This website uses cookies to ensure you get the best experience on our website.
- Table of Contents

Antigen retrieval is a vital technique in immunohistochemistry (IHC) that enhances the visibility of antigens in formalin-fixed, paraffin-embedded (FFPE) tissue samples. This process is especially critical in Multiplex Assay workflows, where multiple antigens must be unmasked uniformly for accurate, simultaneous detection. During fixation, formaldehyde creates cross-links that mask epitopes—the specific regions of antigens recognized by antibodies. This masking impairs antibody access to these critical sites, limiting effective binding and impacting staining results. To ensure optimal detection of gene expression in FFPE tissues, especially in conjunction with antigen retrieval, downstream techniques like QPCR Services can validate the presence and quantity of target transcripts.
To address this, antigen retrieval methods are employed to reverse the cross-linking and restore antigenicity. These techniques allow antibodies to bind more effectively to their target antigens, improving sensitivity and accuracy in detection. While fixation is essential for preserving tissue morphology, it can negatively affect protein biochemistry, altering the conformation or electrostatic charge of epitopes and further hindering antibody interaction. Thus, antigen retrieval is necessary to ensure reliable diagnostic and research outcomes by overcoming the challenges posed by the fixation process. This step becomes even more critical when working with a multiplex IHC service, where consistent epitope exposure is essential for accurate multi-target detection.
Two common methods to retrieve antigens are heat-induced epitope retrieval (HIER) and proteolysis-induced epitope retrieval (PIER).
HIER is the most widely used technique for antigen retrieval. This method utilizes heat to reverse the cross-linking that occurs during tissue fixation. Typically, tissue samples are immersed in a buffer solution, such as sodium citrate or EDTA, and subjected to heating using a microwave, pressure cooker, or water bath. The heat denatures proteins and breaks the cross-links, effectively unmasking the epitopes and allowing antibodies to bind more efficiently.
Microwave heat retrieval is highly effective for most antigens. It works in two ways: microwave radiation induces polar movement in molecules, aiding in the breakdown of aldehyde bonds. The rapid heat generated by this molecular motion, upon reaching a sufficient temperature, can denature proteins fixed with formaldehyde. This technique is known for its quick heat generation, simplicity of use, and the possibility of causing the antigen retrieval solution to boil.
Critical factors in HIER include the temperature, pH, and incubation time, all of which must be carefully optimized to ensure proper antigen unmasking without causing morphological damage. Common buffers used in HIER include sodium citrate (10 mM, pH 6), EDTA (1 mM, pH 8), and Tris/EDTA (pH 9), with temperatures typically reaching 95-100 °C.
In contrast to HIER, PIER employs proteolytic enzymes, such as bromelain, aspronase, pepsin, ficin, trypsin, proteinase K, or other proteases, to digest the proteins surrounding the epitopes. This enzymatic treatment selectively breaks down specific protein structures, thereby exposing the masked epitopes. PIER is often considered a gentler approach compared to HIER, making it suitable for fragile tissues. However, careful optimization of enzyme concentration, incubation temperature, and incubation time is required to achieve optimal results.
Choosing the appropriate proteolytic enzymes depends on the specific antigen components being targeted. Typically, pepsin and bromelain are used to detect interstitial antigens like fibronectin, laminin (LN), and various types of collagen. Other proteolytic enzymes can be used to identify intracellular antigens. It's crucial to consider the optimal pH for each enzyme to ensure effective digestion.
2.3 Heat-induced Epitope Retrieval (HIER) vs Proteolytic-induced Epitope Retrieval (PIER)
| heat-induced epitope retrieval (HIER) | proteolytic-induced epitope retrieval (PIER) | |
| Advantage | A broader range of antigens, especially nuclear and high-molecular-weight proteins less likely to disrupt tissue morphology produce less non-specific staining | Preferred for difficult-to-recover epitopes less damaging to delicate or fragile tissues, suitable for certain types of samples |
| Disadvantage | Overheating can damage tissues or lead to loss of antigenicity, while insufficient heating may result in inadequate retrieval | the low success rate for restoring immunoreactivity potential for destroying both tissue morphology and the antigen of interest |
| Difficulties | Unequal retrieval due to unequal heating | Concentration calibration |
The pH of the retrieval solution used in antigen heat retrieval can significantly impact staining results. The effects of pH on staining can generally be classified into four categories:
1. Stable Type: pH has minimal effect on staining results, as seen with PCNA, AE1, EMA, and CD20.
2. V Type: Both high and low pH values yield good staining results, while pH values around 4-5 result in poorer staining, as observed with ER and Ki-67.
