This website uses cookies to ensure you get the best experience on our website.
- Table of Contents
and ELISA kits, proteins related to developmental biology, stem cell research, lineage commitment, and tissue development.
Developmental biology studies how a single cell gives rise to organized tissues, organs, and body plans through coordinated programs of cell fate specification, proliferation, migration, differentiation, and morphogenesis. Antibodies are essential tools in developmental biology because they enable precise detection of lineage markers, pluripotency regulators, morphogens, transcription factors, adhesion proteins, and signaling nodes across embryos, stem cell systems, organoids, and differentiated tissues. In spatial assays such as IHC and IF, researchers can localize developmental markers within tissue architecture and monitor patterning events. In mechanistic studies, Western blot and ChIP help validate pathway activation, transcriptional control, and chromatin-associated regulation, while flow cytometry supports stem and progenitor cell phenotyping. This developmental biology antibodies hub is designed to help researchers move quickly from developmental context to biomarkers, methods, pathways, and related research areas.
Developmental biology studies often begin with marker selection: pluripotency regulators for stem cell state, morphogens for tissue patterning, transcription factors for lineage commitment, and adhesion or signaling proteins for morphogenesis. The examples below highlight validated product pages and representative developmental biology targets that support embryogenesis, organogenesis, differentiation, and developmental signaling studies.
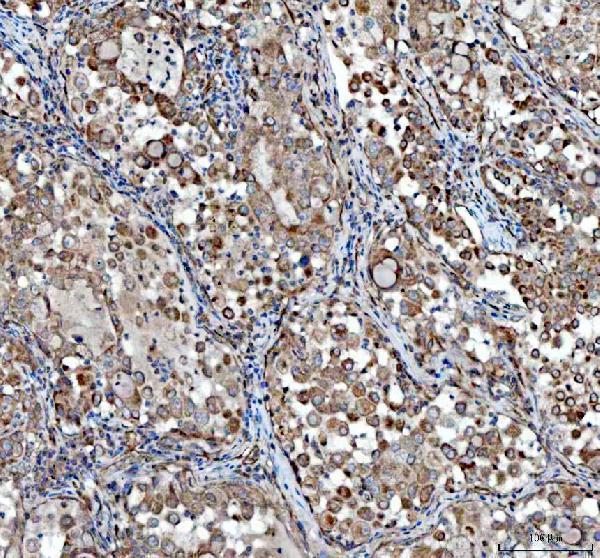
Anti-VEGF/VEGFA Antibody Picoband®, useful for angiogenesis, vascular remodeling, and tissue development studies where vessel formation and growth factor signaling are central readouts.
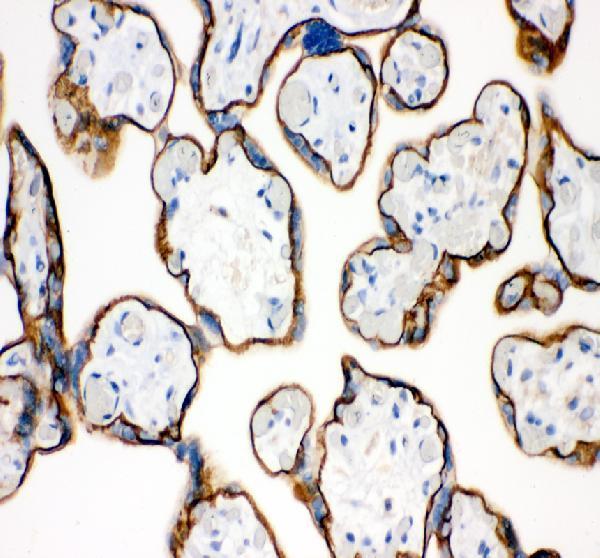
Anti-E Cadherin 1/CDH1 Antibody Picoband®, a core marker for epithelial identity, tissue organization, compaction, and developmental transitions involving cell-cell adhesion.
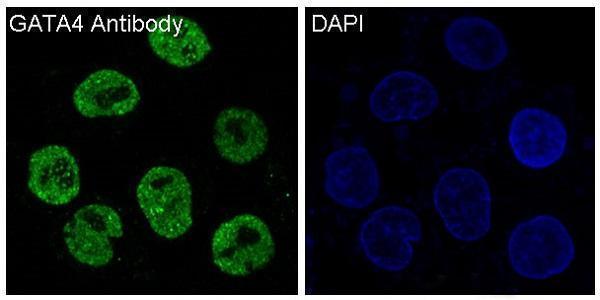
Anti-GATA4 Rabbit Monoclonal Antibody, relevant for developmental programs involving cardiac, endodermal, and organ-specific lineage specification.
