp/sites/webpublish.queensu.ca.qlmpwww/files/files/Ventana%20Pamphlet.pdf
Boster Scientific Support
Answered: 2019-07-05
Question
Our lab used your anti-CD68 antibody for WB on spleen umbilical cord blood in the past. I am using mouse, and I plan to use the antibody for IF next. My lab would like examining spleen umbilical cord blood as well as adipose tissue in our next experiment. Do you have any suggestion on which antibody would work the best for IF?
Verified Customer
Verified customer
Asked: 2018-05-18
Answer
I looked at the website and datasheets of our anti-CD68 antibody and it seems that PA1518 has been validated on mouse in both WB and IF. Thus PA1518 should work for your application. Our Boster satisfaction guarantee will cover this product for IF in mouse even if the specific tissue type has not been validated. We do have a comprehensive range of products for IF detection and you can check out our website bosterbio.com to find out more information about them.
Boster Scientific Support
Answered: 2018-05-18
Question
Our lab want to know about using your anti-CD68 antibody for autocrine signaling studies. Has this antibody been tested with western blotting on rat spleen tissue? We would like to see some validation images before ordering.
R. Evans
Verified customer
Asked: 2016-09-23
Answer
We appreciate your inquiry. This PA1518 anti-CD68 antibody is tested on rat spleen tissue, tissue lysate, mouse spleen tissue. It is guaranteed to work for IF, IHC-P, IHC-F, WB in mouse, rat. Our Boster guarantee will cover your intended experiment even if the sample type has not been be directly tested.
Boster Scientific Support
Answered: 2016-09-23
Question
We are currently using anti-CD68 antibody PA1518 for rat tissue, and we are satisfied with the IHC-F results. The species of reactivity given in the datasheet says mouse, rat. Is it possible that the antibody can work on zebrafish tissues as well?
K. Carter
Verified customer
Asked: 2016-01-08
Answer
The anti-CD68 antibody (PA1518) has not been validated for cross reactivity specifically with zebrafish tissues, but there is a good chance of cross reactivity. We have an innovator award program that if you test this antibody and show it works in zebrafish you can get your next antibody for free. Please contact me if I can help you with anything.
Boster Scientific Support
Answered: 2016-01-08





