This website uses cookies to ensure you get the best experience on our website.
- Table of Contents
and ELISA kits, proteins related to Amyotrophic Lateral Sclerosis.
Amyotrophic Lateral Sclerosis (ALS) is a progressive neurodegenerative disease that affects nerve cells in the brain and spinal cord, leading to the gradual loss of muscle control. Commonly known as Lou Gehrig's disease, ALS disrupts the communication between the brain and muscles, resulting in weakness, paralysis, and eventually, respiratory failure. While the exact cause of ALS remains unclear, genetic and environmental factors are believed to play a role. Currently, there is no cure for ALS, making research into effective treatments and therapies critically important. Advances in biomedical research, particularly in the development of targeted antibodies, offer hope for slowing disease progression and improving the quality of life for those affected. By deepening our understanding of ALS and harnessing innovative scientific approaches, we strive to pave the way toward meaningful breakthroughs in combating this challenging condition.
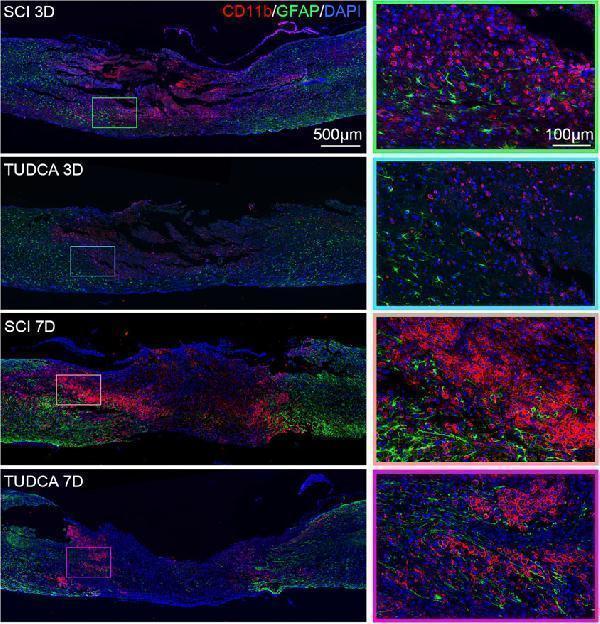
PA1239
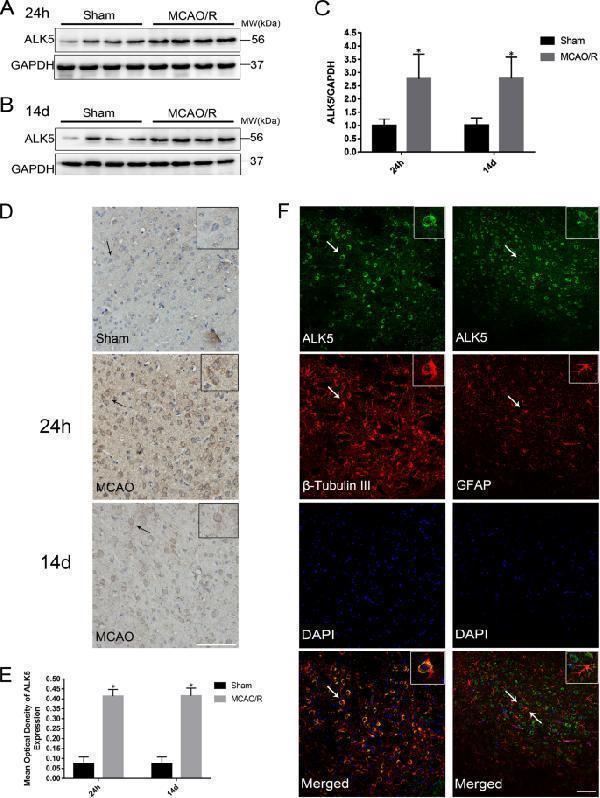
MA1045
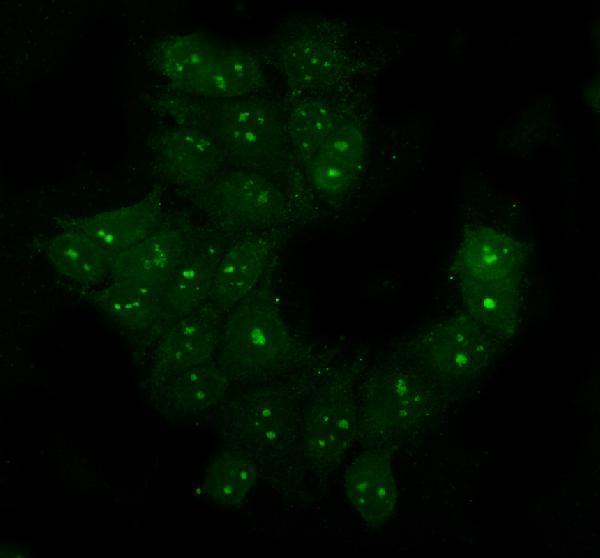
PA1345
| Protein Name | Gene Name | Function |
|---|---|---|
| Neurofilament Light Chain (NfL) | NEFL | Marker of axonal damage and neuronal degeneration. |
| TAR DNA-binding Protein 43 (TDP-43) | TARDBP | Involved in RNA processing; aggregates linked to ALS pathogenesis. |
| Superoxide Dismutase 1 (SOD1) | SOD1 | Enzyme that protects against oxidative stress; mutations associated with familial ALS. |
| C9orf72 | C9orf72 | Genetic expansions linked to ALS and frontotemporal dementia. |
| Fused in Sarcoma (FUS) | FUS | RNA binding protein involved in gene regulation; mutations associated with ALS. |
| Glial Fibrillary Acidic Protein (GFAP) | GFAP | Marker of astrocytic activation and gliosis in neurodegeneration. |
| Ubiquitin Carboxy-terminal Hydrolase L1 (UCHL1) | UCHL1 | Involved in the ubiquitin-proteasome system; implicated in protein degradation. |
| Angiogenin (ANG) | ANG | Promotes angiogenesis and neuronal survival; mutations associated with ALS. |
| Valosin-Containing Protein (VCP) | VCP | Involved in protein homeostasis and autophagy; mutations linked to ALS. |
| S100 Calcium Binding Protein B (S100B) | S100B | Marker of glial activation and neuroinflammation. |
| Chitinase-3-like Protein 1 (CHI3L1) | CHI3L1 | Involved in inflammation and tissue remodeling. |
| Osteopontin (SPP1) | SPP1 | Involved in inflammatory responses and cell signaling. |
| Low-Affinity Nerve Growth Factor Receptor (p75NTR) | NGFR | Regulates neuronal survival and apoptosis. |
| Transactive Response DNA-binding Protein 43 (TDP-43) | TARDBP | RNA metabolism and stress granule formation; pathological aggregation in ALS. |
| TREM2 | TREM2 | Regulates microglial activation and immune responses in the CNS. |
| UNC13A | UNC13A | Associated with neuronal vesicle release and ALS disease progression. |