| Protein Name | Gene Name | Function |
|---|---|---|
| Sonic Hedgehog | SHH | Regulates limb patterning, neural tube development, and morphogen-dependent tissue specification. |
| Wnt3a | WNT3A | Involved in embryonic patterning, cell fate decisions, and developmental signaling. |
| Fibroblast Growth Factor 8 | FGF8 | Critical for limb, brain, and axis development. |
| Bone Morphogenetic Protein 4 | BMP4 | Controls mesoderm formation, tissue patterning, and bone/cartilage development. |
| Notch1 | NOTCH1 | Controls cell fate choice, stem cell maintenance, and differentiation timing. |
| PAX6 | PAX6 | Key regulator of eye, neural, and brain development. |
| HOXA1 | HOXA1 | Determines body patterning along the anterior-posterior axis. |
| Vascular Endothelial Growth Factor | VEGFA | Promotes blood vessel formation and developmental angiogenesis. |
| Nodal | NODAL | Involved in gastrulation, germ layer formation, and mesoderm differentiation. |
| Oct4 | POU5F1 | Maintains pluripotency in embryonic stem cells and early developmental states. |
| Nanog | NANOG | Supports self-renewal and maintenance of undifferentiated stem cells. |
| SOX2 | SOX2 | Maintains stemness and contributes to neural and pluripotency programs. |
| GATA4 | GATA4 | Essential for heart, liver, and endoderm-related developmental programs. |
| E-cadherin | CDH1 | Mediates cell-cell adhesion required for compaction, epithelial organization, and tissue formation. |
| Retinoic Acid Receptor Alpha | RARA | Mediates retinoic acid signaling in differentiation and patterning processes. |
| Transforming Growth Factor Beta | TGFB1 | Regulates cell growth, differentiation, EMT-related processes, and tissue morphogenesis. |
| Nkx2.5 | NKX2-5 | Crucial for cardiac specification and heart development. |
| CDX2 | CDX2 | Involved in intestinal development, trophectoderm biology, and lineage specification. |
| Leukemia Inhibitory Factor | LIF | Supports maintenance of embryonic stem cell pluripotency in culture systems. |
| Embryonic Alkaline Phosphatase | ALPP | Marker of undifferentiated embryonic stem cells and early pluripotent states. |
Use IHC and IF to localize lineage markers, developmental transcription factors, adhesion proteins, and morphogen-responsive targets in embryos, tissues, organoids, and differentiation models.
Explore IHC/IF guideValidate developmental signaling pathways such as Wnt, Notch, Hedgehog, Hippo, and TGF-β/Smad, and confirm marker expression changes during stem cell maintenance or lineage differentiation.
Explore Western blot guideProfile stem cell populations, progenitor enrichment, and differentiation states at single-cell resolution. Flow cytometry is especially useful for sorting developmental subpopulations and monitoring lineage transitions.
Explore flow cytometry guideStudy how transcription factors, chromatin-associated regulators, and histone modifications control developmental gene expression programs during pluripotency, patterning, and differentiation.
Explore ChIP guideDevelopmental biology frequently begins with stem cell models, where researchers track pluripotency factors, early lineage priming, and differentiation cues. These systems are used to study self-renewal, fate restriction, and the earliest decision points in development.
Many developmental workflows focus on how stem or progenitor cells acquire tissue-specific identities. Neural development, cardiogenesis, osteogenesis, and other organ programs require precise marker panels to distinguish progenitors, intermediate states, and mature cell types.
Development does not depend on lineage markers alone. Morphogenesis also requires coordinated cell movement, epithelial integrity, extracellular matrix remodeling, and vascular support. These contexts are important when studying compaction, branching, EMT-like transitions, and tissue maturation.
Early developmental studies often focus on how pluripotent cells maintain stemness or commit toward specific germ layers and lineages. Marker panels in this stage commonly include OCT4, SOX2, NANOG, ALPP, LIF, and signaling readouts linked to pluripotency maintenance.
Once fate decisions begin, embryos rely on morphogens and signaling gradients to coordinate pattern formation. Hedgehog, Wnt, BMP, FGF, and Notch pathways shape tissue domains, regional identity, and developmental timing across multiple systems.
Later developmental stages emphasize organ formation, structural maturation, and tissue remodeling. Researchers often combine lineage markers, organ-specific transcription factors, adhesion proteins, and vascular markers to understand how tissues assemble and stabilize.
Core pathway maps for embryonic stem cell state, lineage bias, and developmental marker interpretation.
Classical developmental pathways that control patterning, growth, self-renewal, differentiation, and tissue identity.
Developmental transitions involving adhesion, extracellular matrix interaction, and cell plasticity.
Gene regulatory networks define how developmental programs are turned on, maintained, and resolved across time and space. These networks integrate transcription factors, enhancers, chromatin regulators, and extracellular signaling inputs to control lineage commitment, tissue patterning, and organ-specific differentiation. In developmental biology, mapping these regulatory relationships helps explain why the same genome can generate highly specialized cell types in a predictable sequence. Antibodies against transcription factors, chromatin-associated proteins, and pathway intermediates are therefore central to mechanistic studies of developmental control.
Stem cells provide the entry point for understanding how undifferentiated cells generate the full diversity of tissues in a developing organism. Developmental biology studies in this area focus on self-renewal, pluripotency maintenance, progenitor expansion, and the signaling events that direct differentiation into neural, mesodermal, endodermal, and organ-specific lineages. Reliable antibodies against pluripotency markers, lineage factors, and developmental signaling proteins are essential for distinguishing intermediate states and validating differentiation trajectories.
Development is not only about cell identity but also about how cells move, adhere, polarize, and assemble into functional tissues. Morphogenesis depends on epithelial organization, extracellular matrix interactions, cytoskeletal dynamics, EMT-like transitions, and vascular remodeling. These mechanisms are especially important when studying compaction, tube formation, branching structures, boundary formation, and organ architecture. Adhesion markers, ECM-related targets, and pathway maps for EMT and junction dynamics are therefore highly relevant for developmental biology workflows.