This website uses cookies to ensure you get the best experience on our website.
- Table of Contents
Antibodies, ELISA kits, and gene targets for stem cell biology, pluripotency, lineage commitment, tissue regeneration, and regenerative biology research.
Stem cell and regenerative biology research explores how pluripotent, multipotent, and tissue-resident progenitor cells maintain self-renewal, respond to niche signals, commit to specific lineages, and coordinate tissue repair, remodeling, and regeneration across developmental and injury-response contexts.
This field includes embryonic stem cells, induced pluripotent stem cells (iPSCs), adult stem cells, mesenchymal stem cells, and organ-specific progenitor populations involved in bone, cartilage, muscle, neural, cardiac, and vascular regeneration. Researchers often study how stemness programs are maintained, how differentiation is triggered, and how regenerative microenvironments influence repair outcomes in vitro and in vivo.
Antibodies are essential tools in stem cell and regenerative biology because they help identify pluripotent states, validate reprogramming, track lineage differentiation, profile regenerative niches, and measure pathway activity across organoids, tissue sections, primary cells, and cell culture models. This page is designed to help researchers navigate common biomarkers, experimental methods, lineage-specific contexts, pathway maps, and related research areas for stem cell and regeneration-focused studies.
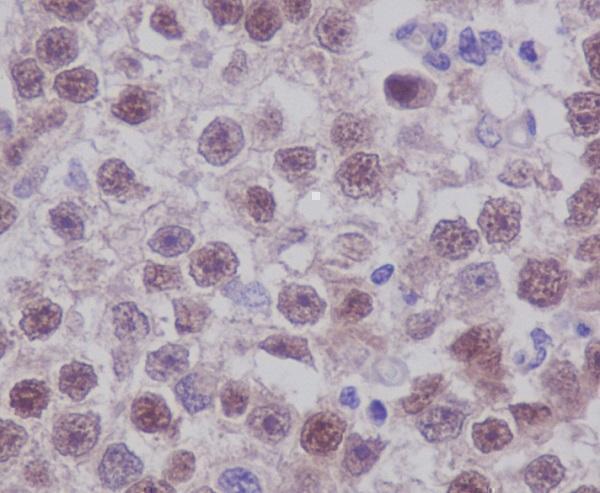
Anti-Oct4 POU5F1 Rabbit Monoclonal Antibody, IHC analysis of Oct4. OCT4 is a core pluripotency regulator frequently used to validate reprogramming and stem cell state maintenance.
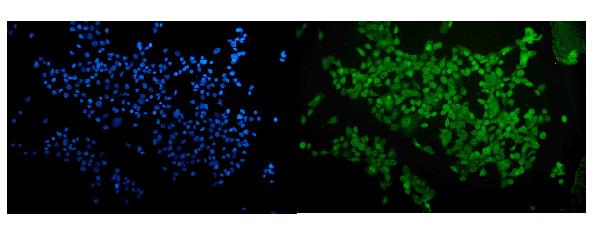
Anti-RUNX2 Antibody Picoband®, IF analysis of RUNX2. RUNX2 is a key osteogenic differentiation marker commonly used in bone regeneration and stem cell lineage commitment studies.
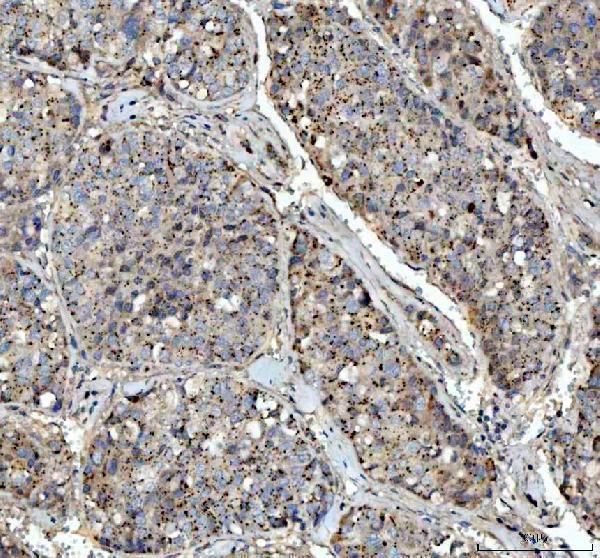
Anti-VEGF/VEGFA Antibody Picoband®, IHC analysis of VEGFA. VEGFA is a widely used target in angiogenesis, tissue repair, and regenerative microenvironment research.
