This website uses cookies to ensure you get the best experience on our website.
- Table of Contents
and ELISA kits, proteins related to Colorectal Cancer.
Colorectal cancer is a major health concern worldwide, affecting both the colon and rectum. It arises when abnormal cells in the lining of the large intestine grow uncontrollably, forming malignant tumors. Early detection through regular screenings, such as colonoscopies, significantly improves the chances of successful treatment and survival. Risk factors include age, genetic predisposition, lifestyle choices, and certain medical conditions. Advances in medical research, particularly in the development of targeted antibodies, are revolutionizing the approach to prevention and therapy. These innovative antibody-based treatments offer more precise and effective options for managing colorectal cancer, leading to better patient outcomes and quality of life. Ongoing research continues to uncover new insights, bringing hope for improved diagnostics and personalized treatments in the fight against this pervasive disease.
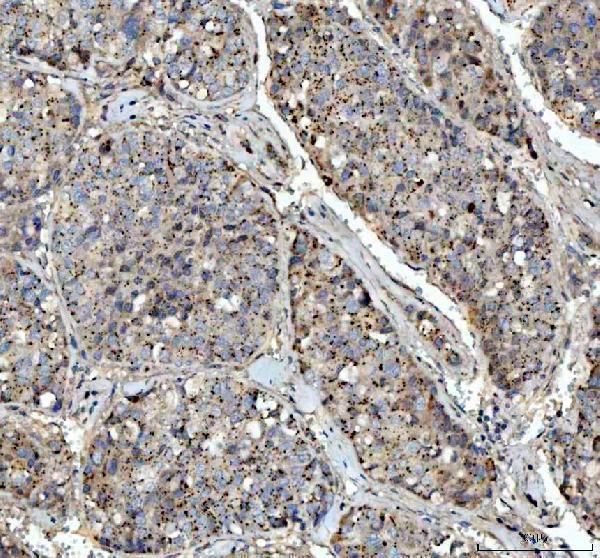
Anti-VEGF/VEGFA Antibody Picoband®, Figure 3. IHC analysis of VEGFA using anti-VEGFA antibody (PA1080).
VEGFA was detected in a paraffin-embedded section of human liver cancer tissue. Hea...
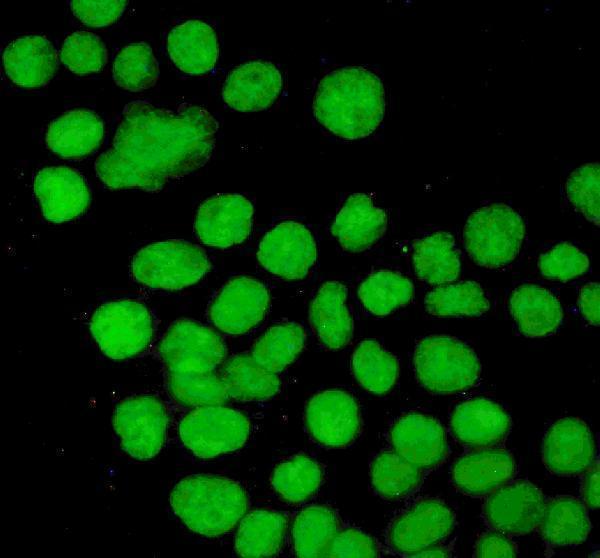
Anti-P53/TP53 Antibody Picoband®, Figure 3. IF analysis of P53 using anti-P53 antibody (PB9008).
P53 was detected in immunocytochemical section of A431 cells. Enzyme antigen...

Anti-P53 Tp53 Monoclonal Antibody, Figure 3. IF analysis of p53 using anti-p53 antibody (MA1078).
p53 was detected in immunocytochemical section of A431 cell. Enzyme antigen retrieva...
| Protein Name | Gene Name | Function |
|---|---|---|
| CEA | CEACAM5 | Tumor marker used for monitoring colorectal cancer treatment and recurrence. |
| KRAS | KRAS | Mutations predict non-responsiveness to EGFR-targeted therapies. |
| BRAF | BRAF | Mutations associated with poor prognosis and resistance to certain therapies. |
| NRAS | NRAS | Mutations indicate resistance to EGFR inhibitors. |
| p53 | TP53 | Tumor suppressor involved in cell cycle regulation; mutations linked to cancer progression. |
| APC | APC | Tumor suppressor gene; mutations lead to uncontrolled cell growth. |
| HER2 | ERBB2 | Overexpression can drive cancer growth; target for specific therapies. |
| VEGF | VEGFA | Promotes angiogenesis; target for anti-angiogenic therapies. |
| EGFR | EGFR | Cell growth receptor; target for EGFR inhibitors. |
| PIK3CA | PIK3CA | Mutations involved in cell growth and survival pathways. |
| PTEN | PTEN | Tumor suppressor regulating cell cycle; loss associated with cancer progression. |
| SMAD4 | SMAD4 | Involved in TGF-beta signaling; mutations linked to poor prognosis. |
| CTNNB1 | CTNNB1 | Encodes beta-catenin, involved in cell adhesion and signaling. |
| PD-L1 | CD274 | Immune checkpoint protein; target for immunotherapy. |
| ALK | ALK | Rearrangements can drive cancer; target for ALK inhibitors. |
| MET | MET | Receptor tyrosine kinase involved in growth and metastasis. |
| HER3 | ERBB3 | Partner for HER2 signaling; potential therapeutic target. |
| FGFR2 | FGFR2 | Receptor involved in cell differentiation and proliferation. |
| ALK | ALK | Rearrangements can drive cancer; target for ALK inhibitors. |
The study of molecular pathogenesis and genetic alterations is fundamental to understanding colorectal cancer (CRC). This sub-area explores the specific genetic mutations and molecular pathways that drive the initiation and progression of CRC. Key mutations in genes such as APC, KRAS, and p53 are frequently observed in colorectal tumors and play critical roles in uncontrolled cell growth, resistance to apoptosis, and metastasis. Additionally, research in this area investigates the epigenetic modifications and microsatellite instability that contribute to tumor heterogeneity and patient prognosis. By elucidating the genetic landscape of CRC, scientists can identify potential biomarkers for early detection, develop targeted therapies that specifically address the molecular abnormalities, and personalize treatment strategies to improve patient outcomes. Advances in genomic sequencing technologies have significantly enhanced our ability to characterize the complex genetic interactions in colorectal cancer, paving the way for innovative approaches in precision medicine.
The tumor microenvironment (TME) and its interaction with the immune system constitute a crucial area of colorectal cancer research. The TME consists of various cell types, including immune cells, fibroblasts, endothelial cells, and the extracellular matrix, which interact with cancer cells to influence tumor growth, invasion, and metastasis. Understanding these interactions is essential for developing effective immunotherapies. Research in this sub-area focuses on how immune checkpoints, cytokines, and cellular signaling pathways within the TME can be manipulated to enhance the immune system's ability to recognize and eliminate cancer cells. Additionally, the presence of specific immune cell populations, such as T cells and macrophages, within the TME has been linked to patient prognosis and response to therapy. By targeting the TME, scientists aim to overcome resistance to conventional treatments and improve the efficacy of immunotherapies like checkpoint inhibitors and CAR T-cell therapy. This approach holds promise for providing more durable and personalized treatment options for patients with colorectal cancer.