3. Increasing Type: Staining results improve progressively with increasing pH, such as with HMB45.
4. Decreasing Type: Staining results weaken as pH increases; this is rare and seen with antibodies like MOC31.
Currently, the most commonly used retrieval solutions are
citrate buffer and EDTA buffer. Studies indicate that for most antibodies, EDTA
(pH 8.0 or 9.0) is more effective than citrate at pH 6.0, especially for
nuclear-positive antibodies. However, due to traditional practices, most hospitals
and laboratories continue to use citrate buffer at pH 6.0.
Optimal retrieval conditions depend on
the type of tissue, fixation, and antibody, necessitating optimization for each
antigen. For labs that lack time or resources for in-house testing, contract IHC services can provide standardized, validated protocols for consistent results. To optimize antigen retrieval, preliminary studies should be conducted
using a matrix of various combinations of time, temperature, and pH.
If staining still appears unexpectedly faint after retrieval optimization, this guide on weak or no staining in IHC is a useful next step for checking whether the main bottleneck comes from fixation, retrieval conditions, antibody performance, or detection sensitivity.
Here is an example of a matrix.
| Time | Antigen Retrieval Solution pH | Antigen Retrieval Solution pH | Antigen Retrieval Solution pH |
|---|---|---|---|
| pH6.0 | pH8.0 | pH9.0 | |
| 4 minutes | Slide #1 | Slide #2 | Slide #3 |
| 8 minutes | Slide #4 | Slide #5 | Slide #6 |
| 12 minutes | Slide #7 | Slide #8 | Slide #9 |
Materials and reagents
Microwave
staining dish
Antigen retrieval buffer (Tris/EDTA pH
9.0, sodium citrate pH 6.0, EDTA pH 8.0, or other)
1.Immerse the slides in staining dish containing antigen retrieval buffer.
2.Microwave the staining dish at 95°C for 8 minutes.
3.Cool the slides for 5 minutes.
4.Microwave the staining dish at 95°C for 4 minutes.
5.Cool the slides to room temperature before proceeding with staining.
Note: Steamer, water bath or
pressure cooker can be used as alternative heating source to replace microwave.
PIER requires careful optimization of
enzyme concentration, incubation temperature, and incubation time to achieve
optimal results. Trypsin is a commonly used enzymatic antigen retrieval
solution. The following protocol uses 0.1% trypsin as an example. Optimal
conditions always have to be determined by each laboratory and in accordance
with the specific product information.
Materials and reagents
37°C incubator
Humidified chamber
Enzymatic antigen retrieval solution
1.Prepare the 0.1% trypsin solution and preheat it to 37°C.
2. Pipette the enzyme solution onto the tissue section.
3. Place the slides in a humidified container and then into a 37°C incubator.
4. After 10-30 minutes, remove the slides and transfer them to a rack in a container of
tap water.