| Protein Name | Gene Name | Function |
|---|---|---|
| OCT4 | POU5F1 | Core pluripotency factor used to maintain embryonic stem cell identity and validate reprogramming. |
| SOX2 | SOX2 | Supports self-renewal, pluripotency, and neural progenitor identity in stem cell models. |
| NANOG | NANOG | Maintains undifferentiated stem cell states and helps prevent premature lineage commitment. |
| KLF4 | KLF4 | Reprogramming-associated transcription factor involved in pluripotency and stem cell maintenance. |
| c-MYC | MYC | Regulates proliferation, metabolism, and reprogramming efficiency in stem cell systems. |
| CD34 | CD34 | Common marker for hematopoietic stem cells and endothelial progenitor populations. |
| CD133 | PROM1 | Surface marker used to identify stem and progenitor cells in multiple tissues. |
| CD73 | NT5E | Mesenchymal stem cell-associated marker linked to stromal signaling and immunomodulatory function. |
| CD90 | THY1 | Widely used mesenchymal stem cell marker involved in adhesion and stromal identity. |
| CD105 | ENG | Endoglin-associated mesenchymal and vascular marker relevant to angiogenesis and regenerative niches. |
| LGR5 | LGR5 | Adult stem cell marker used in intestinal, epithelial, and tissue-resident progenitor studies. |
| ABCG2 | ABCG2 | Associated with side-population stem cells and stem-like efflux phenotypes. |
| Nestin | NES | Intermediate filament marker widely used for neural stem and progenitor cell identification. |
| RUNX2 | RUNX2 | Essential transcription factor for osteogenic differentiation and bone regeneration studies. |
| SOX9 | SOX9 | Key regulator of chondrogenic differentiation and cartilage repair models. |
| GATA4 | GATA4 | Cardiac lineage marker used in heart development, differentiation, and regeneration research. |
| MyoD | MYOD1 | Master regulator of myogenic commitment and skeletal muscle regeneration workflows. |
| Pax7 | PAX7 | Satellite cell marker commonly used in muscle stem cell and repair studies. |
| BMP4 | BMP4 | Developmental and regenerative signaling factor that promotes lineage differentiation. |
| FGF2 | FGF2 | Supports stem cell proliferation, survival, and regenerative signaling across multiple model systems. |
| VEGF | VEGFA | Promotes angiogenesis and vascular remodeling in tissue repair and regenerative medicine studies. |
Use IHC and IF to localize pluripotency factors, lineage markers, and niche-associated proteins in tissue sections, organoids, spheroids, and differentiation models. Typical readouts include OCT4, SOX2, Nestin, RUNX2, SOX9, and GATA4.
Explore IHC/IF assay guideBuild phenotyping panels for hematopoietic, mesenchymal, or tissue-resident stem cell populations using surface markers such as CD34, CD73, CD90, CD105, CD133, and KIT. Flow cytometry is especially useful for enrichment, gating, and population heterogeneity analysis.
Explore flow cytometry guideConfirm stemness programs and signaling activity in cell lysates using markers such as OCT4, SOX2, NANOG, β-catenin, YAP/TAZ, SMADs, and Notch pathway components. Western blot remains a core method for pathway validation during reprogramming and differentiation.
Explore Western blot guideQuantify conditioned-media and model-system readouts such as VEGF, FGF2, BMP-related signals, repair-associated cytokines, and niche factors. ELISA is especially useful for studying angiogenesis, paracrine communication, and regeneration-linked signaling environments.