| Glutamate Transporter 1 (GLT-1) | SLC1A2 | Responsible for glutamate reuptake; dysregulation linked to excitotoxicity in ALS. |
| Methylmalonic Coenzyme A Mutase (MMUT) | MMUT | Involved in mitochondrial metabolism; mutations may contribute to ALS pathology. |
| Brain-Derived Neurotrophic Factor (BDNF) | BDNF | Supports neuron survival and synaptic plasticity; altered levels observed in ALS. |
| Vascular Endothelial Growth Factor (VEGF) | VEGFA | Promotes blood vessel formation; associated with motor neuron survival. |
Amyotrophic Lateral Sclerosis (ALS) exhibits both sporadic and familial forms, with genetic mechanisms playing a crucial role in the latter. Research into the genetic underpinnings of ALS has identified several key mutations, such as those in the SOD1, C9orf72, TARDBP, and FUS genes. The discovery of the C9orf72 hexanucleotide repeat expansion, for instance, has been pivotal in understanding the disease's hereditary aspects and its pathological cascade. These genetic mutations contribute to neuronal degeneration through various pathways, including impaired RNA processing, disrupted protein homeostasis, and dysfunctional cellular transport mechanisms. Studying these genetic factors not only aids in unraveling the complex etiology of ALS but also paves the way for developing targeted therapies and personalized medicine approaches. Additionally, genetic research facilitates the identification of biomarkers for early diagnosis and disease progression, which is essential for improving patient outcomes. By elucidating the hereditary patterns and molecular consequences of these genetic alterations, scientists aim to mitigate the impact of ALS and explore potential avenues for intervention and treatment.
Protein aggregation is a hallmark of ALS pathology, with misfolded proteins accumulating within motor neurons and contributing to cellular dysfunction and death. Key proteins implicated in ALS include TDP-43 and FUS, which normally play roles in RNA metabolism and cellular transport. In ALS, these proteins undergo pathological mislocalization and form insoluble aggregates that disrupt normal cellular functions. The aggregation of these proteins interferes with RNA processing, impairs axonal transport, and induces proteostatic stress, leading to neuronal toxicity. Additionally, impaired protein degradation pathways, such as the ubiquitin-proteasome system and autophagy, exacerbate the accumulation of these toxic aggregates. Understanding the mechanisms behind protein misfolding and aggregation is critical for identifying therapeutic targets aimed at enhancing protein clearance and restoring neuronal health. Research in this area also explores the interplay between protein aggregation and other cellular processes, including mitochondrial dysfunction and oxidative stress, to develop comprehensive strategies for combating ALS. By targeting the underlying cellular mechanisms that drive protein aggregation, scientists hope to halt or reverse the neurodegenerative processes that characterize ALS, ultimately improving the prognosis for affected individuals.