5. Rinse under running water for 3 minutes.
6. Proceed with the immunohistochemical staining.
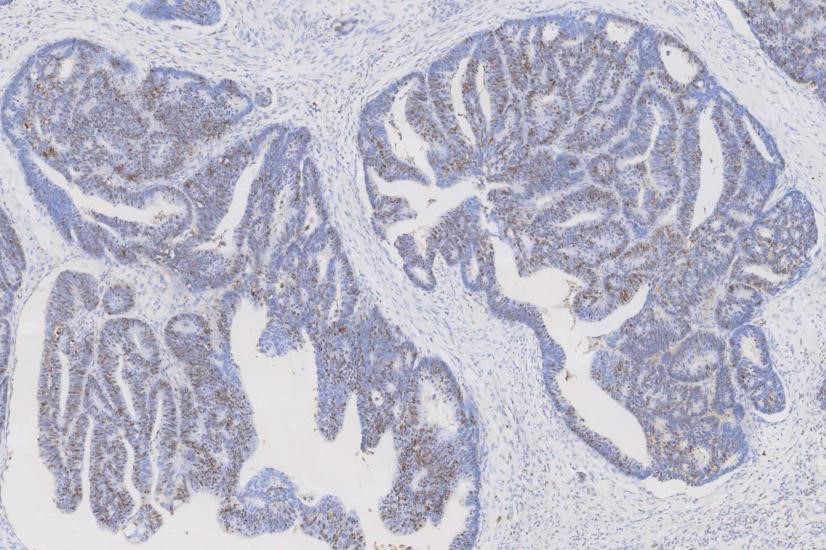
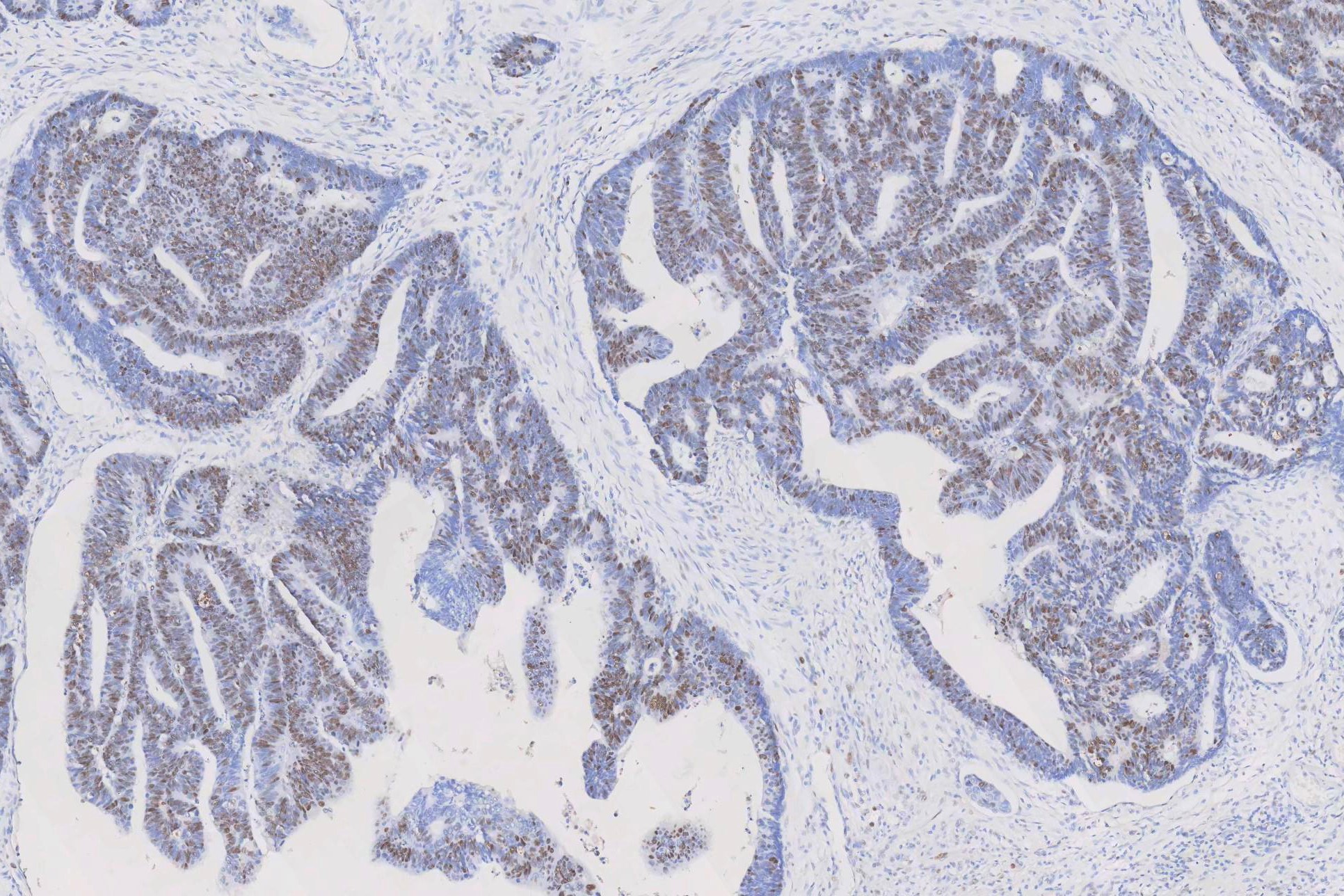
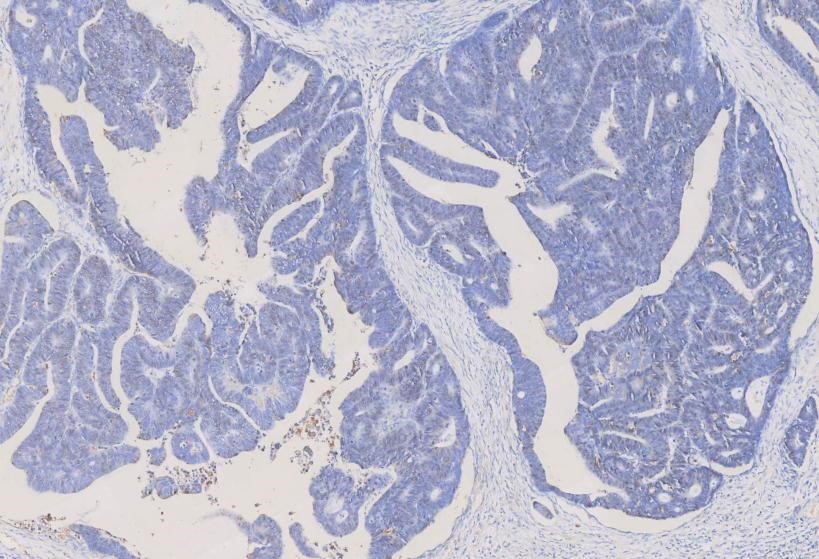
IHC analysis of PCNA using anti-PCNA antibody (MA1083). PCNA was detected in a paraffin-embedded section of human rectal cancer tissue. The tissue section was blocked with 5% BSA. The tissue section was then incubated with 1μg/ml mouse anti-PCNA Antibody (MA1083) overnight at 4°C. Peroxidase Conjugated Goat Anti-Mouse IgG was used as a secondary antibody and incubated for 30 minutes at 37°C. The tissue section was developed using the HRP Conjugated Mouse IgG Super Vision Assay Kit (Catalog # SV0001) with DAB as the chromogen.
When DAB is used as the chromogen, background control becomes a major part of slide interpretability. This practical guide on troubleshooting high background in DAB staining is especially helpful when signal is present but contrast is poor, diffuse staining appears, or the whole slide looks dirty.