Explore ELISA assay guideOne of the most common entry points in stem cell biology is the study of pluripotency and induced reprogramming. Researchers frequently validate OCT4, SOX2, NANOG, KLF4, and c-MYC expression to confirm reprogramming efficiency, monitor stem cell maintenance, and compare undifferentiated versus lineage-committed states in ESC and iPSC workflows.
Adult stem cells and tissue progenitors are central to regeneration studies in hematopoietic, intestinal, neural, epidermal, and muscle systems. These workflows often rely on markers such as CD34, CD133, LGR5, KIT, ABCG2, and Nestin to define progenitor populations, side populations, or lineage-restricted regenerative compartments.
Mesenchymal stem cell research intersects with stromal signaling, extracellular matrix remodeling, immune regulation, and tissue repair. Marker combinations such as CD73, CD90, and CD105 are widely used to define MSC-like populations, while VEGFA and growth factor measurements help connect stromal activity to vascular remodeling and regenerative support.
Regenerative medicine studies frequently combine stem cell differentiation markers with angiogenic, matrix, and repair-associated readouts. Bone, cartilage, muscle, vascular, neural, and cardiac regeneration models often use RUNX2, SOX9, GATA4, MYOD1, PAX7, BMP4, FGF2, and VEGFA to connect lineage specification with functional tissue repair outcomes.
ESC and iPSC models are widely used to study pluripotency, epigenetic resetting, and directed differentiation. These systems are typically characterized using OCT4, SOX2, NANOG, KLF4, and c-MYC, often combined with pathway validation and IF-based morphology readouts.
Adult stem cell studies often focus on hematopoietic stem cells, neural progenitors, intestinal stem cells, and other tissue-specific regenerative pools. Marker combinations such as CD34, KIT, Nestin, LGR5, and ABCG2 help define progenitor identity, niche dependence, and regenerative capacity.
Mesenchymal and stromal models are especially relevant when studying tissue remodeling, scaffold responses, angiogenesis, and immune-regulatory repair environments. These contexts often intersect with fibroblast, endothelial, and matrix-associated biology.
Differentiation-oriented stem cell models often focus on organ-specific regeneration outcomes. These include osteoblast and chondrocyte differentiation, muscle repair, endothelial regeneration, and cardiac lineage commitment. Marker selection should match lineage context and regenerative endpoint.
Core pathways and transcriptional programs that maintain undifferentiated stem cell states and support reprogramming.
Signaling routes that shape differentiation trajectories, lineage commitment, and stem cell niche responses.
Pathways that connect mechanical signaling, angiogenesis, matrix remodeling, and regenerative tissue responses.
A central question in stem cell biology is how pluripotent cell states are established, stabilized, and lost. Core transcription factors such as OCT4, SOX2, and NANOG cooperate with chromatin regulators and signaling pathways to maintain self-renewal, while KLF4 and c-MYC contribute to cellular reprogramming and state transitions. These mechanisms are foundational for embryonic stem cell maintenance, iPSC generation, and studies of plasticity in regenerative systems.
Stem cells do not act in isolation. Their fate is shaped by soluble factors, extracellular matrix interactions, cell–cell contacts, and pathway inputs such as Wnt, Notch, TGF-β/Smad, and Hippo signaling. These niche-dependent mechanisms influence whether stem cells remain undifferentiated, expand, or commit to bone, cartilage, neural, muscle, cardiac, or vascular lineages. Antibody-based validation of lineage markers and pathway nodes is critical for interpreting these state transitions.
Regeneration depends not only on stem cell presence but also on successful integration with angiogenesis, stromal remodeling, and tissue-specific repair programs. Growth factors such as VEGFA, FGF2, and BMP4, along with lineage regulators such as RUNX2, SOX9, GATA4, MYOD1, and PAX7, help connect molecular identity to functional repair. These mechanisms are especially important in regenerative medicine, tissue engineering, and injury-repair model systems.