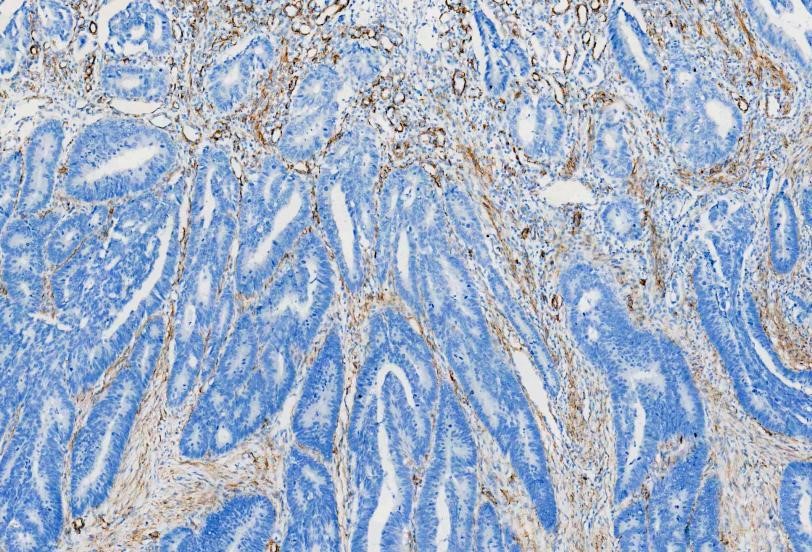
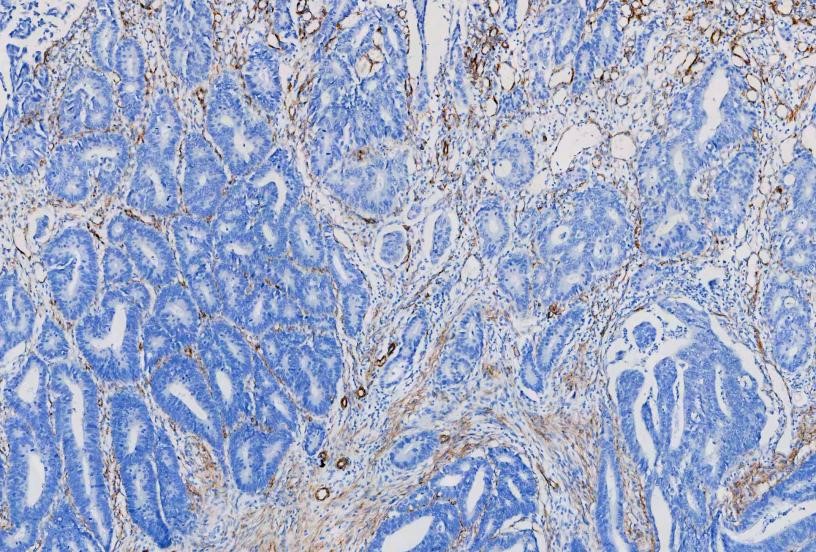
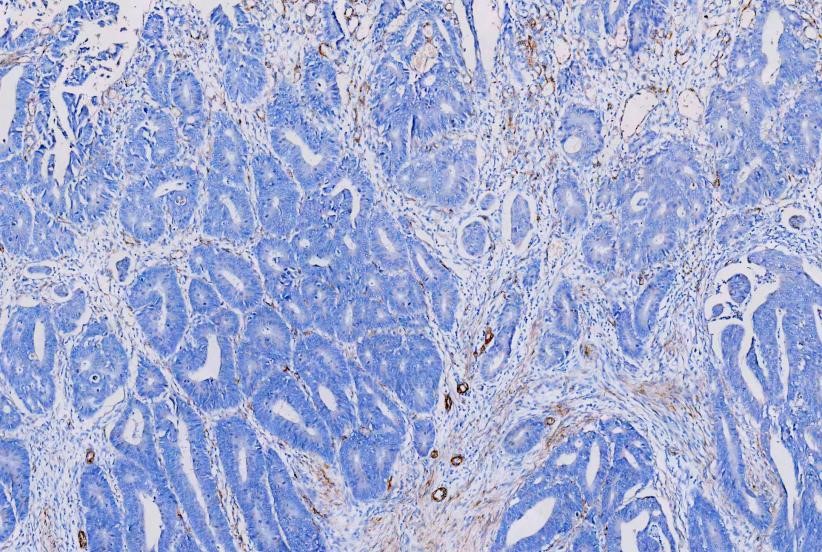
IHC analysis of
Alpha-Smooth Muscle Actin using anti-Alpha-Smooth Muscle Actin antibody
(MA1106). Alpha-Smooth Muscle Actin was detected in a paraffin-embedded section
of human rectal cancer tissues. The tissue section was blocked with 5% BSA. The
tissue section was then incubated with 1μg/ml mouse anti-Alpha-Smooth Muscle
Actin Antibody (MA1106) overnight at 4°C. Peroxidase Conjugated Goat Anti-Mouse
IgG was used as secondary antibody and incubated for 30 minutes at 37°C. The
tissue section was developed using HRP Conjugated Mouse IgG Super Vision Assay
Kit (Catalog # SV0001) with DAB as the chromogen.